Featured
-
-
Article |
Structural mechanism of angiogenin activation by the ribosome
- Anna B. Loveland
- , Cha San Koh
- & Andrei A. Korostelev
-
Article
| Open Access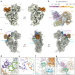 Structural basis of ribosomal 30S subunit degradation by RNase R
Structural basis of ribosomal 30S subunit degradation by RNase R
Cryo-electron microscopy structures of intermediates formed during the degradation of the 30S ribosomal unit shed light on how the 3′ to 5′ exonuclease ribonuclease R controls the ribosomal degradation process.
- Lyudmila Dimitrova-Paternoga
- , Sergo Kasvandik
- & Helge Paternoga
-
Article
| Open Access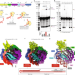 Template and target-site recognition by human LINE-1 in retrotransposition
Template and target-site recognition by human LINE-1 in retrotransposition
Human LINE-1 ORF2p relies on upstream single-stranded target DNA to position the adjacent duplex in the endonuclease active site for nicking of the longer DNA strand, with a single nick generating a staggered DNA break.
- Akanksha Thawani
- , Alfredo Jose Florez Ariza
- & Kathleen Collins
-
Article |
 mRNA reading frame maintenance during eukaryotic ribosome translocation
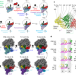
mRNA reading frame maintenance during eukaryotic ribosome translocation
The accuracy of eukaryotic ribosome translocation relies on eukaryote-specific elements of the 80S ribosome, elongation factor 2 and transfer RNAs, all of which contribute to the maintenance of the messenger RNA reading frame.
- Nemanja Milicevic
- , Lasse Jenner
- & Gulnara Yusupova
-
Article
| Open Access Structural insights into intron catalysis and dynamics during splicing
Structural insights into intron catalysis and dynamics during splicing
Analysis of the group II intron ribonucleoprotein shows the molecular interactions involved in branchpoint adenosine recognition, lariat formation and exon ligation, providing clues to the evolutionary conservation of structural components and catalytic mechanisms in premessenger RNA splicing.
- Ling Xu
- , Tianshuo Liu
- & Anna Marie Pyle
-
Article
| Open Access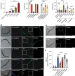 m1A in CAG repeat RNA binds to TDP-43 and induces neurodegeneration
m1A in CAG repeat RNA binds to TDP-43 and induces neurodegeneration
TDP-43 binds to N1-methyladenosine on CAG repeat RNA, resulting in the formation of gel-like TDP-43 aggregates in the cytoplasm that resemble those observed in neurological disease pathology.
- Yuxiang Sun
- , Hui Dai
- & Yinsheng Wang
-
Article
| Open Access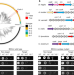 Antiviral type III CRISPR signalling via conjugation of ATP and SAM
Antiviral type III CRISPR signalling via conjugation of ATP and SAM
The Bacteroides fragilis type III CRISPR protein Cmr conjugates ATP to S-adenosyl methionine, generating S-adenosyl methionine (SAM)-AMP, a novel second messenger with a role in antiviral signalling.
- Haotian Chi
- , Ville Hoikkala
- & Malcolm F. White
-
Article
| Open Access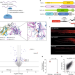 piRNA processing by a trimeric Schlafen-domain nuclease
piRNA processing by a trimeric Schlafen-domain nuclease
The endoribonuclease PUCH, a trimer of Schlafen-like-domain proteins, initiates piRNA processing in the nematode Caenorhabditis elegans through 5′-end piRNA precursor cleavage.
- Nadezda Podvalnaya
- , Alfred W. Bronkhorst
- & René F. Ketting
-
Article
| Open Access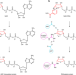 A viral ADP-ribosyltransferase attaches RNA chains to host proteins
A viral ADP-ribosyltransferase attaches RNA chains to host proteins
Bacteriophage T4 uses an enzyme known as ADP-ribosyltransferase ModB to modify the translational apparatus of bacteria it infects, not only by ADP-ribosylating proteins, but also by attaching entire RNA chains in a process known as RNAylation.
- Maik Wolfram-Schauerte
- , Nadiia Pozhydaieva
- & Katharina Höfer
-
Article |
 Oligomerization-mediated activation of a short prokaryotic Argonaute
Oligomerization-mediated activation of a short prokaryotic Argonaute
Cryo-electron microscopy structures and biochemical analyses provide insight into how short prokaryotic Argonaute proteins are assembled and activated, and reveal that oligomerization has a key role in driving catalytic activity.
- Zhangfei Shen
- , Xiao-Yuan Yang
- & Tian-Min Fu
-
Article |
 Hepatitis C virus RNA is 5′-capped with flavin adenine dinucleotide
Hepatitis C virus RNA is 5′-capped with flavin adenine dinucleotide
Hepatitis C virus utilizes flavin adenine dinucleotide as a non-canonical initiating nucleotide for the viral RNA polymerase, resulting in 5′ capping of viral RNA, which provides protection against the host innate immune response.
- Anna V. Sherwood
- , Lizandro R. Rivera-Rangel
- & Jeppe Vinther
-
Article
| Open Access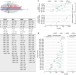 Relaxed targeting rules help PIWI proteins silence transposons
Relaxed targeting rules help PIWI proteins silence transposons
Of the two types of Argonaute proteins produced by animals, AGO and PIWI, PIWI proteins can bind RNAs with less complementarity, enabling efficient silencing of transposons without the need to produce new RNA guides.
- Ildar Gainetdinov
- , Joel Vega-Badillo
- & Phillip D. Zamore
-
Article
| Open Access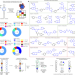 Programming inactive RNA-binding small molecules into bioactive degraders
Programming inactive RNA-binding small molecules into bioactive degraders
Small-molecule RNA-targeted degradation can be leveraged to convert strong, yet inactive, binding interactions into potent and specific modulators of RNA function.
- Yuquan Tong
- , Yeongju Lee
- & Matthew D. Disney
-
Article |
 RNA conformational propensities determine cellular activity
RNA conformational propensities determine cellular activity
Systematic alteration of HIV-1 TAR RNA and quantitative determination of its propensity to bind to the Tat protein establish a key role role for a rare and short-lived RNA state in Tat-dependent transactivation in cells.
- Megan L. Ken
- , Rohit Roy
- & Hashim M. Al-Hashimi
-
Article
| Open Access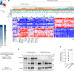 RBFOX2 modulates a metastatic signature of alternative splicing in pancreatic cancer
RBFOX2 modulates a metastatic signature of alternative splicing in pancreatic cancer
Analysis of messenger RNA splicing in a large cohort of pancreatic ductal adenocarcinoma tumours identifies differential splicing correlating with disease progression, associated with the the splicing regulator RBFOX2.
- Amina Jbara
- , Kuan-Ting Lin
- & Rotem Karni
-
Article |
 Nucleolar URB1 ensures 3′ ETS rRNA removal to prevent exosome surveillance
Nucleolar URB1 ensures 3′ ETS rRNA removal to prevent exosome surveillance
A high-resolution microscopy screen of candidate nucleolar proteins identifies URB1 as a protein that is confined to the periphery of the dense fibrillar component, with key roles in pre-ribosomal RNA folding and processing.
- Lin Shan
- , Guang Xu
- & Ling-Ling Chen
-
Article |
 mRNA ageing shapes the Cap2 methylome in mammalian mRNA
mRNA ageing shapes the Cap2 methylome in mammalian mRNA
Cap2 methylation increases on transcripts as they age, reducing activation of innate immunity.
- Vladimir Despic
- & Samie R. Jaffrey
-
Article
| Open Access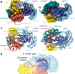 RNA targeting unleashes indiscriminate nuclease activity of CRISPR–Cas12a2
RNA targeting unleashes indiscriminate nuclease activity of CRISPR–Cas12a2
Structural analysis of Cas12a2, a CRISPR-associated nuclease that nonspecifically cleaves ssRNA, ssDNA and dsDNA, reveals a complete activation pathway involved in the abortive infection system protecting cells against invasion.
- Jack P. K. Bravo
- , Thomson Hallmark
- & David W. Taylor
-
Article |
 Structures and mechanisms of tRNA methylation by METTL1–WDR4
Structures and mechanisms of tRNA methylation by METTL1–WDR4
Using cryo-electron microscopy, structural and mechanistic insights into how the METTL1–WDR4 complex catalyses methylation of tRNAs are shown.
- Victor M. Ruiz-Arroyo
- , Rishi Raj
- & Yunsun Nam
-
Article
| Open Access Cas12a2 elicits abortive infection through RNA-triggered destruction of dsDNA
Cas12a2 elicits abortive infection through RNA-triggered destruction of dsDNA
RNA targeting by the Sulfuricurvum type V single-effector nuclease SuCas12a2 drives abortive infection through non-specific cleavage of double-stranded DNA—after recognition of an RNA target through an activating protospacer-flanking sequence, SuCas12a2 efficiently degrades ssRNA, ssDNA and dsDNA.
- Oleg Dmytrenko
- , Gina C. Neumann
- & Chase L. Beisel
-
Article
| Open Access Principles of mitoribosomal small subunit assembly in eukaryotes
Principles of mitoribosomal small subunit assembly in eukaryotes
The structures of both human and yeast mitochondrial ribosomal small subunits undergoing assembly are uncovered.
- Nathan J. Harper
- , Chloe Burnside
- & Sebastian Klinge
-
Article
| Open Access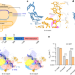 R-loop formation and conformational activation mechanisms of Cas9
R-loop formation and conformational activation mechanisms of Cas9
Cryo-electron microscopy structures of Streptococcus pyogenes Cas9 in multiple DNA-bound states provide insights on the mechanism of Cas9 activation by target DNA.
- Martin Pacesa
- , Luuk Loeff
- & Martin Jinek
-
Article
| Open Access The mechanism of RNA capping by SARS-CoV-2
The mechanism of RNA capping by SARS-CoV-2
Reconstitution of the SARS-CoV-2 RNA 5′ cap reveals the unconventional mechanism by which SARS-CoV-2 caps its RNA genome, providing a new target in the development of antiviral agents to treat COVID-19.
- Gina J. Park
- , Adam Osinski
- & Vincent S. Tagliabracci
-
Article
| Open Access GTSF1 accelerates target RNA cleavage by PIWI-clade Argonaute proteins
GTSF1 accelerates target RNA cleavage by PIWI-clade Argonaute proteins
The evolutionarily conserved RNA-binding protein GTSF1 and its homologues interact with members of the PIWI class of Argonaute proteins, increasing the efficiency of the RNA-cleaving activity of PIWI proteins, an essential function across the animal kingdom.
- Amena Arif
- , Shannon Bailey
- & Phillip D. Zamore
-
Article
| Open Access Structural insights into dsRNA processing by Drosophila Dicer-2–Loqs-PD
Structural insights into dsRNA processing by Drosophila Dicer-2–Loqs-PD
Structures of the Dcr-2–Loqs-PD complex while it is processing a double-stranded RNA (dsRNA) substrate elucidate the interactions between Dcr-2 and Loqs-PD, and show that Dcr-2 undergoes substantial conformational changes during a dsRNA-processing cycle.
- Shichen Su
- , Jia Wang
- & Jinbiao Ma
-
Article
| Open Access Structure of the Dicer-2–R2D2 heterodimer bound to a small RNA duplex
Structure of the Dicer-2–R2D2 heterodimer bound to a small RNA duplex
Cryo-electron microscopy structures of Drosophila Dicer-2–R2D2 complexes with and without small interfering RNA reveal how the RNA is presented to Argonaute in the correct orientation for viral gene silencing.
- Sonomi Yamaguchi
- , Masahiro Naganuma
- & Osamu Nureki
-
Article |
 eIF5B and eIF1A reorient initiator tRNA to allow ribosomal subunit joining
eIF5B and eIF1A reorient initiator tRNA to allow ribosomal subunit joining
Single-molecule spectroscopy and structural studies were used to examine the dynamics of association of eIF1A and eIF5B with the human translation initiation complex and their role in presenting tRNA to the complex to initiate translation.
- Christopher P. Lapointe
- , Rosslyn Grosely
- & Joseph D. Puglisi
-
Article
| Open Access Mechanism of mitoribosomal small subunit biogenesis and preinitiation
Mechanism of mitoribosomal small subunit biogenesis and preinitiation
Structural analysis of several small mitoribosomal subunit intermediates reveals a sequential mechanism of biogenesis, and how assembly links to initiation to form active mitoribosomes.
- Yuzuru Itoh
- , Anas Khawaja
- & Alexey Amunts
-
Article
| Open Access A prebiotically plausible scenario of an RNA–peptide world
A prebiotically plausible scenario of an RNA–peptide world
Peptide synthesis can take place directly on RNA, which suggests how a nucleic acid–protein world might have originated on early Earth.
- Felix Müller
- , Luis Escobar
- & Thomas Carell
-
Article |
 Selective inhibition of miRNA processing by a herpesvirus-encoded miRNA
Selective inhibition of miRNA processing by a herpesvirus-encoded miRNA
Herpesvirus microRNAs interfere directly with host cell microRNA processing, thereby disrupting mitochondrial architecture, evading intrinsic host defences and driving the switch from latent to lytic infection.
- Thomas Hennig
- , Archana B. Prusty
- & Bhupesh K. Prusty
-
Matters Arising |
 Evaluating ribosomal frameshifting in CCR5 mRNA decoding
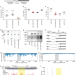
Evaluating ribosomal frameshifting in CCR5 mRNA decoding
- Yousuf A. Khan
- , Gary Loughran
- & John F. Atkins
-
Article |
 Structure of active human telomerase with telomere shelterin protein TPP1
Structure of active human telomerase with telomere shelterin protein TPP1
Cryo-electron microscopy structures of human telomerase and telomerase in complex with TPP1 provide insights into the interactions of these proteins and their activities.
- Baocheng Liu
- , Yao He
- & Juli Feigon
-
Article |
 Target site selection and remodelling by type V CRISPR-transposon systems
Target site selection and remodelling by type V CRISPR-transposon systems
Structural studies on Scytonema hofmanni CRISPR-associated transposon protein complexes indicate a mechanism for RNA-guided DNA transposition involving Cas12k, TnsC and TnsB.
- Irma Querques
- , Michael Schmitz
- & Martin Jinek
-
Article |
 Cryo-EM structures of full-length Tetrahymena ribozyme at 3.1 Å resolution
Cryo-EM structures of full-length Tetrahymena ribozyme at 3.1 Å resolution
Cryo-electron microscopy has been used to determine the structure of the Tetrahymena ribozyme (a catalytic RNA) at sufficiently high resolution to model side chains and metal ions.
- Zhaoming Su
- , Kaiming Zhang
- & Wah Chiu
-
Article |
 NORAD-induced Pumilio phase separation is required for genome stability
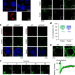
NORAD-induced Pumilio phase separation is required for genome stability
The noncoding RNA NORAD maintains genome stability in mammalian cells by sequestering Pumilio proteins in phase-separated compartments.
- Mahmoud M. Elguindy
- & Joshua T. Mendell
-
Article |
 Structures of telomerase at several steps of telomere repeat synthesis
Structures of telomerase at several steps of telomere repeat synthesis
Cryo-electron microscopy structures of Tetrahymena telomerase with telomeric DNA at several steps of nucleotide addition provide insights into the structural basis of telomere repeat synthesis.
- Yao He
- , Yaqiang Wang
- & Juli Feigon
-
Article |
 Structure of human telomerase holoenzyme with bound telomeric DNA
Structure of human telomerase holoenzyme with bound telomeric DNA
A high-resolution structure of human telomerase bound to telomeric DNA reveals details of telomerase assembly and its active site, and sheds light on how mutations alter telomerase function.
- George E. Ghanim
- , Adam J. Fountain
- & Thi Hoang Duong Nguyen
-
Article |
 The RNA m6A reader YTHDC1 silences retrotransposons and guards ES cell identity
The RNA m6A reader YTHDC1 silences retrotransposons and guards ES cell identity
N6-methyladenosine RNA and its reader YTHDC1 serve as a bridge to silencing retrotransposons through the RNA derived from these retrotransposons in mouse ES cells.
- Jiadong Liu
- , Mingwei Gao
- & Jiekai Chen
-
Article |
 The kinetic landscape of an RNA-binding protein in cells
The kinetic landscape of an RNA-binding protein in cells
Time-resolved RNA–protein cross-linking with a pulsed femtosecond ultraviolet laser, followed by immunoprecipitation and high-throughput sequencing, enables the determination of binding and dissociation kinetics of the RNA-binding protein DAZL within cells.
- Deepak Sharma
- , Leah L. Zagore
- & Eckhard Jankowsky
-
Article |
 BRCA1 and RNAi factors promote repair mediated by small RNAs and PALB2–RAD52
BRCA1 and RNAi factors promote repair mediated by small RNAs and PALB2–RAD52
Single-stranded, DNA-damage-associated small RNAs generated by a BRCA1–RNA-interference complex promote PALB2–RAD52-mediated DNA repair at transcriptional termination pause sites that contain R-loops and are rich in single-stranded DNA breaks in both quiescent and proliferating cells.
- Elodie Hatchi
- , Liana Goehring
- & David M. Livingston
-
Article |
 RNA nucleation by MSL2 induces selective X chromosome compartmentalization
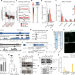
RNA nucleation by MSL2 induces selective X chromosome compartmentalization
Dosage compensation in Drosophila involves nucleation of the dosage compensation complex at the X chromosome by MSL2 and the non-coding RNA roX.
- Claudia Isabelle Keller Valsecchi
- , M. Felicia Basilicata
- & Asifa Akhtar
-
Article |
 Structural basis for the final steps of human 40S ribosome maturation
Structural basis for the final steps of human 40S ribosome maturation
Studies of five cryo-electron microscopy structures reveal the composition and conformational progression in the final maturation events of human 40S ribosomal subunit assembly.
- Michael Ameismeier
- , Ivo Zemp
- & Roland Beckmann
-
Article |
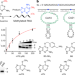 Site-specific RNA methylation by a methyltransferase ribozyme
Site-specific RNA methylation by a methyltransferase ribozyme
A methyltransferase ribozyme, along with the small-molecule cofactor O6-methylguanine, is shown to catalyse the site-specific installation of 1-methyladenosine in various RNAs, providing insights into the catalytic abilities of RNA.
- Carolin P. M. Scheitl
- , Mohammad Ghaem Maghami
- & Claudia Höbartner
-
Article |
 RAD51-dependent recruitment of TERRA lncRNA to telomeres through R-loops
RAD51-dependent recruitment of TERRA lncRNA to telomeres through R-loops
Telomeric-repeat-containing RNA is recruited to telomeres by a mechanism that involves the DNA recombinase RAD51 and the formation of DNA–RNA hybrids, or R-loops—a process similar to that involved in homology-directed DNA repair.
- Marianna Feretzaki
- , Michaela Pospisilova
- & Joachim Lingner
-
Article |
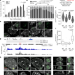 A protein assembly mediates Xist localization and gene silencing
A protein assembly mediates Xist localization and gene silencing
A protein condensate formed by multivalent interactions between the long non-coding RNA Xist and specific RNA-binding proteins drives the compartmentalization required to perpetuate gene silencing on the inactive X chromosome.
- Amy Pandya-Jones
- , Yolanda Markaki
- & Kathrin Plath
-
Article |
 Cryo-EM of elongating ribosome with EF-Tu•GTP elucidates tRNA proofreading
Cryo-EM of elongating ribosome with EF-Tu•GTP elucidates tRNA proofreading
Time-resolved cryogenic electron microscopy structures of a ribosome during the delivery of aminoacyl-tRNA by EF-Tu•GTP capture 33 ribosomal states, enabling visualization of the initial selection, proofreading and peptidyl transfer stages.
- Anna B. Loveland
- , Gabriel Demo
- & Andrei A. Korostelev
-
Article |
 Base-pair conformational switch modulates miR-34a targeting of Sirt1 mRNA
Base-pair conformational switch modulates miR-34a targeting of Sirt1 mRNA
Repression of a messenger RNA by a cognate microRNA depends not only on complementary base pairing, but also on the rearrangement of a single base pair, producing a conformation that fits better within the human Ago2 protein.
- Lorenzo Baronti
- , Ileana Guzzetti
- & Katja Petzold
-
Article |
 Structure of replicating SARS-CoV-2 polymerase
Structure of replicating SARS-CoV-2 polymerase
A cryo-electron microscopy structure of the RNA-dependent RNA polymerase of SARS-CoV-2 sheds light on coronavirus replication and enables the analysis of the inhibitory mechanisms of candidate antiviral drugs.
- Hauke S. Hillen
- , Goran Kokic
- & Patrick Cramer
-
Article |
 Determination of RNA structural diversity and its role in HIV-1 RNA splicing
Determination of RNA structural diversity and its role in HIV-1 RNA splicing
Dimethyl sulfate mutational profiling with sequencing, combined with the newly developed DREEM algorithm, reveals that heterogeneity of RNA structure in HIV-1 regulates the use of splice sites and expression of viral genes.
- Phillip J. Tomezsko
- , Vincent D. A. Corbin
- & Silvi Rouskin
