Abstract
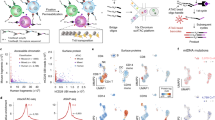
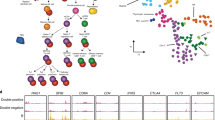
Recent technological advances have enabled researchers to accurately and efficiently assay the chromatin dynamics of scarce cell populations. In this Opinion article, we advocate the application of these technologies to central questions in immunology. Unlike changes to other molecular structures in the cell, chromatin features can reveal the past (developmental history), present (current activity) and future (potential response to challenges) of a given immune cell type; chromatin profiling is therefore an important new tool for studying the immune-regulatory networks of health and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Heng, T. S., Painter, M. W. & Immunological Genome Project Consortium. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Kim, C. C. & Lanier, L. L. Beyond the transcriptome: completion of act one of the Immunological Genome Project. Curr. Opin. Immunol. 25, 593–597 (2013).
Shay, T. & Kang, J. Immunological Genome Project and systems immunology. Trends Immunol. 34, 602–609 (2013).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Schones, D. et al. Dynamic regulation of nucleosome positioning in the human genome. Cell 132, 887–898 (2008).
Boyle, A. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Ostuni, R. et al. Latent enhancers activated by stimulation in differentiated cells. Cell 152, 157–171 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Bauer, D. E. et al. An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science 342, 253–257 (2013).
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 160, 351–352 (2014).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Lara-Astiaso, D. et al. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
Buenrostro, J., Giresi, P., Zaba, L., Chang, H. & Greenleaf, W. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Thurman, R. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Cusanovich, D. A. et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Greenleaf, W. J. Assaying the epigenome in limited numbers of cells. Methods 72, 51–56 (2015).
Cedar, H. & Bergman, Y. Epigenetics of haematopoietic cell development. Nat. Rev. Immunol. 11, 478–488 (2011).
Broske, A.-M. et al. DNA methylation protects hematopoietic stem cell multipotency from myeloerythroid restriction. Nat. Genet. 41, 1207–1215 (2009).
Ji, H. et al. Comprehensive methylome map of lineage commitment from haematopoietic progenitors. Nature 467, 338–342 (2010).
Shlush, L. I. et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature 506, 328–333 (2014).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Sun, D. et al. Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell 14, 673–688 (2014).
Giresi, P., Kim, J., McDaniell, R., Iyer, V. & Lieb, J. FAIRE (formaldehyde-assisted isolation of regulatory elements) isolates active regulatory elements from human chromatin. Genome Res. 17, 877–885 (2007).
Heintzman, N. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Creyghton, M. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Sanyal, A., Lajoie, B., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Kieffer-Kwon, K.-R. et al. Interactome maps of mouse gene regulatory domains reveal basic principles of transcriptional regulation. Cell 155, 1507–1520 (2013).
Cui, K. et al. Chromatin signatures in multipotent human hematopoietic stem cells indicate the fate of bivalent genes during differentiation. Cell Stem Cell 4, 80–93 (2009).
Bernstein, B. E., Mikkelsen, T. S., Xie, X. & Kamal, M. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2010).
Odink, K. et al. Two calcium-binding proteins in infiltrate macrophages of rheumatoid arthritis. Nature 330, 80–82 (1987).
Lam, M. T. Y. et al. Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature 498, 511–515 (2013).
Treiber, T. et al. Early B cell factor 1 regulates B cell gene networks by activation, repression, and transcription- independent poising of chromatin. Immunity 32, 714–725 (2010).
Garber, M. et al. A high-throughput chromatin immunoprecipitation approach reveals principles of dynamic gene regulation in mammals. Mol. Cell 47, 810–822 (2012).
Monticelli, S. & Natoli, G. Short-term memory of danger signals and environmental stimuli in immune cells. Nat. Immunol. 14, 777–784 (2013).
Netea, M. G., Quintin, J. & van der Meer, J. W. M. Trained immunity: a memory for innate host defense. Cell Host Microbe 9, 355–361 (2011).
Foster, S., Hargreaves, D. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007).
Ginhoux, F. & Jung, S. Monocytes and macrophages: developmental pathways and tissue homeostasis. Nat. Rev. Immunol. 14, 392–404 (2014).
Okabe, Y. & Medzhitov, R. Tissue-specific signals control reversible program of localization and functional polarization of macrophages. Cell 157, 1–13 (2014).
Varol, C., Mildner, A. & Jung, S. Macrophages: development and tissue specialization. Annu. Rev. Immunol. 33, 643–675 (2015).
Burzyn, D., Benoist, C. & Mathis, D. Regulatory T cells in nonlymphoid tissues. Nat. Immunol. 14, 1007–1013 (2013).
Yokoyama, W. M., Sojka, D. K., Peng, H. & Tian, Z. Tissue-resident natural killer cells. Cold Spring Harb. Symp. Quant. Biol. 78, 149–156 (2013).
Glasmacher, E. et al. A genomic regulatory element that directs assembly and function of immune-specific AP-1–IRF complexes. Science 338, 975–980 (2012).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Boyle, A. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011).
Pique-Regi, R. et al. Accurate inference of transcription factor binding from DNA sequence and chromatin accessibility data. Genome Res. 21, 447–455 (2011).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Cirillo, L. & Zaret, K. An early developmental transcription factor complex that is more stable on nucleosome core particles than on free DNA. Mol. Cell 4, 961–969 (1999).
Ostuni, R. & Natoli, G. Lineages, cell types and functional states: a genomic view. Curr. Opin. Cell Biol. 25, 759–764 (2013).
Winter, D. R. & Amit, I. The role of chromatin dynamics in immune cell development. Immunol. Rev. 261, 9–22 (2014).
Vilagos, B. et al. Essential role of EBF1 in the generation and function of distinct mature B cell types. J. Exp. Med. 209, 775–792 (2012).
Adams, D. et al. BLUEPRINT to decode the epigenetic signature written in blood. Nat. Biotechnol. 30, 224–226 (2012).
Martens, J. H. A. & Stunnenberg, H. G. BLUEPRINT: mapping human blood cell epigenomes. Haematologica 98, 1487–1489 (2013).
Pulendran, B. Systems vaccinology: Probing humanity's diverse immune systems with vaccines. Proc. Natl Acad. Sci. USA 111, 12300–12306 (2014).
Brodin, P. et al. Variation in the human immune system is largely driven by non-heritable influences. Cell 160, 37–47 (2015).
Ballester, B. et al. Multi-species, multi-transcription factor binding highlights conserved control of tissue-specific biological pathways. eLife 3, e02626 (2014).
Heinz, S. et al. Effect of natural genetic variation on enhancer selection and function. Nature 503, 487–492 (2013).
Rosengren Pielberg, G. et al. A cis-acting regulatory mutation causes premature hair graying and susceptibility to melanoma in the horse. Nat. Genet. 40, 1004–1009 (2008).
Degner, J. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
McDaniell, R. et al. Heritable individual-specific and allele-specific chromatin signatures in humans. Science 328, 235–239 (2010).
Mikkelsen, T. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Suzuki, T. et al. Pulmonary macrophage transplantation therapy. Nature 514, 450–454 (2014).
Richmond, T. & Davey, C. The structure of DNA in the nucleosome core. Nature 423, 145–150 (2003).
Felsenfeld, G. & Groudine, M. Controlling the double helix. Nature 421, 448–453 (2003).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Gross, D. S. & Garrard, W. T. Nuclease hypersensitive sites in chromatin. Annu. Rev. Biochem. 57, 159–197 (1988).
Amendola, M. & Van Steensel, B. Mechanisms and dynamics of nuclear lamina–genome interactions. Curr. Opin. Cell Biol. 28, 61–68 (2014).
Grewal, S. & Moazed, D. Heterochromatin and epigenetic control of gene expression. Science 301, 798–802 (2003).
Dixon, J. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Rao, Suhas, S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-wide mapping of in vivo protein-DNA interactions. Science 316, 1497–1502 (2007).
Acknowledgements
The authors thank G. Brodsky for drafting the artwork. Research in the laboratory of I.A. is supported by the European Research Council (309788) and the Israeli Science Foundation (712163), The Ernest and Bonnie Beutler Research Program of Excellence in Genomic Medicine and the Center for Excellence in Genome Science from the National Human Genome Research Institute (NHGRI; 1P50HG006193). Research in the laboratory of S.J. is supported by the Israeli Science Foundation (887/11), the European Research Council (340345) and the Deutsche Forschungsgemeinschaft (FOR1336). D.R.W. is grateful to the Azrieli Foundation for the Azrieli Fellowship award and for support from the European Molecular Biology Organization (EMBO; ALT766-2014) and the European Commission FP7 (Marie Curie Actions, EMBOCOFUND2012, GA-2012-600394).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Assay for transposase-accessible chromatin
-
(ATAC). A method for identifying regions of open chromatin by using Tn5 transposase to insert paired sequencing adaptors into accessible chromatin.
- Chromatin
-
The 3D complex of DNA and proteins within the nucleus. Features of chromatin (including localization, structure, interacting proteins, accessibility and modifications) regulate cell-type-specific gene expression.
- Chromatin immunoprecipitation
-
(ChIP). A method for identifying genomic regions that associate with a particular protein, including specifically modified histones, through immunoprecipitation of the crosslinked DNA fragments.
- Chromatin interaction analysis using paired-end tag sequencing
-
(ChIA-PET). A method for identifying pairs of regions associated with a particular protein by combining chromatin immunoprecipitation with the isolation of interacting DNA fragments.
- Cis-acting elements
-
Functional regulatory elements (such as enhancers) within the genome that regulate the transcription of genes.
- DNA methylation
-
The addition of a methyl group to a nucleic acid in the genome, typically to cytosine within a CpG pair.
- DNase hypersensitivity assay
-
A method that relies on the preference of the enzyme DNaseI to digest unbound DNA to identify regions of open chromatin.
- Enhancers
-
Distal regulatory elements that may function in combination with promoters or other enhancers to influence the transcription of one or more genes through the binding of transcription factors.
- Epigenome
-
Modifications to the genome that do not change the DNA sequence, including DNA methylation, histone modifications and rearrangements of chromatin structure.
- Expression quantitative trait loci
-
(eQTLs). Genomic loci that correlate with changes in gene expression analogously with the relationship between quantitative trait loci and phenotype.
- Formaldehyde-assisted isolation of regulatory elements
-
(FAIRE). A method for identifying regions of open chromatin by isolating DNA fragments that are not bound to DNA after crosslinking.
- Genome-wide association study
-
(GWAS). A statistical analysis comparing the occurrence of multiple common genetic variants (usually single-nucleotide polymorphisms) with a certain outcome (a phenotype or disease) to identify causal variants.
- Hi-C and 5C
-
High-throughput adaptations of the chromosome conformation capture (3C) method that isolates DNA fragments interacting with each other. 5C searches for interactions between many loci, whereas Hi-C searches for all possible interactions in the genome.
- Pioneer transcription factors
-
Sequence-specific DNA-binding proteins that target closed chromatin sites and recruit chromatin remodellers to transform these sites into an open chromatin state enabling the binding of additional transcription factors.
- Promoter
-
A proximal regulatory element that is typically located upstream of the particular gene it regulates and includes the binding sites of general transcription factors such as RNA polymerase II.
- Single-nucleotide polymorphisms
-
(SNPs). Mutations (substitutions, insertions or deletions) of an individual nucleotide in the genome sequence that are common in the population.
- Trans-acting binding factors
-
Non-DNA molecules (typically referring to transcription factors or chromatin factors) that regulate the transcription of genes.
Rights and permissions
About this article
Cite this article
Winter, D., Jung, S. & Amit, I. Making the case for chromatin profiling: a new tool to investigate the immune-regulatory landscape. Nat Rev Immunol 15, 585–594 (2015). https://doi.org/10.1038/nri3884
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri3884
This article is cited by
-
Impact of rare and common genetic variation in the interleukin-1 pathway on human cytokine responses
Genome Medicine (2021)
-
The origins of memory T cells
Nature (2017)
-
Rapid Recall Ability of Memory T cells is Encoded in their Epigenome
Scientific Reports (2017)
-
Epigenetic landscapes reveal transcription factors that regulate CD8+ T cell differentiation
Nature Immunology (2017)
-
Mathematical Models for Immunology: Current State of the Art and Future Research Directions
Bulletin of Mathematical Biology (2016)