Abstract
We analyzed 370 bp of the first hypervariable region of the mitochondrial DNA (mtDNA) control region in 752 individuals from 17 tribal and four nontribal groups from the Indian subcontinent, to address questions concerning the origins, genetic structure and relationships of these groups. Southern Indian tribes showed reduced diversity and large genetic distances, both among themselves and when compared with other groups, and no signal of prehistoric demographic expansions. These results probably reflect enhanced genetic drift because of small population sizes and/or bottlenecks in these groups. By contrast, northern groups exhibited more diversity and signals of prehistoric demographic expansions. Phylogenetic analyses revealed that southern and northern groups (except northeastern ones) have related mtDNA sequences albeit at different frequencies, further supporting the larger impact of drift on the genetic structure of southern groups. The Indian mtDNA gene pool appears to be more closely related to the east Eurasian gene pool (including central, east and southeast Asian populations) than the west Eurasian one (including European and Caucasian populations). Within India, northeastern tribes are quite distinct from other groups; they are more closely related to east Asians than to other Indians. This is consistent with linguistic evidence in that these populations speak Tibeto-Burman languages of east Asian origin. Otherwise, analyses of molecular variance suggested that caste and tribal groups are genetically similar with respect to mtDNA variation.
Similar content being viewed by others
Introduction
Archeological, fossil and genetic evidence points to a major expansion of anatomically modern humans out of Africa some 100 000 years ago,1,2 but the migration routes remain poorly understood. In this respect, the Indian subcontinent is considered to be a crucial geographic area for human migrations,3,4 since it is located at the crossroads of Africa, the Pacific and west and east Eurasia.
More than one billion people with enormous morphological, genetic, cultural and linguistic diversity inhabit the Indian subcontinent.5,6 At least four potential sources of genes contributing to the current Indian gene pool can be envisaged.3 The first one is an old Paleolithic component, probably almost extinct nowadays. The second component would have witnessed early Neolithic migrations of farmers from the eastern horn of the Fertile Crescent, probably speaking proto-Dravidian languages. The third source is responsible for the arrival of Indo-European speakers ∼3500 years ago, who most probably introduced the caste system that hierarchically organized the vast majority of Indian society. The fourth component is associated with Austro-Asiatic and Tibeto-Burman speakers inhabiting east and northeast India, with ties to east Asia.
The molecular genetic data generated so far concerning the people of the Indian subcontinent have largely focused on caste populations rather than on tribal groups.7,8,9,10,11,12,13 According to the 1991 census, ∼8% of the Indian population belong to tribal communities.14 They represent minorities that have not been absorbed by the caste system.3 They are generally thought to be the aboriginal inhabitants of the Indian subcontinent that were present in the region before the arrival of Indo-European speakers. There are currently about 400 tribes in India that vary in size from a few hundred to a few million; they speak languages belonging to all four of the major language families represented in India (Austro-Asiatic, Dravidian, Indo-European and Tibeto-Burman). Their origins and genetic affinities remain largely unknown, although such information is of primary importance in understanding the possible role of India in early migrations of modern humans, since any remnants of genetic contributions from pre-Indo-European migrants would presumably be present in tribal populations rather than in caste populations.
The molecular genetic evidence on Indian tribal origins and relationships is rather scanty. The mitochondrial DNA (mtDNA) intergenic COII/tRNALys 9-bp deletion marker, present at high frequency in some Asian15 and African16 populations, is found at low frequencies in India and arose multiple times independently.17,18,19 Based on a study of autosomal and mtDNA markers in eight Indian tribes speaking Austro-Asiatic, Dravidian or Tibeto-Burman languages, it was concluded that these different language groups represented distinct founding groups, with Austro-Asiatic speakers being the most ancient inhabitants of the region.19
With the aim of furthering our understanding of their origins and relationships, we report here a comprehensive study of mtDNA variation in Indian tribal groups. We analyzed sequences of the first hypervariable segment (HV1) of the mtDNA control region in more than 750 individuals belonging to different language families and compared these to available data from caste populations and from world populations, in order to obtain a global picture of the genetic structure and the relationships of populations inhabiting the Indian subcontinent.
Subjects and methods
Samples
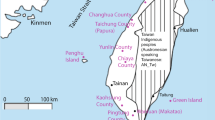
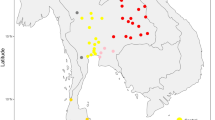
DNA samples from 752 individuals belonging to 21 populations inhabiting the Indian subcontinent were analyzed (Table 1, Figure 1). Additional information on the 17 tribal populations and the four nontribal populations can be found elsewhere.18,20 Published data were also included from 10 tribal populations,12,19 one Indo-Euro-pean-speaking caste from north India,12 eight Dravidian-speaking castes from Andhra Pradesh,10 and two castes from south India.7 Although they were sampled in the southern state of Andhra Pradesh, the Andh, Pardhi, Thoti and Lambadi tribes as well as the eight Dravidian-speaking castes10 will be referred to as central Indian populations in the following text, in order to differentiate them from the populations sampled in the states of Kerala, Karnataka and Tamil Nadu (Figure 1).
mtDNA sequencing
The mtDNA HV1 was amplified using primers L15996 (Vigilant et al21) and H16410 (Handt et al22). Purified PCR products were sequenced on both strands using primers L16001 (5′-ATTAGCACCCAAAGCTAAGA-3′) and H16401 (Vigilant et al21). Sequencing reactions were prepared with the BigDye Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA) and purified by isopropanol precipitation, according to the supplier's recommendations. Sequencing reactions were resolved on either ABI 377 or 3700 automated DNA sequencers (Applied Biosystems). The HV1 sequences were deposited in the HvrBase23 and are also available from the authors.
Data analysis
HV1 sequences were manually aligned with the published reference sequence.24 After removal of sites with insertions and deletions, the software package ARLEQUIN version 2.000 (Schneider et al25) was used to calculate haplotype and nucleotide diversity and their standard deviations (SD), mismatch distributions, mean pairwise differences and their SD, Fu's26 Fs statistic and associated P-values based on 1000 simulated samples, raggedness index r27 and Fst distances between pairs of populations and associated P-values based on 1000 permutations. Analyses of molecular variance28 (AMOVA) were performed using ARLEQUIN to evaluate the genetic structure of the populations, with the significance of variance components tested with 10 000 permutations.
Multidimensional scaling (MDS) was performed by means of STATISTICA, based on Fst distances. Populations from the Indian subcontinent were compared to world-wide populations from Africa (!Kung,29 Dinka,30 Turkana and Yoruba31), Europe (Germans,32 Bulgarians and Turks33), Caucasus (Armenians, Azerbaijanians, Georgians and Kabardinians34), central Asia (Kazakh, Kirghiz and Kirghiz-Talas35), east Asia (Han Chinese-Changsha, Han Chinese-Xi'an, Tottori,36 Japanese,37 Koreans38 and Mongolians39), southeast Asia (Akkha, White and Red Karen, Lahu, Lisu-Chiang Rai, Lisu-Mae Hong Son40 and Vietnamese36), Australia (from Arnhem Land, Sandy Desert,41 Desert and Riverine42), and island southeast Asia (Indonesians,29,43 coastal and highland Papua New Guineans41 (PNG)).
A gene tree was computed using the neighbor-joining method, based on p-distances, as implemented in MEGA version 2.1 (Kumar et al44). The tree was rooted using a Neanderthal HV1 sequence45 and bootstrap analysis was carried out with 500 replications. In addition to Indian sequences, worldwide HV1 sequences9 were included for comparison, as well as PNG highlanders, central Asians and Australians described above. We inferred the frequencies of some major mtDNA haplo-groups in our data from the resulting tree topology, from Indian HV1 sequences whose haplogroup information is available,12 and from haplogroup-diagnostic substitutions in HV1.46
Results
Diversity indices and demographic parameters
Sequence data corresponding to nucleotide positions (np) 16 022–16 391 in the reference sequence24 were obtained from 752 individuals. Nucleotide substitutions were observed at 153 sites, which defined 316 different HV1 sequences. Some individuals exhibited length variation between np 16 181 and 16 183 (these positions were removed from the subsequent analyses). Deletions were observed at five sites (np 16 166, 16 179, 16 194, 16 195 and 16 258) and insertions at two sites (a C between np 16 169 and 16 170 and an A between np 16 189 and 16 190).
The 752 HV1 sequences from the present study were subsequently analyzed together with 219 previously published sequences from 10 Indian tribes, enabling a more comprehensive study of HV1 variation in Indian tribal populations. Since small sample sizes could potentially affect the reliability of the analyses, some populations were pooled. Pooling was performed according to several criteria including geographical proximity, linguistic affiliation, historical record and population relationships deduced from Fst distances. For example, the Mullukurunan and Mullukurumba are both south Indian tribes speaking a Kannada language (a Dravidian language-subfamily). Some scholars have argued that they are the same although they nowadays live in different areas, and it has been reported that the Mullukurunan are also known as ‘Mullu Kurumba’.47 Moreover, the two tribes were separated by an Fst distance of −0.030, which is not significantly different from zero (P=0.84). Therefore the data from these two groups were pooled. In summary, this approach resulted in 23 groups, each of which was composed of at least 20 individuals (Table 2).
Diversity indices and demographic parameters estimated for these groups are reported in Table 2. Overall, haplotype diversity in Indian tribals ranged from 0.671 to 0.995 and nucleotide diversity from 0.005 to 0.023. Haplotype diversity was significantly higher (Mann–Whitney U-test: Z=3.24, P<0.01) in north, east and northeast India (0.940–0.995) than in south India (0.671–0.939). Intermediate haplotype diversity values were observed in central India (0.884–0.985); they were not significantly different from north, east and northeast India (Z=1.51, P=0.13) or south India (Z=1.70, P=0.09). Similarly, nucleotide diversity in north, east and northeast India (0.014–0.023) was significantly higher (Z=2.78, P<0.01) than in south India (0.005–0.017). Again, central India exhibited intermediate values (0.012–0.017); they were not significantly different from north, east and northeast India (Z=1.61, P=0.11) or south India (Z=1.78, P=0.07). Therefore, north, east and northeast Indian tribes showed greater mtDNA diversity than south Indian tribes.
These patterns of genetic diversity in Indian tribes were further strengthened by the analysis of mean pairwise differences (MPD). MPD for south tribes (1.77–5.80) were significantly lower than MPD from north, east and northeast tribes (5.06–7.69; Z=2.78, P<0.01) or from central tribes (4.46–5.91; Z=2.04, P=0.04), whereas MPD from central and north, east and northeast tribes were not significantly different (Z=1.70, P=0.09). Mismatch distributions (Figure 2) were computed for the 14 tribal and four nontribal populations from Table 1 whose sample size is ≥20. Unimodal distributions were observed mostly in northeast tribes and some central tribes, whereas all of the south tribes exhibited multimodal distributions with a higher frequency of the low difference classes (0 and 1): 25–61% of the pairwise differences for the south tribes were in the 0/1 classes vs only 1–13% for the other tribes. Unimodal distributions are interpreted as signs of demographic expansions while multimodal distributions are interpreted as signs of constant population size over time.27 Moreover, the peaks observed at 0/1 classes in the mismatch distributions indicated bottlenecks in these populations.48 In parallel, the raggedness index r was generally less than 0.03 in north, east and northeast tribes but more than 0.07 in south tribes (Table 2). Values of r lower than 0.05 suggest demographic expansions while values of r greater than 0.05 are more consistent with constant population sizes.27 Fu's Fs also support these patterns of demographic history in India (Table 2). Negative values of Fs that differ significantly from zero, indicative of population demographic expansions,26 were obtained in 86% of north, east and northeast tribes, 50% of central tribes and only 25% of south tribes. Therefore, several approaches provided congruent evidence for different demographic histories in Indian tribes. In general, north, east and northeast tribes showed signs of expansion while south tribes, and to a lesser extent central tribes, were likely to have experienced bottlenecks and/or constant population sizes over time.
The four nontribal populations exhibited high gene and nucleotide diversities (Table 2). In addition, the Fs statistic as well as the raggedness index (Table 2) and mismatch distributions (Figure 2) suggested demographic expansions in these populations.
MDS analysis
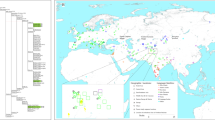
MDS analysis based on Fst distances was performed to investigate relationships between Indian tribes, Indian castes and world populations. In the MDS plot (Figure 3), Indian tribal and caste populations both clustered together with other Eurasian and Australian populations. The Kung were a clear outlier, but removing them from the analysis did not significantly change the results, except as noted below (analysis not shown).
MDS plot of 66 world populations, based on Fst distances. The area enclosed by a solid line is magnified in Figure 4.
Among Indian populations, most of the south tribes were outliers within the Eurasian/Australian cluster. Further information on the affinities of Indian tribes was obtained by focussing on the Eurasian/Australian cluster in the MDS plot (Figure 4). Overall, Indian subcontinent populations were closer to east Eurasians (including central, east and southeast Asians) than to west Eurasians (including Europeans and Caucasians). Since the smaller number of European populations considered might have contributed to suggesting stronger affinities between India and east Eurasia, the analysis was repeated by adding six European populations (from Sweden, Austria, France, Britain, Italy and Spain). However, the trends in the MDS plot did not change (not shown). The smaller average Fst distance separating Indian populations from east Eurasians (0.081) than from west Eurasians (0.118) confirmed the aforementioned affinities. The removal of northeast tribes from the analysis led to similar average Fst values (0.087 for India–east Eurasia vs 0.116 for India–west Eurasia) and the removal of south tribes resulted in ∼25% smaller average Fst values (0.058 for India–east Eurasia vs 0.093 for India–west Eurasia). The removal of both northeast and south tribes was almost equivalent to removing only south tribes (0.061 for India–east Eurasia vs 0.083 for India–west Eurasia). In all cases, then, Indian populations showed closer affinities to east than to west Eurasians.
In contrast to other tribal groups, the five northeast Indian groups were closer to east Eurasian populations (average Fst distance: 0.049) than to other Indian tribes (average Fst distance: 0.118, dropping to 0.084 when south tribes were removed). The same trend was observed for east Indian tribes; however, when the !Kung were removed from the analysis, east Indian tribes were closer to other Indian groups than to east Asians in an MDS plot (not shown). Average Fst values also supported their closer affinities to Indian tribes (0.048 with south and northeast tribes excluded; 0.084 if they are included) than to east Eurasians (0.068).
Gene tree
The outlier positions of south tribes in the MDS plot (Figure 3), coupled with demographic inferences indicating bottlenecks in these populations, mean that either south tribes have different HV1 sequences than other groups, or related sequences at very different frequencies. To distinguish between these two hypotheses, a neighbor-joining tree was constructed for 553 Indian and 420 worldwide HV1 sequences and rooted with a Neanderthal sequence (Figure 5). To facilitate the analysis, it was subdivided into 14 clusters (I–XIV). The deepest cluster (cluster I) was almost exclusively African-specific, with the exception of a single sequence from a south Indian tribe (Kuruchian). Sequences from the south tribes were found in all of the clusters, as were sequences from at least one other Indian population (with the exception of cluster I). This suggested that south tribes do not have different HV1 sequences than other groups, but rather related sequences at different frequencies.
Summary information of a gene tree of 973 mtDNA HV1 sequences, rooted with a Neanderthal sequence. The topology of the tree is represented on the left-hand side; n is the number of sequences in each cluster (I–XIV). The table also provides the number of sequences in each cluster sampled in the following geographic areas: Africa, Australia, Papua-New-Guinea (PNG), West Eurasia (W Eurasia), East Eurasia (E Eurasia) and India (‘NE tribes’: northeast tribes; ‘S tribes’: south tribes).
The gene tree also suggested a close relationship between east Eurasians and northeast Indians in that 90% of the sequences in cluster XIII belonged either to east Eurasians or northeast Indians. In addition, 93% of the sequences from cluster XIV belonged either to Indian tribes (excluding northeast tribes) or castes, suggesting close relationships between these groups.
mtDNA haplogroup affiliation
mtDNA haplogroups are defined by RFLPs, but by using information from the gene tree, published data on Indian HV1 sequences for which the mtDNA haplogroup was known, and diagnostic HV1 mutations, we were able to infer the haplogroup affiliation for 90% of the Indian sequences (Table 3). Most Indian sequences belonged to the Asian haplogroup M, as found previously.9,12,19 South, central and east tribes exhibited very similar high frequencies of haplogroup M (∼75%). North tribes showed a somewhat lower frequency (∼67%), with a correspondingly higher frequency of the west Eurasian haplogroup JT (∼7%). Haplogroup M frequency in northeast tribes was found to be lower (∼56%) than in other Indian regions. The northeast tribes were also distinguished by a combined frequency of the east Asian haplogroups A and F of ∼32%, while these two haplogroups were virtually absent elsewhere in the Indian subcontinent.
Genetic structure of Indian populations
AMOVA was used to investigate the genetic structure of Indian populations, focussing either on tribes only or on both tribes and castes (Table 4). In the total tribal sample (model 1), 88% of the variance was found within populations and 12% among populations. Indian tribes were then grouped according to geographic proximity (model 2), to linguistic affinities (model 3) and to the results suggested by the MDS analysis, namely two groups separating northeast tribes from all others (model 4). Under these models, 86–88% of the variance was found within populations, 10% among populations within groups and 2–4% among groups. A model that accurately reflects the genetic structure should maximize the variance among groups and minimize the variance among populations within groups; therefore, none of these models provides a good description of the genetic structure, although model 4 is the best.
When the northeast, central and south groups of populations were analyzed separately, 22% of the variance was among populations in south tribes but only 3–5% in central and northeast tribes. These results emphasize the distinctiveness of south tribes from one another that was also evident in the MDS plots (Figure 3 and Figure 4). Therefore, the analyses corresponding to models 1–4 were repeated with the south tribes excluded (not shown). Again, none of these models provided a good description of the genetic structure. The best model grouped populations on the basis of linguistic criteria, and was the only model for which the ‘among groups’ component was larger than the ‘among populations within groups’ component.
We also compared tribes to caste populations. East and northeast tribes were excluded from the analyses since: (i) no HV1 data from east and northeast castes are avail-able, and (ii) these tribes speak Austro-Asiatic and Tibeto-Burman languages (respectively) which are spoken exclusively by tribal populations (preventing linguistically based analyses between tribes and castes). For the model based on social criteria (ie castes vs tribes; Table 4, model 5), 91% of the variance was within populations and 9% among populations within groups. The variance among groups did not differ significantly from zero (P=0.33), suggesting that the social distinction of castes vs tribes does not accurately reflect the genetic structure of Indian populations. Removing south groups from the analysis did not change the picture (not shown), with the variance among groups still not differing significantly from zero (P=0.57).
Discussion
Origins of tribal groups
Our analyses of mtDNA variation in tribal populations of India indicate that groups in different geographic regions have different demographic histories. In general, southern tribes have reduced mtDNA diversity and mismatch distributions strongly indicative of recent bottlenecks. The distinctiveness of southern groups is also emphasized by the MDS analyses and AMOVA. However, it is difficult to distinguish from these data between old and severe bottlenecks or more recent and less severe bottlenecks. Present-day population sizes in southern tribes tend to be small (ie generally less than 30 000; Table 1), as compared to northern tribes (ie generally over 100 000; Table 1). Consequently, genetic drift could have generated large genetic distances both among southern groups and between southern and other groups, thereby masking their real affinities to other populations.48 According to this scenario, southern populations have related mtDNA sequences (albeit at different frequencies) and hence a shared history with other Indian populations. An alternative hypothesis is that southern tribes have a specific mtDNA gene pool as compared to other Indian populations, indicating a long period of isolation and/or different history from other tribal groups. An example of the latter is PNG,41,49 in which most sequences are found in two clusters (within clusters III and XI, Figure 5) characterized by long branches (not shown). However, southern Indian sequences are intermingled throughout the tree, clustering with sequences from multiple populations. In addition, south, central, and east Indian tribes all have similar mtDNA haplogroup compositions (Table 3). These results provide strong support for the hypothesis that southern tribes have mtDNA sequences closely related to those of other tribes, but with different frequencies, which would suggest fairly recent bottlenecks occurring in these populations.
A possible cause of these bottlenecks, put forth by Excoffier and Schneider,48 involves Neolithic human expansions. According to this hypothesis, the recent settlement of Indo-European speakers in India some 3500 years ago3 might have had a major impact on the demography of south tribes. West Eurasian mtDNA haplogroups H, JT and W represent 6–7% of north and central tribes (Table 3), which are located in the area where Indo-European languages are spoken. In contrast, these west Eurasian mtDNA types are virtually absent in south tribes, which are located where Dravidian languages are spoken. This might reflect different responses of local people to the Indo-European settlement of India. In the north and center, Indo-Europeans may have admixed with local people,50 concomitant with the spread of Indo-European languages. In contrast, in the southern part of India, local populations may have challenged the arrival of Indo-European newcomers, resulting in limited admixture, reduction of tribal population sizes and retention of their original languages, thus explaining why Dravidian languages survived the spread of Indo-European languages in south India.
Tibeto-Burman speakers from northeast India show closer genetic affinities with east Asian groups than with other Indian groups. This is suggested by the MDS analyses and mtDNA haplogroup composition, in that northeast Indian tribes possess haplogroups A and F, which are frequent in east Asians but virtually absent from other regions of India. The mtDNA evidence (Clark et al18, this study) thus agrees with Y chromosome evidence51 as well as linguistic evidence,52 indicating a probable east Asian origin of these particular tribes. Archeological, linguistic and genetic evidence51 suggests that proto-Tibeto-Burman languages arose 5000–6000 years ago in east Asia. The fact that Tibeto-Burman speaking tribes from India have retained genetic traces of east Asian origins for such a long time suggests that, despite the more recent migrations to India, these populations remained relatively isolated, explaining the close correlation between genetic and linguistic results. This contrasts with the situation observed in other regions in the world, for example in Scandinavia53 and the Caucasus,34 where migrations led to language replacements and hence to incongruencies between genetic and linguistic results.
Apart from northeast tribes, all other Indian tribes exhibited a similar and high frequency of mtDNA haplo-group M (ie 56% in northeast vs ∼75% for others). These results are strikingly similar to a previous study based on RFLP analysis of a smaller set of Indian tribes,19 according to which haplogroup M had a frequency of ∼51% in northeast tribes and ∼76% in east, central and south tribes. This homogeneity in frequency of haplogroup M further supports the view that south tribes have sequences closely related to those of their neighboring populations. Thus, our mtDNA data are compatible with at least three major sources for the present-day mtDNA gene pool of Indian tribes: (i) a major one associated with all non-northeast tribes (whatever their linguistic or geographic ties), with a high frequency of mtDNA haplogroup M; (ii) one associated with Tibeto-Burman speakers from northeast India, with affinities to east Asians; and (iii) a third one associated with the presence of west Eurasian-typical mtDNA haplogroups (ie haplogroups H, JT and W, which represent 6–7% of mtDNA types in north and central tribes), most probably attributable to admixture with recent Indo-European-speaking migrants to India.50
Relationships between castes and tribes
The comparison between Indian castes and tribes revealed no strong difference between them, as pointed out by the AMOVA. A possible explanation for the observed similarities in caste and tribal mtDNA gene pools is common ancestry, with a proto-Asian origin of Indian castes.9 An alternative hypothesis involves a proto-west-Eurasian origin of castes, with the present-day similarities in caste and tribal mtDNA gene pools then being attributable to recent admixture with local Indian populations. The latter hypothesis would require extensive gene flow, which could seem a priori to be incompatible with the mating practices imposed by the caste system in India.6 However, there is evidence for female gene flow between Hindu castes10 and it has been suggested that male gene flow in south Indian populations may not be as negligible54 as previously thought.11 Our results show the presence of west-Eurasian typical mtDNA haplogroups in Indian tribes, presumably resulting from admixture with Indo-Europeans (ie who probably introduced the caste system in India). This interpretation would suggest that caste people initially possessed west-Eurasian mtDNAs rather than Asian mtDNAs. This view is reinforced by the fact that caste groups are more similar to west Eurasians (average Fst: 0.080) than are the tribals (average Fst: 0.149; and 0.117 if south tribes are excluded). Therefore, the similarities in caste and tribal mtDNA gene pools might reflect extensive maternal gene flow rather than common ancestry. However, caste and tribal populations are separated by an average Fst distance of 0.049 (if south tribes are excluded; 0.082 if they are included), suggesting that overall, castes are closer to Indian tribes than to west Eurasians. This makes the hypothesis of proto-Asian ancestry of castes equally plausible. The mtDNA data alone do not support one hypothesis over the other. Moreover, caste populations from different regions of India may have different origins,55 some of them being derived from west Eurasian ancestors with subsequent admixture with local populations, others being derived from local population ancestors via acculturation.
Relationships with other populations
mtDNA variation in India suggests that overall, Indian tribes show more affinities to east Eurasians than to west Eurasians. This means that migrations from the west (ie involving Indo-Europeans and Neolithic expansions of farmers) have not had a major distorting impact on the original gene pool. This view is consistent with the relatively small proportion of west Eurasian typical mtDNA haplogroups present in Indian tribes. On the other hand, three typical east-Asian mtDNA haplogroups (A, B and F) are absent or virtually absent from non-northeast India (Bamshad et al9 Kivisild et al,12 Roychoudhury et al,19 this study). Furthermore, the fourth typical east Asian mtDNA haplogroup M has a different structure in India as compared to other Asian areas.9 This suggests that, although they show close affinities, the east Asian and Indian mtDNA gene pools are fairly distinct. This result is consistent with the suggestion that the east Asian and Indian mtDNA pools have been separated from each other for about 30 000 years.49
It has been hypothesized that the peopling of Sahul (PNG and Australia) may have been the result of an early migration from east Africa through the Indian subcontinent following the ‘southern route’.1,3,56 Australian populations exhibited an average Fst distance of 0.067 with east Eurasians and of 0.089 with Indians (but only 0.062 if South tribes excluded), whereas the average Fst values separating Australians from PNG or African (!Kung excluded) populations were 0.194 and 0.145, respectively. These results suggested close genetic affinities between Australian populations and both Indian and east Eurasian populations. An India–Australia connection is consistent with other mtDNA41 and Y chromosome57 evidence. Taken together with other conclusions,41,57 the present results give credence to the trihybrid model of peopling of Australia58 involving ‘Negrito’, east Asian and Indian sources. The Indian influence on Australia may be recent (ie <5000 years),41,57 thus much later than (and therefore independent from) the early migration that would have followed the southern route ∼60 000 years ago.
In addition, Forster et al49 have proposed an mtDNA control region motif (16223C and 16357C) which could represent a signature of an early migration from Africa to Sahul through the southern route. This motif was not found in any Indian tribal mtDNA; 16357C had a frequency of only 2.1% and was always associated with 16223T, while 16223C had a frequency of 27.9%. Furthermore, Indians do not show particular affinities to Africans. A possible exception is the typical African HV1 sequence found in a Kuruchian from south India. However, there are communities in India such as the Siddis, who are known to be recent migrants from Africa.6 The African-like sequence found in India could therefore originate from admixture between recent African migrants and Indian tribals, or it may represent a remnant of an ancient migration from Africa to India; it is difficult to draw conclusions from a single sequence.
In summary, although the data support a recent India–Australia connection, we could not find in Indian tribals any unquestionable genetic signature of the ∼60 000 year-old migration from Africa to Sahul following the postulated southern route. A possible explanation would be that such migration never occurred along that route. Alternatively, the early migrants from Africa may have made their way to Sahul following the southern route without settling in India. Another possibility, which is probably the most reasonable one, is that in India the genetic traces of early migrations along the southern route were erased by the subsequent migrations which shaped the present-day mtDNA gene pool of India.3
References
Lahr MM, Foley R : Multiple dispersals and modern human origins. Evol Anthropol 1994; 3: 48–60.
Templeton AR : Out of Africa again and again. Nature 2002; 416: 45–51.
Cavalli-Sforza LL, Menozzi P, Piazza A : History and geography of human genes. Princeton, Princeton University Press, 1994.
Cann RL : Genetic clues to dispersal of human populations: retracing the past from the present. Science 2001; 291: 1742–1748.
Papiha SS : Genetic variation in India. Hum Biol 1996; 68: 607–628.
Majumder PP : People of India: biological diversity and affinities. Evol Anthropol 1998; 6: 100–113.
Mountain JL, Hebert JM, Bhattacharyya S et al: Demographic history of India and mtDNA-sequence diversity. Am J Hum Genet 1995; 56: 979–992.
Bamshad M, Fraley AE, Crawford MH et al: MtDNA variation in caste populations of Andhra Pradesh. India. Hum Biol 1996; 68: 1–28.
Bamshad M, Kivisild T, Watkins WS et al: Genetic evidence on the origins of Indian caste populations. Genome Res 2001; 11: 994–1004.
Bamshad MJ, Watkins WS, Dixon ME et al: Female gene flow stratifies Hindu castes. Nature 1998; 395: 651–652.
Bhattacharyya NP, Basu P, Das M et al: Negligible male gene flow across ethnic boundaries in India, revealed by analysis of Y-chromosomal DNA polymorphisms. Genome Res 1999; 9: 711–719.
Kivisild T, Bamshad MJ, Kaldma K et al: Deep common ancestry of Indian and western-Eurasian mitochondrial DNA lineages. Curr Biol 1999; 9: 1331–1334.
Majumder PP, Roy B, Banerjee S et al: Human-specific insertion/deletion polymorphisms in Indian populations and their possible evolutionary implications. Eur J Hum Genet 1999; 7: 435–446.
Singh KS : People of India: the scheduled tribes. Delhi: Oxford University Press, 1994, (vol III.)
Merriwether DA, Friedlaender JS, Mediavilla J, Mgone C, Gentz F, Ferrel RE : Mitochondrial DNA variation is an indicator of Austronesian influence in Island Melanesia. Am J Phys Anthropol 1999; 110: 243–270.
Soodyall H, Vigilant L, Hill AV, Stoneking M, Jenkins T : MtDNA control-region sequence variation suggests multiple independent origins of an ‘Asian-specific’ 9-bp deletion in Sub-Saharan Africans. Am J Hum Genet 1996; 58: 595–608.
Watkins WS, Bamshad M, Dixon ME et al: Multiple origins of the mtDNA 9-bp deletion in populations of South India. Am J Phys Anthropol 1999; 109: 147–158.
Clark VJ, Sivendren S, Saha N et al: The 9-bp deletion between the mitochondrial lysine tRNA and COII genes in tribal populations of India. Hum Biol 2000; 72: 273–285.
Roychoudhury S, Roy S, Basu A et al: Genomic structures and population histories of linguistically distinct tribal groups of India. Hum Genet 2001; 109: 339–350.
Melton T, Peterson R, Redd AJ et al: Polynesian genetic affinities with Southeast Asian populations as identified by mtDNA analysis. Am J Hum Genet 1995; 57: 403–414.
Vigilant L, Pennington R, Harpending H, Kocher TD, Wilson AC : Mitochondrial DNA sequences in single hairs from a southern African population. Proc Natl Acad Sci USA 1989; 86: 9350–9354.
Handt O, Krings M, Ward RL, Pääbo S : The retrieval of ancient human DNA sequences. Am J Hum Genet 1996; 59: 368–376.
Burckhardt F, von Haeseler A, Meyer S : HvrBase: compilation of mtDNA control region sequences from primates. Nucleic Acids Res 1999; 27: 138–142.
Anderson S, Bankier AT, Barrell BG et al: Sequence and organization of the human mitochondrial genome. Nature 1981; 290: 457–465.
Schneider S, Roessli D, Excoffier L : Arlequin ver. 2.000: a software for population genetics data analysis. University of Geneva, Switzerland: Genetics and Biochemistry Laboratory, 2000.
Fu YX : Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 1997; 147: 915–925.
Harpending HC, Sherry ST, Rogers AR, Stoneking M : The genetic structure of ancient human populations. Curr Anthropol 1993; 34: 483–496.
Excoffier L, Smouse P, Quattro J : Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 1992; 131: 479–491.
Vigilant L, Stoneking M, Harpending H, Hawkes K, Wilson AC : African populations and the evolution of human mitochondrial DNA. Science 1991; 253: 1503–1507.
Krings M, Salem AH, Bauer K et al: MtDNA analysis of Nile River Valley populations: a genetic corridor or a barrier to migration? Am J Hum Genet 1999; 64: 1166–1176.
Watson E, Bauer K, Aman R, Weiss G, von Haeseler A, Pääbo S : MtDNA sequence diversity in Africa. Am J Hum Genet 1996; 59: 437–444.
Hofmann S, Jaksch M, Bezold R et al: Population genetics and disease susceptibility: characterization of central European haplogroups by mtDNA gene mutations, correlation with D-loop variants and association with disease. Hum Mol Genet 1997; 6: 1835–1846.
Calafell F, Underhill P, Tolun A, Angelicheva D, Kalaydjieva L : From Asia to Europe: mitochondrial DNA sequence variability in Bulgarians and Turks. Ann Hum Genet 1996; 60: 35–49.
Nasidze I, Stoneking M : Mitochondrial DNA variation and language replacements in the Caucasus. Proc R Soc London B 2001; 268: 1197–1206.
Comas D, Calafell F, Mateu E et al: Trading genes along the Silk Road: mtDNA sequences and the origin of Central Asian populations. Am J Hum Genet 1998; 63: 1824–1838.
Oota H, Kitano T, Jin F et al: Extreme mtDNA homogeneity in continental Asian populations. Am J Phys Anthropol 2002; 118: 146–153.
Seo Y, Stradmann-Bellinghausen B, Rittner C, Takahama K, Schneider PM : Sequence polymorphism of mitochondrial DNA control region in Japanese. Forensic Sci Int 1998; 97: 155–164.
Lee SD, Shin CH, Kim KB, Lee YS, Lee JB : Sequence variation of mitochondrial DNA control region in Koreans. Forensic Sci Int 1997; 87: 99–116.
Kolman CJ, Sambuughin N, Bermingham E : Mitochondrial DNA analysis of mongolian populations and implications for the origin of New World founders. Genetics 1996; 142: 1321–1334.
Oota H, Settheetham-Ishida W, Tiwawech D, Ishida T, Stoneking M : Human mtDNA and Y-chromosome variation is correlated with matrilocal vs. patrilocal residence. Nat Genet 2001; 29: 20–21.
Redd AJ, Stoneking M : Peopling of Sahul: mtDNA variation in Aboriginal Australian and Papua New Guinean populations. Am J Hum Genet 1999; 65: 808–828.
van Holst Pellekaan SM, Frommer M, Sved JA, Boettcher B : Mitochondrial control-region sequence variation in Aboriginal Australians. Am J Hum Genet 1998; 62: 435–449.
Redd AJ, Takezaki N, Sherry S, McGarvey S, Sofro ASM, Stoneking M : Evolutionary history of the COII/tRNALys intergenic 9 base pair deletion in human mitochondrial DNAs from the Pacific. Mol Biol Evol 1995; 12: 604–615.
Kumar S, Tamura K, Jakobsen IB, Nei M : MEGA2: Molecular evolutionary genetics analysis software. Tempe, AZ, USA: Arizona State University, 2001.
Krings M, Stone A, Schmitz RW, Krainitzki H, Stoneking M, Pääbo S : Neandertal DNA sequences and the origin of modern humans. Cell 1997; 90: 19–30.
Macaulay V, Richards M, Hickey E et al: The emerging tree of West Eurasian mtDNAs: a synthesis of control-region sequences and RFLPs. Am J Hum Genet 1999; 64: 232–249.
Singh KS : People of India: India's communities. Delhi: Oxford University Press, 1998, Vol IV–VI.
Excoffier L, Schneider S : Why hunter-gatherer populations do not show signs of Pleistocene demographic expansions? Proc Natl Acad Sci USA 1999; 96: 10597–10602.
Forster P, Torroni A, Renfrew C, Röhl A : Phylogenetic star contraction applied to Asian and Papuan mtDNA evolution. Mol Biol Evol 2001; 18: 1864–1881.
Passarino G, Semino O, Bernini LF, Santachiara-Benerecetti AS : Pre-Caucasoid and Caucasoid genetic features of the Indian population, revealed by mtDNA polymorphisms. Am J Hum Genet 1996; 59: 927–934.
Su B, Xiao C, Deka R et al: Y chromosome haplotypes reveal prehistorical migrations to the Himalayas. Hum Genet 2000; 107: 582–590.
Matisoff JA : Sino-Tibetan linguistics: present state and future prospects. Annu Rev Anthropol 1991; 20: 469–504.
Sajantila A, Pääbo S : Language replacement in Scandinavia. Nat Genet 1995; 11: 359–360.
Ramana GV, Su B, Jin L et al: Y-chromosome SNP haplotypes suggest evidence of gene flow among caste, tribe, and the migrant Siddi populations of Andhra Pradesh, South India. Eur J Hum Genet 2001; 9: 695–700.
Majumder PP : Indian caste origins: genomic insights and future outlook. Genome Res 2001; 11: 931–932.
Quintana-Murci L, Semino O, Bandelt HJ, Passarino G, McElreavey K, Santachiara-Benerecetti AS : Genetic evidence on an early exit of Homo sapiens sapiens from Africa through eastern Africa. Nat Genet 1999; 23: 437–441.
Redd AJ, Roberts-Thomson J, Karafet T et al: Gene flow from the Indian subcontinent to Australia: evidence from the Y chromosome. Curr Biol 2002; 12: 673–677.
Birdsell JB : Preliminary data on the tri-hybrid origin of the Australian Aborigines. Archeol Phys Anthropol Oceania 1967; 2: 100–155.
Acknowledgements
We are grateful to the original donors of samples. We thank Birgit Nickel and Carsten Schwarz for technical assistance. We also thank Partha Majumder, Michael Bamshad and Scott Watkins for kindly providing HV1 sequences, and Silke Brauer, Manfred Kayser, Vano Nasidze, Hiroki Oota, Brigitte Pakendorf, Anthony Ryan and Alice Salzat for fruitful discussions during the course of this study. This project was supported by funds from the Max Planck Society, Germany.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cordaux, R., Saha, N., Bentley, G. et al. Mitochondrial DNA analysis reveals diverse histories of tribal populations from India. Eur J Hum Genet 11, 253–264 (2003). https://doi.org/10.1038/sj.ejhg.5200949
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200949
Keywords
This article is cited by
-
Complete mitogenomes document substantial genetic contribution from the Eurasian Steppe into northern Pakistani Indo-Iranian speakers
European Journal of Human Genetics (2021)
-
Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations
Communications Biology (2020)
-
Carriers of mitochondrial DNA macrohaplogroup L3 basal lineages migrated back to Africa from Asia around 70,000 years ago
BMC Evolutionary Biology (2018)
-
Neolithic phylogenetic continuity inferred from complete mitochondrial DNA sequences in a tribal population of Southern India
Genetica (2018)
-
Carriers of human mitochondrial DNA macrohaplogroup M colonized India from southeastern Asia
BMC Evolutionary Biology (2016)