« Prev Next »

Ribosomes, Transcription, and Translation
The genetic information stored in DNA is a living archive of instructions that cells use to accomplish the functions of life. Inside each cell, catalysts seek out the appropriate information from this archive and use it to build new proteins — proteins that make up the structures of the cell, run the biochemical reactions in the cell, and are sometimes manufactured for export. Although all of the cells that make up a multicellular organism contain identical genetic information, functionally different cells within the organism use different sets of catalysts to express only specific portions of these instructions to accomplish the functions of life.
How Is Genetic Information Passed on in Dividing Cells?
When a cell divides, it creates one copy of its genetic information — in the form of DNA molecules — for each of the two resulting daughter cells. The accuracy of these copies determines the health and inherited features of the nascent cells, so it is essential that the process of DNA replication be as accurate as possible (Figure 1).

One factor that helps ensure precise replication is the double-helical structure of DNA itself. In particular, the two strands of the DNA double helix are made up of combinations of molecules called nucleotides. DNA is constructed from just four different nucleotides — adenine (A), thymine (T), cytosine (C), and guanine (G) — each of which is named for the nitrogenous base it contains. Moreover, the nucleotides that form one strand of the DNA double helix always bond with the nucleotides in the other strand according to a pattern known as complementary base-pairing — specifically, A always pairs with T, and C always pairs with G (Figure 2). Thus, during cell division, the paired strands unravel and each strand serves as the template for synthesis of a new complementary strand.

What Are the Initial Steps in Accessing Genetic Information?
![]()
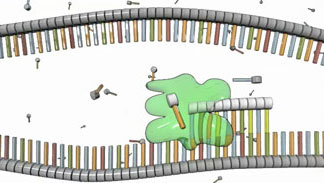
RNA molecules differ from DNA molecules in several important ways: They are single stranded rather than double stranded; their sugar component is a ribose rather than a deoxyribose; and they include uracil (U) nucleotides rather than thymine (T) nucleotides (Figure 4). Also, because they are single strands, RNA molecules don't form helices; rather, they fold into complex structures that are stabilized by internal complementary base-pairing.

mRNA is the most variable class of RNA, and there are literally thousands of different mRNA molecules present in a cell at any given time. Some mRNA molecules are abundant, numbering in the hundreds or thousands, as is often true of transcripts encoding structural proteins. Other mRNAs are quite rare, with perhaps only a single copy present, as is sometimes the case for transcripts that encode signaling proteins. mRNAs also vary in how long-lived they are. In eukaryotes, transcripts for structural proteins may remain intact for over ten hours, whereas transcripts for signaling proteins may be degraded in less than ten minutes.
Cells can be characterized by the spectrum of mRNA molecules present within them; this spectrum is called the transcriptome. Whereas each cell in a multicellular organism carries the same DNA or genome, its transcriptome varies widely according to cell type and function. For instance, the insulin-producing cells of the pancreas contain transcripts for insulin, but bone cells do not. Even though bone cells carry the gene for insulin, this gene is not transcribed. Therefore, the transcriptome functions as a kind of catalog of all of the genes that are being expressed in a cell at a particular point in time.
What Is the Function of Ribosomes?

Ribosomes are complexes of rRNA molecules and proteins, and they can be observed in electron micrographs of cells. Sometimes, ribosomes are visible as clusters, called polyribosomes. In eukaryotes (but not in prokaryotes), some of the ribosomes are attached to internal membranes, where they synthesize the proteins that will later reside in those membranes, or are destined for secretion (Figure 6). Although only a few rRNA molecules are present in each ribosome, these molecules make up about half of the ribosomal mass. The remaining mass consists of a number of proteins — nearly 60 in prokaryotic cells and over 80 in eukaryotic cells.
Within the ribosome, the rRNA molecules direct the catalytic steps of protein synthesis — the stitching together of amino acids to make a protein molecule. In fact, rRNA is sometimes called a ribozyme or catalytic RNA to reflect this function.
Eukaryotic and prokaryotic ribosomes are different from each other as a result of divergent evolution. These differences are exploited by antibiotics, which are designed to inhibit the prokaryotic ribosomes of infectious bacteria without affecting eukaryotic ribosomes, thereby not interfering with the cells of the sick host.

How Does the Whole Process Result in New Proteins?
After the transcription of DNA to mRNA is complete, translation — or the reading of these mRNAs to make proteins — begins. Recall that mRNA molecules are single stranded, and the order of their bases — A, U, C, and G — is complementary to that in specific portions of the cell's DNA. Each mRNA dictates the order in which amino acids should be added to a growing protein as it is synthesized. In fact, every amino acid is represented by a three-nucleotide sequence or codon along the mRNA molecule. For example, AGC is the mRNA codon for the amino acid serine, and UAA is a signal to stop translating a protein — also called the stop codon (Figure 7).

Molecules of tRNA are responsible for matching amino acids with the appropriate codons in mRNA. Each tRNA molecule has two distinct ends, one of which binds to a specific amino acid, and the other which binds to the corresponding mRNA codon. During translation, these tRNAs carry amino acids to the ribosome and join with their complementary codons. Then, the assembled amino acids are joined together as the ribosome, with its resident rRNAs, moves along the mRNA molecule in a ratchet-like motion. The resulting protein chains can be hundreds of amino acids in length, and synthesizing these molecules requires a huge amount of chemical energy (Figure 8).
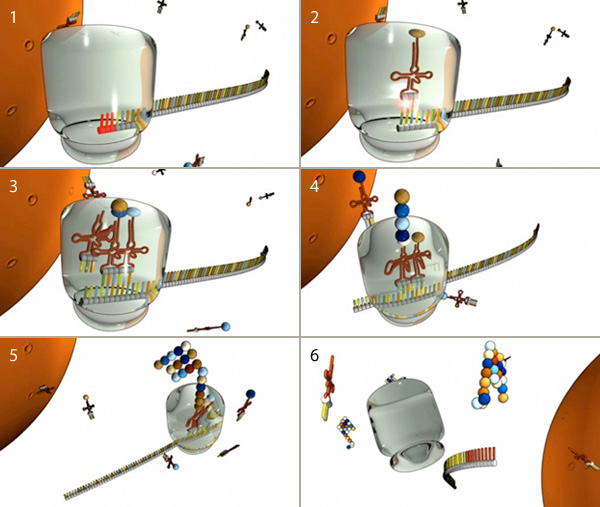
In prokaryotic cells, transcription (DNA to mRNA) and translation (mRNA to protein) are so closely linked that translation usually begins before transcription is complete. In eukaryotic cells, however, the two processes are separated in both space and time: mRNAs are synthesized in the nucleus, and proteins are later made in the cytoplasm.
Conclusion
eBooks
This page appears in the following eBook