Abstract
This protocol is an extension to: Nat. Protoc. 9, 2395–2410 (2014); doi:10.1038/nprot.2014.157; published online 18 September 2014
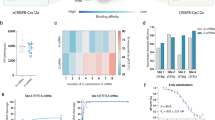
In recent years, CRISPR/Cas9 has emerged as a powerful tool for improving crop traits. Conventional plant genome editing mainly relies on plasmid-carrying cassettes delivered by Agrobacterium or particle bombardment. Here, we describe DNA-free editing of bread wheat by delivering in vitro transcripts (IVTs) or ribonucleoprotein complexes (RNPs) of CRISPR/Cas9 by particle bombardment. This protocol serves as an extension of our previously published protocol on genome editing in bread wheat using CRISPR/Cas9 plasmids delivered by particle bombardment. The methods we describe not only eliminate random integration of CRISPR/Cas9 into genomic DNA, but also reduce off-target effects. In this protocol extension article, we present detailed protocols for preparation of IVTs and RNPs; validation by PCR/restriction enzyme (RE) and next-generation sequencing; delivery by biolistics; and recovery of mutants and identification of mutants by pooling methods and Sanger sequencing. To use these protocols, researchers should have basic skills and experience in molecular biology and biolistic transformation. By using these protocols, plants edited without the use of any foreign DNA can be generated and identified within 9–11 weeks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Uauy, C. Wheat genomics comes of age. Curr. Opin. Plant Biol. 36, 142–148 (2017).
Weeks, D.P., Spalding, M.H. & Yang, B. Use of designer nucleases for targeted gene and genome editing in plants. Plant Biotechnol. J. 14, 483–495 (2016).
Voytas, D.F. Plant genome engineering with sequence-specific nucleases. Annu. Rev. Plant Biol. 64, 327–350 (2013).
Chen, K. & Gao, C. Targeted genome modification technologies and their applications in crop improvements. Plant Cell Rep. 33, 575–583 (2014).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Wang, H., La Russa, M. & Qi, L.S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Barrangou, R. & Doudna, J.A. Applications of CRISPR technologies in research and beyond. Nat. Biotechnol. 34, 933–941 (2016).
Sander, J.D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Wright, A.V., Nunez, J.K. & Doudna, J. Biology and applications of CRISPR systems: harnessing nature's toolbox for genome engineering. Cell 164, 29–44 (2016).
Wang, Y. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Xie, K., Minkenberg, B. & Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl. Acad. Sci. USA 112, 3570–3575 (2015).
Liang, Z., Zhang, K., Chen, K. & Gao, C. Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. J. Genet. Genomics 41, 63–68 (2014).
Svitashev, S. et al. Targeted mutagenesis, precise gene editing, and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol. 169, 931–945 (2015).
Char, S.N. et al. An Agrobacterium-delivered CRISPR/Cas9 system for high-frequency targeted mutagenesis in maize. Plant Biotechnol. J. 15, 257–268 (2017).
Li, Z. et al. Cas9-guide RNA directed genome editing in soybean. Plant Physiol. 169, 960–970 (2015).
Jacobs, T.B., LaFayette, P.R., Schmitz, R.J. & Parrott, W. A. Targeted genome modifications in soybean with CRISPR/Cas9. BMC Biotechnol. 15, 16 (2015).
Butler, N.M., Atkins, P.A., Voytas, D.F. & Douches, D. Generation and inheritance of targeted mutations in potato (Solanum tuberosum L.) using the CRISPR/Cas system. PLoS ONE 10, e0144591 (2015).
Brooks, C., Nekrasov, V., Lippman, Z.B. & Van Eck, J. Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol. 166, 1292–1297 (2014).
Zhang, D., Li, Z. & Li, J. Targeted gene manipulation in plants using the CRISPR/Cas technology. J. Genet. Genomics 43, 251–262 (2016).
Ding, Y., Li, H., Chen, L.L. & Xie, K. Recent advances in genome editing using CRISPR/Cas9. Front. Plant Sci. 7, 703 (2016).
Belhaj, K., Chaparro-Garcia, A., Kamoun, S., Patron, N.J. & Nekrasov, V. Editing plant genomes with CRISPR/Cas9. Curr. Opin. Biotechnol. 32, 76–84 (2015).
Kouranova, E. et al. CRISPRs for optimal targeting: delivery of CRISPR components as DNA, RNA, and protein into cultured cells and single-cell embryos. Hum. Gene Ther. 27, 464–475 (2016).
Bortesi, L. & Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 33, 41–52 (2015).
Gorbunova, V. & Levy, A. Non-homologous DNA end joining in plant cells is associated with deletions and filler DNA insertions. Nucleic Acids Res. 25, 4650–4657 (1997).
Kim, J. & Kim, J.S. Bypassing GMO regulations with CRISPR gene editing. Nat. Biotechnol. 34, 1014–1015 (2016).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Staahl, B.T. et al. Efficient genome editing in the mouse brain by local delivery of engineered Cas9 ribonucleoprotein complexes. Nat. Biotechnol. 35, 431–434 (2017).
Cho, S.W., Lee, J., Carroll, D., Kim, J.S. & Lee, J. Heritable gene knockout in Caenorhabditis elegans by direct injection of Cas9-sgRNA ribonucleoproteins. Genetics 195, 1177–1180 (2013).
Burger, A. et al. Maximizing mutagenesis with solubilized CRISPR-Cas9 ribonucleoprotein complexes. Development 143, 2025–2037 (2016).
DeWitt, M.A. et al. Selection-free genome editing of the sickle mutation in human adult hematopoietic stem/progenitor cells. Sci. Transl. Med. 8, 360ra134 (2016).
Hultquist, J.F. et al. A Cas9 ribonucleoprotein platform for functional genetic studies of HIV-host interactions in primary human T cells. Cell Rep. 17, 1438–1452 (2016).
Kim, S., Kim, D., Cho, S.W., Kim, J. & Kim, J.S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Schumann, K. et al. Generation of knock-in primary human T cells using Cas9 ribonucleoproteins. Proc. Natl. Acad. Sci. USA 112, 10437–10442 (2015).
Woo, J.W. et al. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 33, 1162–1164 (2015).
Subburaj, S. et al. Site-directed mutagenesis in Petunia x hybrida protoplast system using direct delivery of purified recombinant Cas9 ribonucleoproteins. Plant Cell Rep. 35, 1535–1544 (2016).
Malnoy, M. et al. DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front. Plant Sci. 7, 1904 (2016).
Shan, Q., Wang, Y., Li, J. & Gao, C. Genome editing in rice and wheat using the CRISPR/Cas system. Nat. Protoc. 9, 2395–2410 (2014).
Zhang, Y. et al. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016).
Liang, Z. et al. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 8, 14261 (2017).
Li, J. et al. Registration of a high-yielding and broad-adaptive winter wheat cultivar-Kenong 199. J. Triticeae Crops 27, 368 (2007).
Svitashev, S., Schwartz, C., Lenderts, B., Young, J.K. & Mark Cigan, A. Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes. Nat. Commun. 7, 13274 (2016).
Xiao, A. et al. CasOT: a genome-wide Cas9/gRNA off-target searching tool. Bioinformatics 30, 1180–1182 (2014).
Zuris, J.A. et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat. Biotechnol. 33, 73–80 (2015).
Liu, J. et al. Efficient delivery of nuclease proteins for genome editing in human stem cells and primary cells. Nat. Protoc. 10, 1842–1859 (2015).
Zhang, H. et al. The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol. J. 12, 797–807 (2014).
Kruger, N.J. The Bradford method for protein quantitation. Methods in Molecular Biology Vol. 32 (ed. J.M. Walker) 9–15 (Humana Press, 1994).
Allen, G.C., Flores-Vergara, M.A., Krasynanski, S., Kumar, S. & Thompson, W.F. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat. Protoc. 1, 2320–2325 (2006).
Acknowledgements
This work was supported by grants from the National Key Research and Development Program of China (2016YFD0101804), the National Transgenic Science and Technology Program (2016ZX08010-002), the Chinese Academy of Sciences (QYZDY-SSW-SMC030 and GJHZ1602) and the National Natural Science Foundation of China (31788103, 31420103912 and 31570369).
Author information
Authors and Affiliations
Contributions
Z.L., K.C. and C.G. designed the experiments for the protocol; Z.L. and Y.Z. performed the experiments with assistance from K.C., J.L., K.Y. and J.-L.Q.; and Z.L., K.C. and C.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Notes 1–3. (PDF 449 kb)
Rights and permissions
About this article
Cite this article
Liang, Z., Chen, K., Zhang, Y. et al. Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins. Nat Protoc 13, 413–430 (2018). https://doi.org/10.1038/nprot.2017.145
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.145
This article is cited by
-
Optimized protocols for protoplast isolation, transfection, and regeneration in the Solanum genus for the CRISPR/Cas-mediated transgene-free genome editing
Applied Biological Chemistry (2024)
-
Targeted genome-modification tools and their advanced applications in crop breeding
Nature Reviews Genetics (2024)
-
Fine mapping of a major QTL, qKl-1BL controlling kernel length in common wheat
Theoretical and Applied Genetics (2024)
-
Optimized prime editing in monocot plants using PlantPegDesigner and engineered plant prime editors (ePPEs)
Nature Protocols (2023)
-
The sonication-assisted whisker method enables CRISPR-Cas9 ribonucleoprotein delivery to induce genome editing in rice
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.