Abstract
Abnormal Transmembrane protein 9 (TMEM9) expression has been identified in various human tumors. However, the prognostic potential and mechanistic role of TMEM9 in lung adenocarcinoma (LUAD) remain unclear. Here, we first found a significant upregulation of TMEM9 in LUAD tissues, and TMEM9 expression was positively correlated with microvessel density (MVD), T stage, and clinical stage. Survival analysis demonstrated TMEM9 was an independent indicator of poor prognosis in LUAD patients. In addition, downregulation of TMEM9 suppressed tumor growth and metastasis in vitro and in vivo models, and reduced HUVEC proliferation, migration, and tube formation in a cancer cell/HUVEC coculture model. Furthermore, TMEM9 upregulated VEGF expression, and VEGF-neutralizing antibodies reversed HUVEC angiogenesis and cancer cell migration ability caused by overexpression of TMEM9. In contrast, recombinant VEGF (rVEGF) abolished the inhibitory effect of TMEM9-knockdown LUAD cells on HUVEC angiogenesis and tumor cell migration. Moreover, we showed that TMEM9 upregulated VEGF expression by activating the mitogen-activated protein kinase/extracellular signal-regulated kinase/STAT3 (MEK/ERK/STAT3) pathway. Together, our study provides mechanistic insights into the role of TMEM9 in LUAD and highlights the potential of targeting the TMEM9/MEK/ERK/STAT3/VEGF pathway as a novel therapy for preventing LUAD progression.
Similar content being viewed by others
Introduction
Lung cancer is one of the most common cancers globally, with high morbidity and mortality [1]. Non-small cell lung cancer (NSCLC) accounts for 80%–85% of histological types of lung cancers [2]. The most common type of NSCLC is lung adenocarcinoma (LUAD) [3]. Despite significant advances in diagnosing and treating LUAD in recent years, the prognosis of affected patients remains gloomy, particularly in those with recurrent or metastatic LUAD with a poor long-term prognosis [4]. Therefore, it is of great significance to explore the molecular mechanisms leading to an increased metastatic phenotype.
Transmembrane protein 9 (TMEM9), a type I transmembrane protein situated in late endosomes and lysosomes, spans 183 amino acids and features a signal peptide and three glycosylation sites at the amino terminus [5]. TMEM9 has been implicated in various cellular mechanisms, including inflammation [6], tissue regeneration [7], cell differentiation, and tumorigenesis [8]. Many studies have demonstrated that TMEM9 is highly expressed in multiple human cancers, such as mammary tumors and hepatocellular and colorectal cancer [9]. It is critical in cancer cell proliferation, apoptosis, invasion, metastasis [10], and drug resistance [11]. Therefore, a major function for TMEM9 in regulating tumorigenesis and development has been suggested. However, little is known about the pathological role of TMEM9 in LUAD. In addition, the molecular mechanisms of TMEM9-elicited tumor development remain unclear.
In the present study, we evaluated TMEM9 expression and its prognostic value in LUAD. We also provided a novel insight into a mechanism by which TMEM9 mediated tumor growth via MEK/ERK/STAT3/VEGF pathway in LUAD. These findings may represent a novel strategy for therapeutic interventions for LUAD in the future.
Materials and methods
Cell culture and transfection
Human umbilical vein endothelial cells (HUVECs), LUAD cell lines A549 and Anip973, and Human lung epithelial cell line BEAS-2B were donated from the Central Laboratory of Affiliated Central Hospital of Qingdao University. A549 was cultured in the RPMI 1640 medium (GIBCO), and Anip973 and BEAS-2B were cultured in the DMEM medium (GIBCO). HUVECs were cultured in HUVEC media (Haixing Biosciences, Jiangsu, China, TCH-G406). All cells were cultured in a 37 °C, 5% CO2 incubator supplemented with 10% fetal bovine serum, 100 U/mL of penicillin, and 100 mg/mL of streptomycin.
A549 and Anip973 cells were transfected with short hairpin RNAs (shRNA) targeting TMEM9 using lentivirus vector GV493 (Genechem, Shanghai, China). The shRNA sequences were as follows: shTMEM9#1, GCATCTGTCCACCTTATAGAA, shTMEM9#2, GACAGTCTTCGATCGGCACAA. The TMEM9 over-expressing cell lines were generated by lentivirus vector GV703 containing the full-length sequence of TMEM9 (Genechem, Shanghai, China). For the rescue experiments, an RNAi-resistant TMEM9-expression plasmid (oe-TMEM9) or its empty vector (oe-NC) was purchased from Youbio Biological Technology (Hunan, China).
Clinical specimens
This study collected 43 pairs of fresh LUAD tissues and adjacent normal tissues (ANT) from the Affiliated Hospital of Qingdao University from April to October 2021. Clinicopathological information for these patients is shown in Supplementary Table S1. This study was approved by the Ethics Committee Medical College of Qingdao University (approval number: QDU-HEC-2021145), and informed consent was obtained from each patient. Three LUAD cancer tissue microarrays were purchased from Shanghai Outdo Biotech (Shanghai, China). One tissue microarray (HLugA180Su11) contained 90 pairs of carcinoma tissues and matched para-carcinoma tissues. The other two identical sets of tissue microarray chips (HLugA120PG01) each had 120 LUAD tissue dots, of which 103 could be used as samples for further analysis. The clinical characteristics of these patients are provided in Supplementary Tables S2 and S3.
Immunohistochemistry (IHC)
Immunohistochemistry was conducted according to the previous protocols [12]. Briefly, the tissue array specimen was first dewaxed, repaired with EDTA, and incubated with primary antibody (anti-VEGF, ab1316, 1:200; anti-CD31, ab182981, 1:2000 from Abcam; anti-TMEM9, A61446, 1:2000 from EpigenTek) overnight at 4 °C. Next, a secondary antibody was added and incubated at room temperature for 60 min. After diluted diaminobenzidine (DAB) was added to pathological sections, hematoxylin (SIGMA) was used for inhibition. Finally, the slices were sealed after dehydration.
The specimens were scored by 3 independent pathologists who did not know any prognosis or clinicopathologic variables. The staining intensity was scored as previously described: 0, no staining; 1, faint cytoplasmic staining; 2, moderate cytoplasmic staining; and 3, strong cytoplasmic staining, and the percentage of stained cancer cells was recorded. The expression level of TMEM9 and VEGF was evaluated by the percentage of positive cells and the staining intensity score. Based on CD31 staining, the microvascular density (MVD) of LUAD tumor sections was assessed according to the previous literature [13].
In vivo tumorigenesis and metastasis assays
Purchased 4 week-old BALB/c female nude mice were randomly divided into different groups after adaptive feeding for 1 week. A549 cells stably expressing shNC or shTMEM9 lentivirus vectors were injected subcutaneously into the left flank of 5-week-old mice (2 × 107 mL−1, 0.1 mL per mice, n= 5 in each group). The tumor volume was measured every 4 days and calculated as follows: volume = (width2 × length) / 2. 28 days later, the tumor tissue was fixed in 10% formalin solution for 24 h, embedded in paraffin. IHC staining was performed.
To verify the effect of TMEM9 on tumor metastasis in vivo, A549 cells stably expressing shNC or shTMEM9 lentivirus vectors were injected into the tail vein of nude mice (1 × 107 mL−1, 0.15 mL per mice, n = 5 in each group). After 6 weeks, lung tissues were dissected, and metastatic nodules were counted. Then it was fixed in 4% paraformaldehyde and analyzed by H&E staining after paraffin embedding. The animal study was approved by the Committee on Qingdao University (approval number: QDU-AEC-2021184).
Enzyme-linked immunosorbent assay (ELISA)
The cell culture supernatant was collected, and the VEGF concentration was determined according to the ELISA kit instructions (Liankebio, Hangzhou, China).
RNA isolation and quantitative real-time PCR analysis
Total RNA from LUAD cells and tumor tissues was extracted using TRIzol Reagent (Vazyme, Nanjing, China), and cDNA was synthesized by reverse transcription. Finally, quantitative detection was performed with ChamQ Universal SYBR qPCR Master Mix (Vazyme Biotech, Nanjing, China). Relative gene expression was determined by normalizing the expression of each target gene to β-actin. The data were analyzed by using 2△△Ct.
Primers were shown as follows: TMEM9, Forward (F): 5′-GGGCACATTTACAACCAG-3′, Reverse (R): 5′-ATCAGGAAGGCCATG-TAG-3′; VEGF, Forward (F): 5′- TTCTGGGCTGTTCTCGCTTC-3′, Reverse (R): 5′- CTCTCCTCTTCCTTCTCTTCTTCC -3′.
Immunofluorescence
A549 and Anip973 cells were seeded on glass coverslips in 24-well plates. When the cells grew to 80% confluence, they were fixed with 4% paraformaldehyde and permeabilized with 0.1% Triton X-100 in PBS. Fixed cells were incubated with primary TMEM9 antibody (HPA008483, 1:500 from ATLAS ANTIBODIES) overnight at 4 °C and followed by incubation with FITC-conjugated anti-rabbit IgG for 1 h. DAPI was used for nuclear counterstaining. And cells were observed under a fluorescence microscope.
CCK8 assay
Cells were seeded in 96-well cell culture plates at 3000 cells per well, and three duplicate wells were set in each group. After 24–72 h of culture, CCK8 (USA, MedChem Express, HY-K0301) reagent was added and cultured for 4 h. The absorbance (OD) at 450 nm was detected with a microplate reader. All experiments were repeated at least 3 times.
EdU assay
HUVEC proliferation was measured using an EdU assay kit (Beyotime Biotechnology, Shanghai, China). HUVECs were seeded at the appropriate density in a 24-well plate. Subsequently, Transwell chambers containing the transfected LUAD cell were inserted into the 24-well plates and cocultured at 37 °C, 5% CO2 for 48 h. EdU staining was performed following the manufacturer’s instructions. Finally, fluorescence images were obtained using a fluorescence microscope.
Protein extraction and western blotting
LUAD cells were lysed in RIPA buffer containing protease inhibitors and phosphatase inhibitors. Cell lysates were electrophoresed in 12% SDS-PAGE and then transferred onto a nitrocellulose membrane (MerrckMillipore, Ireland). Membranes were blocked with TBST (Tris-buffered saline, 0.1% Tween-20) containing 5% skim milk for 2 h and were incubated with the indicated antibodies, including anti-MEK1/2 (Abcam, CAT# ab178876);anti-Phospho-MEK1/2 (CST, CAT# 9154 T);anti-ERK1/2 (CST, CAT#4695 T); anti-Phospho-ERK1/2(CST, CAT#4370 T); anti-AKT (CST, CAT#4691 T); anti-Phospho-AKT (CST, CAT#4060 T);anti-STAT3 (Huaan Biotech, ET1607-38); anti-Phospho-STAT3 (CST, CAT#9145 T) overnight at 4 °C. Then, the membranes were washed with TBST and incubated with a peroxidase-conjugated second antibody for 1 h. The protein bands were detected by enhanced chemiluminescence.
Chromatin immunoprecipitation (ChIP)
The cells were used to carry out ChIP assays with the SimpleChIP® Enzymatic Chromatin IP Kit (Agarose Beads) (CST, CAT#9002) and an anti-Phospho-STAT3(CST, CAT#9145 T) following the provided protocol. The enriched DNA was measured using qRT-PCR with the primers 5′-AAGGCCAGGGGTCACTCCAG-3′ (Forward) and 5′-CCCGCGGGGCATTGGCGAGG-3′ (Reverse) to detect VEGF promoter.
Wound Healing
LUAD cells or HUVECs were seeded in 6-well plates in a CO2 incubator at 37 °C. When the cells reached 90–100% confluence, a scratch was made in the monolayer using a sterile p200 tip, and cell debris was washed with PBS. The fresh culture medium containing 2% serum was added to the cells. The cells were cultured for 24–72 h. Images of the scratch were taken at 0 h,24 h, and 72 h using a phase contrast inverted microscope. Then wound healing rates were calculated.
Transwell coculture system
For transwell chamber coculture, HUVECs were added into each well of a six-plate well, and the transfected LUAD cells were seeded into the upper compartment. In some cases, 150 ng/ml Bevacizumab (Roche Pharmaceuticals, Basel, Switzerland) or 2 ng/ml recombinant human VEGF165 (Peprotech, Suzhou, China) was added into the conditioned medium. These two cells were cocultured for 48 h, and HUVECs were collected for further study.
Tube formation
HUVECs (6 × 105 cells/ml) were seeded into 96-well plates containing matrix (50uL matrix per well, placed in 37 °C, 5% CO2 incubator for 30 min and then used), 50uL cell suspension was added into each well, and 3 repeat wells were set up. The cells were cultured in 37 °C, 5% CO2 incubator for 6 h and then photographed under the microscope. The images were evaluated using the Image J software.
Bioinformatics
The expression level of TMEM9 in tumor tissues and normal tissues was obtained from the Oncogene Atlas (TCGA) and Genotype-Tissue Expression (GTEx). Kaplan-Meier plotter (www.kmplot.com) [14] was used to examine correlations between TMEM9 expression and survival. The R software package ggstatsplot realized the correlation chart between TMEM9 and VEGF. RNA-sequencing expression (level 3) profiles for LUAD were downloaded from the TCGA Dataset (https://portal.gdc.com).
Statistical analysis
All data were obtained from 3 independent experiments. The t-test was used to compare the groups. A Cox regression analysis was used to perform univariate and multivariate survival analyses. Data were analyzed using GraphPad Prism 8.0 and IBM SPSS Statistics 25.0 software. NS, no significant difference, *P < 0.05, **P < 0.01.
Results
High TMEM9 mRNA expression is correlated with poor prognosis in LUAD patients
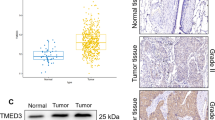
To investigate the differences in TMEM9 expression between tumors and normal tissues, we analyzed the expression levels of TMEM9 in different tumors and their corresponding normal tissues of pan‐cancer using transcriptomic data from TCGA and GTEx databases. The pan-cancer analysis demonstrated that TMEM9 was highly expressed in most tumor tissues (Fig. 1A). We also analyzed TMEM9 mRNA expression in LUAD and pair-matched adjacent normal tissues using the TCGA database and found that TMEM9 expression was substantially higher in LUAD relative to adjacent normal lung tissues (P = 2.2e-05, Supplementary Fig. S1A). We further detected TMEM9 mRNA expression in 43 LUAD tissues and paired normal lung tissues. The result showed that TMEM9 was highly expressed in the tumor tissues (P < 0.01, Fig. 1B). Subsequently, we evaluated the correlation between TMEM9 mRNA expression and clinicopathologic parameters in LUAD patients. As shown in Fig. 1C, D, TMEM9 expression was significantly correlated with the T stage (P = 0.029) and spread through air spaces (STAS) (P = 0.001) of LUAD (Table S1). The Kaplan-Meier plotter database was then used to evaluate whether TMEM9 mRNA expression was correlated with the prognosis of LUAD patients. The Data showed that patients with low TMEM9 expression had longer overall survival (OS) compared to patients with high TMEM9 expression (P = 5.8e-06, Fig. 1E). Therefore, these results suggest that elevated TMEM9 mRNA expression is a risk factor for LUAD and may be a potential prognostic biomarker for patients with LUAD.
A TMEM9 mRNA expression levels in pan‐cancer from TCGA + GTEx database. B The mRNA level of TMEM9 in LUAD and para-carcinoma tissues was measured by qPCR. C TMEM9 mRNA levels in tumor tissues of patients with different T stages. D TMEM9 mRNA levels in tumor tissues of LUAD patients in the STAS negative and STAS positive groups. E Kaplan-Meier curves for LUAD patients’ OS depend on TMEM9 mRNA levels in the TCGA database. Ns, no significant difference, *P < 0.05, **P < 0.01.
TMEM9 was an independent indicator of poor prognosis of LUAD patients
The immunohistochemistry for TMEM9 was performed in the LUAD specimens chip. The staining intensity was higher in LUAD compared with peritumor tissues (P < 0.01, Fig. 2A, B). As shown in Fig. 2C, D, high TMEM9 expression was significantly associated with high microvessel density (MVD) in LUAD tissues (P < 0.01). In addition, the expression of TMEM9 increased gradually with the progress of the clinical stage, T stage, and grade (Fig. 2E and Supplementary Fig. S1B, C). Kaplan-Meier curves showed that LUAD patients with high TMEM9 expression had shorter overall survival (P < 0.01, Fig. 2E). Furthermore, univariate and multivariate Cox regression analysis showed that TMEM9 expression was an independent prognostic factor for overall survival of LUAD patients (Table 1). Together, these results suggest that increased TMEM9 expression is a poor prognostic factor for LUAD.
A Representative images of immunohistochemical staining for TMEM9 expression in LUAD tissues and adjacent normal tissues (ANT). Scale bar, 100 μm. B The percentages of high-TMEM9 cases in LUAD and ANT. C TMEM9 expression levels were positively correlated with MVD levels in tissue samples from patients with LUAD (n = 103). D The expression of MVD in patients with high TMEM9 expression was higher than those in patients with low TMEM9 expression (n = 103). Scale bar, 100 μm. E The associations between TMEM9 expression and the clinicopathological parameters of LUAD patients were analyzed. F Kaplan-Meier curves of TMEM9 expression in LUAD (n = 90). Data are presented as the mean ± SD. Ns, no significant difference, *P < 0.05, **P < 0.01.
TMEM9 promotes LUAD cell growth, migration, and angiogenesis
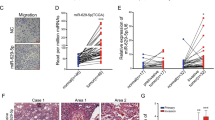
TMEM9 expression was examined in five different LUAD cell lines as well as normal lung epithelial cells BEAS-2B. The expression level of TMEM9 in LUAD cell lines was significantly increased compared with normal lung epithelial cells (Fig. 3A). Immunofluorescent staining showed that TMEM9 predominantly localized to the cytoplasm and the perinucleus with a speckled pattern in A549 and Anip973 cells (Fig. 3B). Next, we determined the biological function of TMEM9 in LUAD cells. TMEM9 was knocked down in A549 and Anip973 (Fig. 3C, D). As shown in Fig. 3E, F, CCK8 and wound-healing assay indicated that down-regulation of TMEM9 inhibited the proliferation and migration of LUAD cells.
A The mRNA levels of TMEM9 in LUAD and normal cells were measured by qPCR. B Immunofluorescent staining showed subcellular localization of TMEM9 in A549 and Anip973. Scale bar, 100 μm. C QPCR analysis of TMEM9 in LUAD stable cell lines with TMEM9 knockdown (shTMEM9) or their control cell lines (sh-NC). D Western blot analysis of TMEM9 in LUAD stable cell lines with TMEM9 knockdown (shTMEM9) or their control cell lines (sh-NC). E The CCK8 assays showed that TMEM9 knockdown significantly inhibited LUAD cell proliferation. F Scratch wound healing assay showed that TMEM9 knockdown inhibited the migration of LUAD cells. G–I ShNC cells or shTMEM9 cells were cocultured with HUVECs for 48 h, and changes in angiogenesis were evaluated by EdU (G), wound-healing (H), and tube formation (I) assays. Scale bar, 100 μm. Results are shown for three experiments performed in triplicate.Data are presented as the mean ± SD. Ns, no significant difference, *P < 0.05, **P < 0.01.
To explore the effects of TMEM9 on the Matrigel capillary structure formation of HUVECs, an indirect coculture model was established based on a transwell chamber. TMEM9 knockdown LUAD cells were cocultured with HUVECs, and then the proliferation, migration, and tube formation of HUVECs were determined. These results showed that TMEM9 interfering in LUAD cells significantly decreased the proliferation, migration, and tube formation of HUVECs compared to scramble shRNA interfering (Fig. 3G–I).
Next, Rescue experiments were performed to further eliminate the possibility that the decrease in proliferation, migration, and angiogenesis was due to shRNA‐mediated off-target effects. ShTMEM9#2 was chosen for the following experiment due to its higher interference efficiency. TMEM9 stable knockdown cells were transfected with a vector expressing TMEM9 (Fig. 4A). As shown in Fig. 4B–F, the rescue of TMEM9 expression could partly reverse the inhibitory effects of TMEM9 knockdown on cell growth, migration, and angiogenesis. In addition, we established LUAD cells stably overexpressing TMEM9 (Supplementary Fig. S2A). TMEM9 overexpression promotes LUAD cell growth, migration, and angiogenesis (Supplementary Fig. S2B–F). These results indicate that TMEM9 may serve an important role in LUAD progression.
A TMEM9 stable knockdown LUAD cells were transfected with a vector expressing TMEM9.The expression of TMEM9 in A549 and Anip973 cells after transfection was detected by Westen blot. B The CCK8 assays showed that upregulated TMEM9 expression could reverse the inhibitory effects of TMEM9 knockdown on cell proliferation. C Scratch wound healing assay showed that upregulated TMEM9 expression could reverse the inhibitory effects of TMEM9 knockdown on cell migration. D–F LUAD cells were cocultured with HUVECs for 48 h, and the effect of rescued TMEM9 expression in LUAD cells on angiogenesis were evaluated by EdU (D), wound-healing (E), and tube formation (F) assays. Scale bar, 100 μm. Results are shown for three experiments performed in triplicate.Data are presented as the mean ± SD. Ns, no significant difference, *P < 0.05, **P < 0.01.
TMEM9 increases the migration of LUAD cells and promotes angiogenesis through VEGF secretion
VEGF is one of the critical regulators responsible for angiogenesis [15]. To test the hypothesis that VEGF, as a downstream of TMEM9, might enhance the angiogenesis, we first analyzed the association of TMEM9 with VEGF using public databases. The data showed a positive correlation between VEGF expression level and TMEM9 level in LUAD (P = 0.004, Supplementary Fig. S3). We further detected the mRNA expression of TMEM9 and VEGF in LUAD tissues by qPCR. As expected, a significant correlation between TMEM9 and VEGF was observed (P = 0.012, r = 0.3795, Fig. 5A). Overexpression of TMEM9 significantly increased the expression of VEGF in A549 and Anip973 cells (Fig. 5B and Supplementary Fig. S4A). At the same time, the knockdown of TMEM9 decreased VEGF expression (Supplementary Fig. S4B, C). Subsequently, VEGF levels in the cell supernatant were detected by ELISA. The results showed that inhibition of TMEM9 significantly reduced the secretion of VEGF, while overexpression of TMEM9 upregulated VEGF (Fig. 5C).
A A significant association of TMEM9 expression with VEGF in LUAD tissue was observed (n = 43). B TMEM9 overexpressing increased VEGF expression in LUAD cells. C Culture supernatant VEGF levels were determined by ELISA. D–F The EdU assay (D), tube formation (E), and scratch surface healing (F) were examined. antiVEGF was added to the culture medium of HUVECs coculture with TMEM9 overexpressed LUAD cells or control cells. G Effect of Bevacizumab (Bev) on LUAD cell migration. Results are shown for three experiments performed in triplicate. Data are presented as the mean ± SD. Ns, no significant difference, *P < 0.05, **P < 0.01.
To verify whether TMEM9 expression in LUAD cell lines promoted tumor cell migration and angiogenesis through VEGF, VEGF neutralizing antibody (Bevacizumab) was added into the coculture system. The results showed that Bevacizumab reversed the promoting effect of TMEM9-overexpressed LUAD cells on HUVEC angiogenesis (Fig. 5D–F and Supplementary Fig. S5B). Moreover, Bevacizumab blocked the positive effect of TMEM9 on the migration of LUAD cell lines(Fig. 5G). Furthermore, when compared to control, recombinant VEGF (rVEGF) abolished the inhibitory effect of TMEM9-knockdown LUAD cells on HUVEC angiogenesis and tumor cell migration (Supplementary Fig. S5). Together, these results indicate that VEGF is a downstream molecule of TMEM9, which plays an essential role in migration and angiogenesis.
TMEM9 promotes VEGF expression via MEK/ERK/STAT3 signaling pathway in LUAD
Previous studies suggested that VEGF expression is regulated by several signaling pathways, such as the transcription factor STAT3, MEK/ERK, or AKT pathways [16,17,18,19,20]. To elucidate the mechanism of TMEM9 induced angiogenesis, the activation of STAT3, AKT, and MEK/ERK signaling was observed in LUAD cells treated with shTMEM9 LUAD cells. Our results revealed that TMEM9 knockdown reduced the phosphorylation levels of MEK1/2, ERK1/2, and STAT3 in both A549 and Anip973 cells (Fig. 6A). Conversely, TMEM9 overexpression increased the levels of phosphorylated MEK1/2, ERK1/2 and STAT3 (Fig. 6B). Next, we investigated the effect of MEK inhibitor (U0126) and STAT3 inhibitor (cryptotanshinone, CTS) on VEGF expression in LUAD cells mediated by TEME9. As shown in Fig. 6C, D, MEK inhibitor (U0126) inhibited the promoting effect of TMEM9 on STAT3 phosphorylation and VEGF expression in LUAD cells. In addition, STAT3 inhibitor (CTS) inhibited VEGF expression mediated by TMEM9. Furthermore, we determined the effect of TMEM9 on VEGF expression by p-STAT3. ChIP-qPCR showed that the knockdown of TMEM9 reduced the binding of p-STAT3 to the VEGF promoter (Fig. 6E). Consistently, overexpression of TMEM9 produced the opposite results (Fig. 6F). Thus, these results suggest that TMEM9 partially promotes VEGF expression through the MEK/ERK/STAT3 pathway in LUAD.
A The expression of p-MEK1/2, MEK1/2, p-ERK1/2, ERK1/2, p-AKT, AKT, p-STAT3, and STAT3 in LUAD stable cell lines with TMEM9 inhibition (shTMEM9) or their control cell lines (sh-NC) was analyzed by Western blot. B Western blot analysis of p-MEK1/2, MEK1/2, p-ERK1/2, ERK1/2, p-AKT, AKT, p-STAT3, and STAT3 in LUAD stable cell lines overexpressing TMEM9 (TMEM9) or their control cell lines (vector). C The protein expression of p-STAT3, STAT3, and VEGF in TMEM9 overexpressing LUAD cells treated with U0126 was detected by Western blot. D The protein expression of VEGF in TMEM9 overexpressing LUAD cells after treatment with cryptotanshinone (CTS) was detected by Western blot. E-F ChIP assays were performed in TMEM9 knockdown (E) and overexpression (F) cells with an anti-p-STAT3 antibody or IgG. Precipitated DNAs were measured by qRT-PCR for VEGF promoter.
TMEM9 knockdown inhibits tumor progression in a xenograft model and a lung metastasis model
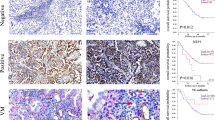
To further test the role of TMEM9 in vivo, TMEM9 knockdown A549 and control cells were subcutaneously injected into nude mice, respectively. The mice were sacrificed 28 days after injection. As shown in Fig. 7A–C, TMEM9 knockdown significantly reduced tumor volume and weight compared with the control group. Immunostaining for A549 cells in xenografts revealed that the expression of Ki67 was decreased markedly in the TMEM9 knockdown group (Fig. 7D, G). Next, we investigated the association of angiogenesis with TMEM9 in vivo. IHC staining showed that the knockdown of TMEM9 also led to a reduced expression of VEGF and MVD compared with the control (Fig. 7E–F, H–I).
A–C TMEM9 knockdown A549 cells and control cells were injected subcutaneously into nude mice. Tumor sizes (A), weight (B), and growth (C) were shown. D-I A lower expression of Ki67, CD31, and VEGF was detected in the xenograft with TMEM9 knockdown. J-K LUAD cells were injected into the tail vein of nude mice. TMEM9 knockdown significantly reduced the number of pulmonary metastatic foci. L Schematic diagram of this study. Each bar represents the mean ± SD. *P < 0.05, **P < 0.01.
In addition, we explored the effect of TMEM9 on lung metastasis. LUAD cells were injected into the tail vein of nude mice, and lung metastases were estimated. We found that TMEM9 knockdown resulted in fewer metastatic lesions than the control group (Fig. 7J–K). However, there were no significant differences in the expression of CD31 and VEGF in lung metastases between the two groups (Supplementary Fig. S6).
Discussion
TMEM9, a new member of the transmembrane protein family, is involved in inflammation, tissue regeneration, cell differentiation, and proliferation. The role of TMEM9 in tumors has garnered increasing attention in recent years. Zhang et al. found that high expression of TMEM9 in HCC patients had a shorter survival period. Downregulation of TMEM9 in HCC cell lines can inhibit cell growth and metastasis and promote apoptosis, which may be a potential target for HCC therapy [10]. In addition, several studies showed that TMEM9 could facilitate the assembly of v‐ATPase, which leads to vesicular acidification and lysosomal dysfunction that promotes tumor occurrence in hepatocellular carcinoma and colorectal cancer [7, 9]. Zhang et al. have recently established the significance of the lysosomal TMEM9-V-ATPase - regulator - Rag axis as a crucial regulator of mTOR signal integrity and proliferation of breast cancer cells [8]. This axis also influences the sensitivity of breast cancer cells to mTOR inhibitors, thereby presenting itself as a promising molecular target for breast cancer treatment and a therapeutic target for mTOR inhibitor-resistant patients. Another study reported that FOXD2-AS1 regulates TMEM9 to mediate sorafenib resistance in HCC cells [11]. The findings indicate that TMEM9 may have crucial implications in the processes of tumorigenesis and development. But the role of TMEM9 in LUAD remains to be investigated. In the study, we investigated the expression and function of TMEM9 in LUAD and demonstrated that TMEM9 promoted tumor cell proliferation, migration, and angiogenesis via increasing VEGF expression and secretion. Recently, several studies have reported that TMEM9 was overexpressed in multiple cancers and significantly correlated with poor prognosis [8,9,10]. Our results showed that TMEM9 mRNA and protein expression in LUAD tissues was higher than those in normal peritumorial tissues. The expression of TMEM9 was significantly correlated with clinicopathologic characteristics, including higher T stage, TNM stage, histological grade, and STAS. In addition, we found that TMEM9 expression was an independent indicator of poor prognosis of LUAD patients, suggesting TMEM9 was involved in tumorigenesis and progression of LUAD.
It has been reported that TMEM9 promotes the proliferation of breast cancer and HCC cells [7, 8]. In the study, we demonstrated that TMEM9 increased the proliferation and migration of LUAD. Angiogenesis has been considered a characteristic feature of rapidly growing solid tumors [21] and is closely associated with tumor growth and metastasis [22, 23]. We first observed a positive correlation between the expression level of TMEM9 and MVD in patient tumor tissues. In vitro experiments further demonstrated that TMEM9 knockdown in LUAD cells reduced the proliferation, migration, and tube formation ability of HUVEC. In an animal model, we also found that TMEM9 knockdown significantly reduced CD31 expression in the xenograft. These data suggest that TMEM9 may play an essential role in promoting angiogenesis.
Tumor angiogenesis is a complex process in which multiple cells and cytokines are involved [24]. The interactions between tumor cells and endothelial cells (ECs) influence tumor angiogenesis [25].VEGF is one of the most critical regulatory factors of tumor angiogenesis and promotes tumor progression, making it a key target of anticancer therapy in various malignant tumors [26,27,28]. We discovered that TMEM9 knockdown inhibited the expression and secretion of VEGF in LUAD cells. Bevacizumab reversed the positive effect of TMEM9 overexpressed LUAD cells on HUVEC angiogenesis, while recombinant VEGF abolished the inhibitory effects of shTMEM9 LUAD cells on HUVEC angiogenesis. These findings indicate that TMEM9 promotes LUAD angiogenesis in a VEGF dependent manner.
Furthermore, VEGF driven from tumor cells also enhances its ability to invade and migrate [29,30,31,32,33]. The present study revealed that the effect of TMEM9 on the migration of LUAD cells was partially dependent on VEGF. Previous studies suggested that VEGF expression is regulated by several signaling pathways, such as the transcription factor STAT3, MEK/ERK, or AKT pathways [34,35,36]. Our results showed that TMEM9 increased the phosphorylation levels of MEK, ERK, and STAT3. Previous studies have shown that STAT3 is a downstream target of the ERK signaling pathway in various tumors [37,38,39,40]. Consistent with previous findings, we also found that U0126 treatment reduced protein levels of p-STAT3 in TMEM9-overexpressing LUAD cells. Studies have confirmed that STAT3 is a direct transcriptional activator of VEGF gene [41]. In this study, we demonstrated that MEK inhibitor (U0126) and STAT3 inhibitor (CTS) inhibited the promoting effect of TMEM9 on VEGF expression in LUAD cells. In addition, we confirmed that TMEM9 can promote p-STAT3 binding to VEGF promoters. Taken together, our data established TMEM9 as a candidate biomarker for the prognosis of lung adenocarcinoma and favored tumor progression. Targeting TMEM9/MEK/ERK/STAT3/VEGF pathway may exert some antitumor effect.
Data availability
The raw data acquired for this study are available from the corresponding author upon reasonable request.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2021;71:209–49.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA: Cancer J Clin. 2019;69:7–34.
Lortet-Tieulent J, Soerjomataram I, Ferlay J, Rutherford M, Weiderpass E, Bray F. International trends in lung cancer incidence by histological subtype: adenocarcinoma stabilizing in men but still increasing in women. Lung cancer (Amst, Neth). 2014;84:13–22.
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ, Hodis E, et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell. 2012;150:1107–20.
Kveine M, Tenstad E, Døsen G, Funderud S, Rian E. Characterization of the novel human transmembrane protein 9 (TMEM9) that localizes to lysosomes and late endosomes. Biochem Biophys Res Commun. 2002;297:912–7.
Wei W, Jiang F, Liu XC, Su Q. TMEM9 mediates IL-6 and IL-1β secretion and is modulated by the Wnt pathway. Int Immunopharmacol. 2018;63:253–60.
Jung YS, Stratton SA, Lee SH, Kim MJ, Jun S, Zhang J, et al. TMEM9-v-ATPase ativates Wnt/β-catenin signaling via APC isosomal degradation for liver regeneration and tumorigenesis. Hepatol (Baltim, Md). 2021;73:776–94.
Zhang S, Lee SH, Nie L, Huang Y, Zou G, Jung YS, et al. Lysosomal TMEM9-LAMTOR4-controlled mTOR signaling integrity is required for mammary tumorigenesis. Cancer Commun (Lond, Engl). 2023;43:159–63.
Jung YS, Jun S, Kim MJ, Lee SH, Suh HN, Lien EM, et al. TMEM9 promotes intestinal tumorigenesis through vacuolar-ATPase-activated Wnt/β-catenin signalling. Nat Cell Biol. 2018;20:1421–33.
Zhang Y, Ran Y, Xiong Y, Zhong ZB, Wang ZH, Fan XL, et al. Effects of TMEM9 gene on cell progression in hepatocellular carcinoma by RNA interference. Oncol Rep. 2016;36:299–305.
Sui C, Dong Z, Yang C, Zhang M, Dai B, Geng L, et al. LncRNA FOXD2-AS1 as a competitive endogenous RNA against miR-150-5p reverses resistance to sorafenib in hepatocellular carcinoma. J Cell Mol Med. 2019;23:6024–33.
Yang Z, Liang X, Fu Y, Liu Y, Zheng L, Liu F, et al. Identification of AUNIP as a candidate diagnostic and prognostic biomarker for oral squamous cell carcinoma. EBioMedicine. 2019;47:44–57.
Akashi Y, Oda T, Ohara Y, Miyamoto R, Kurokawa T, Hashimoto S, et al. Anticancer effects of gemcitabine are enhanced by co-administered iRGD peptide in murine pancreatic cancer models that overexpressed neuropilin-1. Br J Cancer. 2014;110:1481–7.
Győrffy B, Surowiak P, Budczies J, Lánczky A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One. 2013;8:e82241.
Ferrara N. VEGF and the quest for tumour angiogenesis factors. Nat Rev Cancer. 2002;2:795–803.
Yang S, Yang C, Yu F, Ding W, Hu Y, Cheng F, et al. Endoplasmic reticulum resident oxidase ERO1-lalpha promotes hepatocellular carcinoma metastasis and angiogenesis through the S1PR1/STAT3/VEGF-A pathway. Cell Death Dis. 2018;9:1105.
Zhu CC, Chen C, Xu ZQ, Zhao JK, Ou BC, Sun J, et al. CCR6 promotes tumor angiogenesis via the AKT/NF-κB/VEGF pathway in colorectal cancer. Biochim et Biophys Acta Mol Basis Dis. 2018;1864:387–97.
Ding C, Luo J, Fan X, Li L, Li S, Wen K, et al. Elevated Gab2 induces tumor growth and angiogenesis in colorectal cancer through upregulating VEGF levels. J Exp Clin Cancer Res: CR. 2017;36:56.
Wang Y, Crisostomo PR, Wang M, Markel TA, Novotny NM, Meldrum DR. TGF-alpha increases human mesenchymal stem cell-secreted VEGF by MEK- and PI3-K- but not JNK- or ERK-dependent mechanisms. Am J Physiol Regul Integr Comp Physiol. 2008;295:R1115–1123.
Zeng FM, Li YW, Deng ZH, He JZ, Li W, Wang L, et al. SARS-CoV-2 spike spurs intestinal inflammation via VEGF production in enterocytes. EMBO Mol Med. 2022;14:e14844.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Weidenaar AC, ter Elst A, Kampen KR, Meeuwsen-de Boer TG, de Jonge HJ, Scherpen FJ, et al. Stromal interaction essential for vascular endothelial growth factor A-induced tumour growth via transforming growth factor-β signalling. Br J cancer. 2011;105:1856–63.
Bielenberg DR, Zetter BR. The contribution of angiogenesis to the process of metastasis. Cancer J (Sudbury, Mass). 2015;21:267–73.
De Palma M, Biziato D, Petrova TV. Microenvironmental regulation of tumour angiogenesis. Nat Rev Cancer. 2017;17:457–74.
Bergers G, Benjamin LE. Tumorigenesis and the angiogenic switch. Nat Rev Cancer. 2003;3:401–10.
Plate KH, Breier G, Weich HA, Risau W. Vascular endothelial growth factor is a potential tumour angiogenesis factor in human gliomas in vivo. Nature. 1992;359:845–8.
Ferrara N, Gerber HP, LeCouter J. The biology of VEGF and its receptors. Nat Med. 2003;9:669–76.
Chen J, Liu A, Wang Z, Wang B, Chai X, Lu W, et al. LINC00173.v1 promotes angiogenesis and progression of lung squamous cell carcinoma by sponging miR-511-5p to regulate VEGFA expression. Mol Cancer. 2020;19:98.
Bachelder RE, Wendt MA, Mercurio AM. Vascular endothelial growth factor promotes breast carcinoma invasion in an autocrine manner by regulating the chemokine receptor CXCR4. Cancer Res. 2002;62:7203–6.
Beck B, Driessens G, Goossens S, Youssef KK, Kuchnio A, Caauwe A, et al. A vascular niche and a VEGF-Nrp1 loop regulate the initiation and stemness of skin tumours. Nature. 2011;478:399–403.
Xu W, Huang JJ, Cheung PC. Extract of pleurotus pulmonarius suppresses liver cancer development and progression through inhibition of VEGF-induced PI3K/AKT signaling pathway. PLoS One. 2012;7:e34406.
Su JC, Mar AC, Wu SH, Tai WT, Chu PY, Wu CY, et al. Disrupting VEGF-A paracrine and autocrine loops by targeting SHP-1 suppresses triple negative breast cancer metastasis. Sci Rep. 2016;6:28888.
Darrington E, Zhong M, Vo BH, Khan SA. Vascular endothelial growth factor A, secreted in response to transforming growth factor-β1 under hypoxic conditions, induces autocrine effects on migration of prostate cancer cells. Asian J Androl. 2012;14:745–51.
Zhao J, Du P, Cui P, Qin Y, Hu C, Wu J, et al. LncRNA PVT1 promotes angiogenesis via activating the STAT3/VEGFA axis in gastric cancer. Oncogene. 2018;37:4094–109.
Wu X, Chen Z, Zeng W, Zhong Y, Liu Q, Wu J. Silencing of Eag1 gene inhibits osteosarcoma proliferation and migration by targeting STAT3-VEGF pathway. BioMed Res Int. 2015;2015:617316.
Lin J, Cao S, Wang Y, Hu Y, Liu H, Li J, et al. Long non-coding RNA UBE2CP3 enhances HCC cell secretion of VEGFA and promotes angiogenesis by activating ERK1/2/HIF-1α/VEGFA signalling in hepatocellular carcinoma. J Exp Clin Cancer Res: CR. 2018;37:113.
Siddharth S, Kuppusamy P, Wu Q, Nagalingam A, Saxena NK, Sharma D. Metformin enhances the anti-cancer efficacy of sorafenib via suppressing MAPK/ERK/Stat3 axis in hepatocellular carcinoma. Int J Mol Sci. 2022;23:8083.
Lee GW, Hur W, Kim JH, Park DJ, Kim SM, Kang BY, et al. Nardostachys jatamansi root extract attenuates tumor progression in hepatocellular carcinoma via inhibition of ERK/STAT3 pathways. Anticancer Res. 2021;41:1883–93.
Wang LL, Luo J, He ZH, Liu YQ, Li HG, Xie D, et al. STEAP3 promotes cancer cell proliferation by facilitating nuclear trafficking of EGFR to enhance RAC1-ERK-STAT3 signaling in hepatocellular carcinoma. Cell Death Dis. 2021;12:1052.
Yoon JH, Pham TH, Lee J, Lee J, Ryu HW, Oh SR, et al. Methyl linderone suppresses TPA-stimulated IL-8 and MMP-9 expression via the ERK/STAT3 pathway in MCF-7 breast cancer cells. J Microbiol Biotechnol. 2020;30:325–32.
Wei D, Le X, Zheng L, Wang L, Frey JA, Gao AC, et al. Stat3 activation regulates the expression of vascular endothelial growth factor and human pancreatic cancer angiogenesis and metastasis. Oncogene. 2003;22:319–29.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NO. 82072927), Shandong Provincial Natural Science Foundation (No.ZR2022MH301), and the Clinical Scientific Research Program of Wu Jieping Medical Funding (NO.320.6750.2021-02-66).
Author information
Authors and Affiliations
Contributions
XM and YS conceived and designed the experiments. ZW and PZ performed the experiments. HD, JL and ZG participated in the animal experiments.KT provided tissue specimens and patient clinical information. ZQ and HS analyzed the data. ZW and PZ wrote the paper. YS reviewed and edited the manuscript. All authors reviewed the manuscript, and XM and YS approved publishing the review.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Stephen Tait
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Z., Zhao, P., Tian, K. et al. TMEM9 promotes lung adenocarcinoma progression via activating the MEK/ERK/STAT3 pathway to induce VEGF expression. Cell Death Dis 15, 295 (2024). https://doi.org/10.1038/s41419-024-06669-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06669-8