Abstract
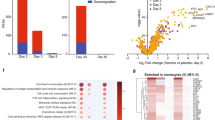
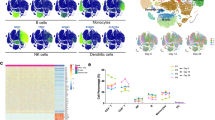
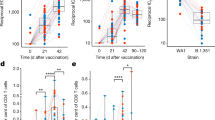
In this study, antibody response and a single-cell RNA-seq analysis were conducted on peripheral blood mononuclear cells from five different groups: naïve subjects vaccinated with AZD1222 (AZ) or Ad5-nCoV (Cso), individuals previously infected and later vaccinated (hybrid) with AZD1222 (AZ-hb) or Ad5-nCoV (Cso-hb), and those who were infected and had recovered from COVID-19 (Inf). The results showed that AZ induced more robust neutralizing antibody responses than Cso. The single-cell RNA data revealed a high frequency of memory B cells in the Cso and Cso-hb. In contrast, AZ and AZ-hb groups exhibited the highest proportion of activated naïve B cells expressing CXCR4. Transcriptomic analysis of CD4+ and CD8+ T cells demonstrated a heterogeneous response following vaccination, hybrid immunity, or natural infection. However, a single dose of Ad5-nCoV was sufficient to strongly activate CD4+ T cells (naïve and memory) expressing ANX1 and FOS, similar to the hybrid response observed with AZ. An interesting finding was the robust activation of a subset of CD8+ T cells expressing GZMB, GZMH, and IFNG genes in the Cso-hb group. Our findings suggest that both vaccines effectively stimulated the cellular immune response; however, the Ad5-nCoV induced a more robust CD8+ T-cell response in previously infected individuals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the article and its Supplementary information files (Table Source data). The single-cell RNA-seq data and BCR-seq data can be accessed in Zenodo (https://doi.org/10.5281/zenodo.7904759).
Code availability
The codes used to generate the results in the manuscript are available at: https://github.com/wanhui5867/adenovirus_C19_vaccines.
References
Baden LR, El Sahly HM, Essink B, Kotloff K, Frey S, Novak R, et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 Vaccine. N Engl J Med. 2021;384:403–16.
Widge AT, Rouphael NG, Jackson LA, Anderson EJ, Roberts PC, Makhene M, et al. Durability of responses after SARS-CoV-2 mRNA-1273 vaccination. N Engl J Med. 2021;384:80–2.
Andreano E, Paciello I, Piccini G, Manganaro N, Pileri P, Hyseni I, et al. Hybrid immunity improves B cells and antibodies against SARS-CoV-2 variants. Nature. 2021;600:530–5.
Chen Y, Tong P, Whiteman N, Moghaddam AS, Zarghami M, Zuiani A, et al. Immune recall improves antibody durability and breadth to SARS-CoV-2 variants. Sci Immunol. 2022;7:eabp8328.
Stamatatos L, Czartoski J, Wan YH, Homad LJ, Rubin V, Glantz H, et al. mRNA vaccination boosts cross-variant neutralizing antibodies elicited by SARS-CoV-2 infection. Science. 2021;372:1413–8.
Reynolds CJ, Pade C, Gibbons JM, Butler DK, Otter AD, Menacho K, et al. Prior SARS-CoV-2 infection rescues B and T cell responses to variants after first vaccine dose. Science. 2021;372:1418–23.
Wratil PR, Stern M, Priller A, Willmann A, Almanzar G, Vogel E, et al. Three exposures to the spike protein of SARS-CoV-2 by either infection or vaccination elicit superior neutralizing immunity to all variants of concern. Nat Med. 2022;28:496–503.
Gelanew T, Mulu A, Abebe M, Bates TA, Wassie L, Tefer M, et al. A single dose ChAdOx1 nCoV-19 vaccine elicits high antibody responses in individuals with prior SARS-CoV-2 infection comparable to that of double dose vaccinated SARS-CoV-2 infection naive individuals. Vaccines. 2022;10:859.
Guzman-Lopez S, Darwich-Salazar A, Bocanegra-Ibarias P, Salas-Trevino D, Flores-Trevino S, Perez-Alba E, et al. Clinical and immunologic efficacy of the recombinant adenovirus Type-5-Vectored (CanSino Bio) vaccine in university professors during the COVID-19 delta wave. Vaccines. 2022;10:656.
Cao Q, Wu S, Xiao C, Chen S, Chi X, Cui X, et al. Integrated single-cell analysis revealed immune dynamics during Ad5-nCoV immunization. Cell Discov. 2021;7:64.
Xiaojie S, Yu L, Lei Y, Guang Y, Min Q. Neutralizing antibodies targeting SARS-CoV-2 spike protein. Stem Cell Res. 2020;50:102125.
Fagiani F, Catanzaro M, Lanni C. Molecular features of IGHV3-53-encoded antibodies elicited by SARS-CoV-2. Signal Transduct Target Ther. 2020;5:170.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–87.e29.
Borcherding N, Bormann NL, Kraus G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res. 2020;9:47.
Bewley KR, Coombes NS, Gagnon L, McInroy L, Baker N, Shaik I, et al. Quantification of SARS-CoV-2 neutralizing antibody by wild-type plaque reduction neutralization, microneutralization and pseudotyped virus neutralization assays. Nat Protoc. 2021;16:3114–40.
Melgoza-Gonzalez EA, Hinojosa-Trujillo D, Resendiz-Sandoval M, Mata-Haro V, Hernandez-Valenzuela S, Garcia-Vega M, et al. Analysis of IgG, IgA and IgM antibodies against SARS-CoV-2 spike protein S1 in convalescent and vaccinated patients with the Pfizer-BioNTech and CanSinoBio vaccines. Transbound Emerg Dis. 2022;69:e734–45.
Bates TA, McBride SK, Leier HC, Guzman G, Lyski ZL, Schoen D, et al. Vaccination before or after SARS-CoV-2 infection leads to robust humoral response and antibodies that effectively neutralize variants. Sci Immunol. 2022;7:eabn8014.
Hernández J, Dehesa-Canseco F, Vázquez-López AB, Reséndiz-Sandoval M, Caire-Juvera G, Solís-Hernández M, et al. Neutralization of Omicron BA.1, BA.5.1.6, BQ.1.3 and XBB1.1 induced by heterologous vaccination Ad5-nCoV and mRNA-1273. Signal Transduct Target Ther. 2023;8:174.
Sherina N, Piralla A, Du L, Wan H, Kumagai-Braesch M, Andréll J, et al. Persistence of SARS-CoV-2-specific B and T cell responses in convalescent COVID-19 patients 6-8 months after the infection. Med. 2021;2:281–95.e4.
Marcotte H, Piralla A, Zuo F, Du L, Cassaniti I, Wan H, et al. Immunity to SARS-CoV-2 up to 15 months after infection. iScience. 2022;25:103743.
He B, Liu S, Wang Y, Xu M, Cai W, Liu J, et al. Rapid isolation and immune profiling of SARS-CoV-2 specific memory B cell in convalescent COVID-19 patients via LIBRA-seq. Signal Transduct Target Ther. 2021;6:195.
Nie Y, Waite J, Brewer F, Sunshine MJ, Littman DR, Zou YR. The role of CXCR4 in maintaining peripheral B cell compartments and humoral immunity. J Exp Med. 2004;200:1145–56.
Xiang H, Zhao Y, Li X, Liu P, Wang L, Wang M, et al. Landscapes and dynamic diversifications of B-cell receptor repertoires in COVID-19 patients. Hum Immunol. 2022;83:119–29.
Yuan M, Wu NC, Zhu X, Lee CD, So RTY, Lv H, et al. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science. 2020;368:630–3.
Liu H, Yuan M, Huang D, Bangaru S, Zhao F, Lee CD, et al. A combination of cross-neutralizing antibodies synergizes to prevent SARS-CoV-2 and SARS-CoV pseudovirus infection. Cell Host Microbe. 2021;29:806–18.e6.
Wen W, Su W, Tang H, Le W, Zhang X, Zheng Y, et al. Immune cell profiling of COVID-19 patients in the recovery stage by single-cell sequencing. Cell Discov. 2020;6:31.
Kalia V, Sarkar S. Regulation of effector and memory CD8 T cell differentiation by IL-2-A balancing act. Front Immunol. 2018;9:2987.
Yeh N, Glosson NL, Wang N, Guindon L, McKinley C, Hamada H, et al. Tc17 cells are capable of mediating immunity to vaccinia virus by acquisition of a cytotoxic phenotype. J Immunol. 2010;185:2089–98.
Hamada H, de la Luz Garcia-Hernandez M, Reome JB, Misra SK, Strutt TM, McKinstry KK, et al. Tc17, a unique subset of CD8 T cells that can protect against lethal influenza challenge. J Immunol. 2009;182:3469–81.
Notarbartolo S, Ranzani V, Bandera A, Gruarin P, Bevilacqua V, Putignano AR, et al. Integrated longitudinal immunophenotypic, transcriptional and repertoire analyses delineate immune responses in COVID-19 patients. Sci Immunol. 2021;6:eabg5021.
Acknowledgements
This research was funded by Consejo Nacional de Ciencia y Tecnología (CONACyT), grant number 312677 (JH), the Swedish Research Council (2019-01302, 2020-06116, QPH), and the European Union’s Horizon 2020 research and innovation program (ATAC, grant number 101003650, QPH). The authors thank Roberto Carvente and Analitek for technical support during the single-cell experiments and the SENASICA and WHO Collaborating Centre on Antimicrobial Resistance in Foodborne and Environmental Bacteria (MEX-33), especially Mayrén Zamora Nava and Fabiola Hernández Pérez, for helping us to sequence our libraries.
Funding
This work was supported by CONACyT (grant number 312677, JH), the Swedish Research Council (grants 2019-01302 and 2020-06116, QPH), and the European Union’s Horizon 2020 research and innovation program through the ATAC project under grant number 101003650 (QPH).
Author information
Authors and Affiliations
Contributions
Conceptualization, JH; methodology, MGV, HW, MRS, DHT, OV, VMH, FDC, MSH; formal analysis, MGV, HW; investigation, MGV, HW; resources, MSH, QPH, JH; writing—original draft preparation, MGV and HW, and JH; writing—review and editing; HM, QPH, JH; funding acquisition, QPH, JH. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
García-Vega, M., Wan, H., Reséndiz-Sandoval, M. et al. Comparative single-cell transcriptomic profile of hybrid immunity induced by adenovirus vector-based COVID-19 vaccines. Genes Immun 25, 158–167 (2024). https://doi.org/10.1038/s41435-024-00270-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41435-024-00270-x