Abstract
The small muscle protein, X-linked (SMPX) gene encodes a cytoskeleton-associated protein, highly expressed in the inner ear hair cells (HCs), possibly regulating auditory function. In the last decade, several mutations in SMPX have been associated with X-chromosomal progressive non syndromic hearing loss in humans and, in line with this, Smpx-deficient animal models, namely zebrafish and mouse, showed significant impairment of inner ear HCs development, maintenance, and functioning. In this work, we uncovered smpx expression in the neuromast mechanosensory HCs of both Anterior and Posterior Lateral Line (ALL and PLL, respectively) of zebrafish larvae and focused our attention on the PLL. Smpx was subcellularly localized throughout the cytoplasm of the HCs, as well as in their primary cilium. Loss-of-function experiments, via both morpholino-mediated gene knockdown and CRISPR/Cas9 F0 gene knockout, revealed that the lack of Smpx led to fewer properly differentiated and functional neuromasts, as well as to a smaller PLL primordium (PLLp), the latter also Smpx-positive. In addition, the kinocilia of Smpx-deficient neuromast HCs appeared structurally and numerically altered. Such phenotypes were associated with a significant reduction in the mechanotransduction activity of the neuromast HCs, in line with their positivity for Smpx. In summary, this work highlights the importance of Smpx in lateral line development and, specifically, in proper HCs differentiation and/or maintenance, and in the mechanotransduction process carried out by the neuromast HCs. Because lateral line HCs are both functionally and structurally analogous to the cochlear HCs, the neuromasts might represent an invaluable—and easily accessible—tool to dissect the role of Smpx in HCs development/functioning and shed light on the underlying mechanisms involved in hearing loss.
Similar content being viewed by others
Introduction
Human SMPX gene was first identified in 1999 when its whole genomic structure was revealed to be made of five exons and four introns, spanning a region of about 52.1 kb1. The gene encodes a 9 kDA prolin-rich protein of 88 aa that contains two casein kinase II phosphorylation sites (CKII), a PEST sequence, and a nuclear localization signal (NLS)2,3,4. Human SMPX was initially reported to be predominantly expressed in both skeletal and cardiac muscle, whilst the murine Smpx exhibited a more complex pattern of expression as in that its expression was described also in the liver, testis, kidney and brain1.
In the last decade, mutations in the SMPX gene have been associated with nonsyndromic X-linked hearing loss DFNX44,5,6,7,8,9,10,11,12,13,14. Interestingly, SMPX was shown to be abundantly expressed in both fetal and adult human inner ear4, while mouse Smpx was concordantly expressed with the master regulator of inner ear hair cell (HC) specification, the transcription factor Atoh1, in all vestibular organs and all cochlear turns of the developing mouse inner ear, thus suggesting a Smpx putative role during HC differentiation and/or functioning15. Furthermore, HC development was found to be severely affected in zebrafish embryos lacking Rbm24a, an RNA-binding protein that directly regulates smpx mRNA stability16. As we previously demonstrated, the zebrafish Smpx protein inside the inner ear HC was mainly localized at the level of the cuticular plate (CP), an actin-rich structure that is necessary for the correct development of both stereocilia and kinocilia, and to carry out mechanotransduction17. Employing a zebrafish model of Smpx deficiency, we showed that the protein exerted a primary function in the proper structural organization of the inner ear HCs sensory bundle as in that both stereocilia and kinocilia displayed evident morphological alterations in terms of number and length11, as later also confirmed in a knock-out mouse model18. Moreover, we demonstrated that inner ear HCs of Smpx-deficient embryos lack of mechanotransduction activity, providing the first possible explanation for the reported cases of SMPX-linked hearing loss in human patients4,5,6,7,8,9,10,11,12,13,14. In this work we focused our attention on the zebrafish lateral line, a sensory system positioned on the outside of the organism, which grants the animal the ability to detect water flow changes in terms of motion and pressure19. The lateral line organ forms, together with the inner ear, the octavolateralis sensory system of fishes20. The functional units of the lateral line are the neuromasts, a series of sensory organs arrayed along the head and body of the fish. Each neuromast contains a central cluster of HCs which are structurally and functionally analogous to the HCs of the cochlea, but more easily accessible than the latter19,21,22. During development, the first neuromasts are formed through the rostrocaudal migration of the posterior lateral line primordium (from now on PLLp)—namely a transient structure generated from placodal cells adjacent to the ear—occurring between 24 and 40 h post fertilization (hpf)19,23. At around 42 hpf, other posterior lateral line primordia (primII and primD) are formed and their migration will give rise to accessory/secondary neuromasts24. Here, we describe the activity of zebrafish smpx, that is expressed in all neuromasts of the PLL. We also depicted the distribution of the encoded protein in the neuromast, which occurred exclusively in the HCs, throughout the cytoplasm and the kinocilium of the cells. Also, by means of loss-of-function experiments, we elucidated the role played by the gene in controlling the proper development and function of the neuromasts. Indeed, lack of Smpx was associated with a decrease in size of the PLLp (in which the Smpx protein was also present), fewer HCs-positive neuromasts, reduced density of kinocilia per neuromast, the alteration of the kinocilium structure and, finally, the substantial impairment in the mechanotransduction activity of the neuromast HCs.
Results
smpx expression in the neuromasts
Through an in-situ hybridization protocol with slight modifications to avoid potential neuromasts degradation, the smpx-specific transcripts were detected in the zebrafish neuromast cells (Fig. 1). The first evidence of smpx expression occurred at 96 hpf, where the signal was clearly visible in the cells of the deposited neuromasts of the head (data not shown), and along the trunk and tail of the larva (Fig. 1A). At 120 hpf, the smpx-specific signal was observed in all neuromasts of the posterior lateral line (PLL, Fig. 1A’). At 96 hpf and 120 hpf, we counted 7 neuromasts of the PLL per each side of the larva (Fig. 1A,A’) with the typical rosette-like structure (Fig. 1B,B’), plus 1–2 accessory neuromasts and 2 terminal neuromasts at the end of the tail (Fig. 1C,C’). A barely distinguishable signal was observed also at 48 and 72 hpf (data not shown), suggesting that the gene was already expressed at these earlier developmental stages, as confirmed by the presence of the protein since 48 hpf (see below).
smpx expression in the neuromasts of the posterior lateral line. Representative stereomicroscope images of in-situ hybridizations performed with the smpx riboprobe on whole mount larvae at 96 and 120 hpf. (A,A’) The gene is expressed in all the neuromasts (L1-L7, black arrowheads) establishing the lateral line at both stages analyzed. The smpx-specific signal is present also in the accessory neuromasts (green arrowheads) and in the terminal neuromasts of the tail (black arrows). (B,B’) Detailed view of smpx signal in the L2, L3 and L6 neuromasts of the trunk at 96 and 120 hpf. The white-dotted rectangles show magnified neuromasts with the typical rosette-like structure, the green arrowheads indicate the accessory neuromasts. (C,C’) Detailed view of smpx signal in the L6, L7, and terminal neuromasts (black arrows) of the tail at 96 and 120 hpf. (A,B) Neuromasts appear slightly below the median line because they were flat-mounted under a coverslip. Lateral views, anterior to the left. Scale bars are 160 µm.
Smpx localization in the neuromast hair cells
The in-situ hybridization data regarding the expression of smpx in the neuromasts were fully confirmed at the protein level. Indeed, whole mount immunofluorescence experiments with an anti-Smpx polyclonal antibody (Supp. Fig. S1), proved the localization of Smpx in the neuromasts of the PLL at 48 (Supp. Fig. S1A), 72 (Supp. Fig. S1B), 96 (Supp. Fig. S1C), 110 (Supp. Fig. S1D), and 120 hpf (Supp. Fig. S1E,F). Moreover, the Smpx protein was also localized at the level of the head neuromasts (Supp. Fig. S1B-E) that form the anterior lateral line (ALL), a structure that develops independently from the PLL25. In all 120 hpf neuromasts, Smpx localized quite exclusively in the cytoplasm of the cells (Supp. Fig. S1F, Fig. 2, Supp. Fig. S2) and never in the nucleus, in contrast to an earlier study that found the protein accumulated within the nucleus of C2C12 cells2. By employing the transgenic zebrafish line tg (sod1:sod1; hsp70:DsRed)26, in which the red fluorescence mainly paints the outermost cells of the neuromasts (Supp. Fig. S3), we precisely localized Smpx in the innermost cluster of cells (Fig. 2A) and not in the DsRed-positive external cell layer (Fig. 2B), as demonstrated by the lack of overlapping signals (Fig. 2C). Furthermore, it is clear the absence of Smpx in the presumptive accessory cells’ territory, encompassed in between the DsRed-positive outer layer and the inmost cells of the neuromast (Fig. 2C). In addition to displaying a massive signal throughout the cytoplasm, Smpx was strongly present in the microtubule-based kinocilium (Fig. 2D-F, Supp. Fig. S2). The distribution of the signal appeared quite uniform in some hair cells, while in others the protein become visible in a ‘beads-on-a-string’ configuration, or not visible at all (Fig. 2F). The Smpx localization in the primary cilium strongly supports the observation that the Smpx-positive cells are the actual mechanosensory HCs responsible for the conversion of mechanical stimuli into electric signal27. This data is in line to what we previously reported in the inner ear, where Smpx was localized exclusively in the HCs of the sensory patches11,17.
Smpx protein localization in the neuromast mechanosensory hair cells. Confocal Z-stacks of immunofluorescence on whole mount 120 hpf larvae. (A–C) Smpx (green) localizes in the most internal cells of the neuromast, namely the mechanosensory hair cells, while it is excluded from the outermost cells (red) of the transgenic line tg (sod1:sod1; hsp70:DsRed) and the presumptive accessory cell layers (white bracket) encompassed in between the outmost DsRed-positive cells and the inmost hair cells. (D–F) Smpx (red) co-localizes with acetylated tubulin (green) in the kinocilium, with Smpx showing a ‘beads-on-a-string’ distribution (arrowheads in D). In some cases, the kinocilium is not labeled by the anti-Smpx antibody (arrow in F). (A–F) Larvae were counterstained with DAPI (blue) to visualize cell nuclei. Lateral views, anterior to the left. Scale bars are 15 µm in A–C and 10 µm in D–F.
The number of intact PLL neuromasts is affected by lack of Smpx
The role of Smpx in proper neuromast development was analyzed, at first, by gene knockdown through the microinjection of a previously validated ATG-targeting smpx-specific morpholino11, then by CRISPR/Cas9-mediated F0 gene knockout. Being Smpx expressed in the mechanosensory HCs of the neuromasts, we employed the FM 4–64 vital dye to stain such cells in vivo28,29. The reduction in number of FM 4–64-positive HCs clusters was observed in the morphant larvae (data not shown) and confirmed in the smpx F0 knockout model (smpxcrispants, Supp. Fig. S4A-C). To gain insight into these observations, we performed a DAPI/acetylated tubulin co-staining in smpx morphant and control embryos, revealing a diminished number of acetylated tubulin-positive neuromasts in the entire PLL (Fig. 3A-C), and the presence of hole-like structures surrounded by nuclei in the center of the regions where homeostatic neuromasts should be deposited (insets in Fig. 3B,C). To further increase our confidence, we carried out an anti-GFP staining in the smpxcrispant larvae generated in the tg (cldnb:GFP) transgenic line background, where the GFP is expressed in all neuromast cell types30; the IHC on such transgenic line was performed to amplify the low fluorescence signal detected in live larvae, that represented a major setback for the analyses. The number of intact neuromasts appeared lower also in the smpxcrispants, supporting our above-mentioned observations (Fig. 3D-F). As for the previous case, the hypothesized lack of HCs from the center of the neuromasts left hole-like structures encircled by the other cell types (insets in Fig. 3E,F). Finally, smpx deletion performed on the tg (sod1:sod1; hsp70:DsRed) line revealed that the outer cell layer, and in turn, the overall number of PLLp-derived neuromasts, was unaltered by the lack of the protein (Supp. Figure 4D-F), confirming that Smpx deficiency does not affect all the neuromast cell types.
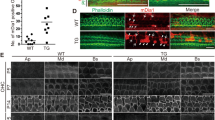
Smpx-deficiency leads to a lower number of intact neuromasts in morphant and crispant larvae. (A) Quantification of the reduced number of intact neuromasts in smpxMO versus ctrlMO larvae. (B,C) Representative confocal images of ctrlMO and smpxMO larvae labeled with an anti-acetylated tubulin antibody (magenta). Larvae were counterstained with DAPI to visualize cell nuclei. Insets in C show magnified neuromasts lacking the innermost nuclei and the knocilia of the HCs (compare to inset in B). (D) Quantification of the reduced number of intact neuromasts in smpxcrispant versus ctrl larvae. (E,F) Representative confocal images of immunofluorescence experiments with an anti-GFP antibody in ctrl and smpxcrispant tg (cldnb:GFP) transgenic larvae. Insets in F display magnified neuromasts showing the depletion of the innermost cell cluster (compare to inset in E). (A,D) Unpaired t-test was performed to assess statistical significance by setting p ≤ 0.05 (*), p ≤ 0.01 (**), p ≤ 0.001 (***) and p ≤ 0.0001 (****) as significant. (A) ctrlMO n = 8, smpxMO n = 8. (D) ctrl n = 19, smpxcrispant n = 22. (B,C and E,F) Confocal Z-stacks images taken from 120 hpf whole mount larvae; lateral views, anterior to the left. Scale bars are 500 µm.
Smpx is expressed in the posterior lateral line primordium and controls its size
Because PLL proper development was severely altered by Smpx-deficiency, we decided to study earlier steps of the organ assembly analyzing the migrating PLL primordium (PLLp). Interestingly, we found that Smpx localized also in this structure, starting from ~ 26 hpf (Fig. 4A,B). To test whether the lack of Smpx might cause an improper shaping of the PLLp, eventually leading to a defective deposition of neuromast cell precursors as development proceeds, we investigated the overall morphology of the PLLp following smpx downregulation/knockout. The prox1a RNA probe, a well-known marker for neuromast hair and supporting cells31, was used to visualize the PLLp, which appeared smaller in size as a consequence of Smpx deficiency (Fig. 4C,D). Indeed, the quantification of the prox1a-positive PLLp area showed that smpx downregulation caused a significant reduction of 1585 ± 261.8 µm2 (Fig. 4E), strongly supporting the above-mentioned hypothesis. The same outcome was obtained through smpx knockout (Fig. 4F,G), with a significant reduction of the PLLp area of 1337 ± 361.2 µm2 (Fig. 4H). Moreover, because prox1a is a selective HC/supporting cell marker, its expression permits the visualization of the rosette-like structures which guarantee the proper formation and deposition of protoneuromasts during PLL organ morphogenesis. Interestingly, such structures were not detectable in smpx morphants (compare Fig. 4C,D) and crispants (compare Fig. 4F,G). Through an in-depth analysis of the tg (cldnb:GFP) line, the rosette-like structures of the PLLp in smpxcrispants appeared numerically less dense and structurally altered compared to the normal organization of the controls (Fig. 4I,J).
Smpx-deficiency causes the decrease in the PLLp size. (A,B) Immunofluorescence with the anti-Smpx antibody unveiling the localization of the protein in the PLLp at 26 and 29 hpf. (C,D) In-situ hybridization with a prox1a probe to compare the PLLp size (white-dotted encircled areas) in 30 hpf ctrlMO and smpxMO; the PLLp of Smpx-deficient embryos appears smaller in size; the typical rosette-like structures present in the ctrlMO sample (white arrowheads in C) are not clearly distinguishable in the smpxMO sample. (E) Size quantification in ctrlMO and smpxMO showing a drastic decrease in the PLLp area following the downregulation of smpx. (F,G) In-situ hybridization with a prox1a probe to compare the PLLp size (white-dotted encircled areas) in 30 hpf ctrl and smpxcrispant embryos; the PLLp of the smpxcrispant embryos appears smaller in size and the typical rosette-like structures present in the ctrl sample (white arrowheads in F) are not clearly distinguishable in the smpxcrispant preparation. (H) Size quantification in ctrlMO and smpxcrispant showing a drastic decrease in the PLLp area following the ablation of smpx. (I,J) Z-stack confocal images of the PPLp at 30 hpf in tg (cldnb:GFP) embryos; the shape of the epithelial rosettes in control embryos are lost in smpxcrispants. (A,B) Confocal Z-stacks taken from whole mount embryos; lateral views, anterior to the left. Embryos were counterstained with DAPI to visualize cell nuclei. (E,H) Unpaired t-test was performed to assess statistical significance by setting p ≤ 0.05 (*), p ≤ 0.01 (**), p ≤ 0.001 (***) and p ≤ 0.0001 (****) as significant. (E) ctrlMO n = 22, smpxMO n = 18. (H) ctrl n = 10, smpxcrispant n = 13. Scale bars are 50 µm.
The mechanotransduction activity of the neuromast ciliated cells depends on Smpx
Kindt and colleagues demonstrated that neuromast HCs require kinocilia for proper mechanosensation32. In line with the evidence that Smpx localizes also along the kinocilium, we tested the role of the protein in lateral line mechanotransduction by means of gene loss-of-function. Smpx downregulation caused a reduction in the mechanotransduction activity of the deposited neuromast HCs, at 120 hpf, as demonstrated by incubating live larvae in embryo water containing the FM 4–64 vital dye. Indeed, the lower mechanotransduction activity was highlighted by the defective uptake of the dye (Fig. 5A–C), resulting in a significant decrease in the overall fluorescence intensity in smpx morphant larvae compared to controls (Fig. 5D).
Smpx-deficiency causes a reduced mechanotransduction activity of the neuromast hair cells. (A–C) Representative images of FM 4–64 incorporation in one control neuromast (ctrlMO) and two Smpx-deficient (smpxMO) neuromasts at 120 hpf (white rectangles on the right of each single image are magnifications of the corresponding, white-dotted rectangles on the left); compared to the control (A), smpx-morphants (B,C) display a reduced FM 4–64 dye uptake. (D) Quantification of the FM 4–64 dye uptake calculated as fluorescence intensity per each neuromast (n = 10 for both controls and morphants). (A–C) Confocal Z-stacks taken from whole mount larvae. (D) Unpaired t-test was performed to assess statistical significance by setting p ≤ 0.05 (*), p ≤ 0.01 (**), p ≤ 0.001 (***) and p ≤ 0.0001 (****) as significant. Scale bars are 50 µm.
Smpx is required for normal kinocilium structure
Based on the lack of mechanosensitivity following Smpx deficiency, we analyzed, by immunohistochemistry, both the general morphology of the primary cilium of the HC and the structure of the neuromasts. The number of kinocilia in morphant neuromasts was clearly decreased in comparison to controls, and such massive reduction was accompanied by the severe alteration in the overall length of the kinocilia (Fig. 6A,B). Again, to strengthen such evidence, we investigated the consequence of smpx knockout on neuromasts assembly, gaining comparable results. Indeed, in smpxcrispant larvae, the primary cilia of the neuromasts appeared dramatically less dense compared to controls (Fig. 6C,D).
Smpx is required for normal kinocilium structure. Immunofluorescence performed on whole mount 120 hpf larvae with an antibody against acetylated tubulin to label the neuromast kinocilia. (A,B) Compared to controls (ctrlMO), the kinocilia of morphant larvae (smpxMO) appear clearly shorter in length and numerically less dense. (C,D) As for the knockdown experiment, when compared to controls (ctrl), the kinocilia of crispant larvae (smpxcrispant) are shorter and numerically lower. (A,B) ctrlMO n = 5, smpxMO n = 4. (C,D) ctrlMO n = 5, smpxMO n = 5. (A–D) Confocal Z-stack images of neuromast L4 for each larva. n is equal to the sum of the larvae from two independent experiments. Embryos were counterstained with DAPI to visualize cell nuclei. Scale bars are 6 µm.
Discussion
In addition to our previous expression data17, whole mount in-situ hybridization (WISH) experiments revealed that smpx was also expressed in the neuromasts, namely the specialized ciliated structures located along the trunk of the animal which establish the lateral line organ33. The expression was present, up to the latter stage analyzed (120 hpf), in the neuromasts of the head, in those placed all along the trunk and tail of the embryo, as well as in the terminal neuromasts of the tail. In line with the mRNA expression, the protein was also localized in the neuromasts starting from 48 hpf, the earlier stage analyzed. Specifically, by employing the transgenic line tg (sod1:sod1, hsp70:DsRed), that we used as a reporter line for its ability to drive DsRed expression in the outer cell layer of the neuromasts, we demonstrated that Smpx is specifically localized in the most internal cells of the ciliated structures; these cells are the actual sensors responsible for mechanosensation34. Strongly reinforcing our data, previous scRNA-seq experiments revealed smpx mRNA to be highly enriched in the transcriptome of both young and mature HCs of homeostatic neuromasts, when compared to the transcriptome profiles of supporting and mantle cells35. However, both our most in-depth immunofluorescence experiments for Smpx protein localization and the scRNA-seq by Lush and colleagues35 were performed only with 120 hpf neuromasts and it is not possible to exclude a priori the expression of smpx also in other neuromast cell types at different developmental stages. Quite interestingly, as clearly visible at 120 hpf, the protein was localized not only throughout the cytoplasm of the neuromast sensory HCs, but also at the level of the kinocilium. This last data is extremely surprising, because Smpx was never detectable in kinocilium of the HCs of the zebrafish larva inner ear at all stages analyzed17. A possible explanation for such apparent discrepancy could be represented by the different timing at which the HCs in the two districts acquire a more and more mature-type organization, as previously suggested for later stages of development36. Notably, some neuromasts displayed Smpx arranged in a ‘beads-on-a-string’ fashion all along the kinocilium and in each of the HC of the structure, while in other neuromasts Smpx signal painted the kinocilium of some HCs but was completely absent from others. This is very interesting considering that primary cilia are highly dynamic organelles in which their functioning is strictly dependent on ciliary proteins turnover and subcellular localization37. Therefore, the intense trafficking of ciliary membrane proteins from the Golgi apparatus towards ciliary compartments may represent the event underlying the heterogeneity of Smpx signal in the kinocilia observed in this work. Functional experiments via gene loss-of-function, by using both morpholino and CRISPR/Cas9 approaches, showed that the lack of Smpx led to a reduction in the number of fully formed neuromasts. Interestingly, it was recently demonstrated in homozygous zebrafish mutants for rbm24a—a gene encoding an RNA-binding protein that directly regulates the stability of the smpx transcript—a similar defective HCs morphogenesis16, further supporting the link between these two genes and their putative co-presence in the same pathway during neuromast HCs development/maintenance. Indeed, smpx was found downregulated in rbm24a mutants, indicating that the reason for the reduced number of neuromast HCs in the mutant larvae16 might be, in fact, the insufficient amount of Smpx synthesized by the HCs. On the other hand, while in rbm24a mutants the number of neuromasts appeared lower16, our experiments revealed that the overall number of deposited PLLp-derived neuromasts at 120 hpf was mostly unaffected in Smpx-deficient larvae, thus ruling out the hypothesis that rbm24a and smpx might act together also during the gross deposition of protoneuromasts. Interestingly, Smpx starts to be detectable in the PLLp of control embryos already at 26 hpf, as well as few hours later, when the PLLp starts to release the first neuromast. Thus, the localization of Smpx in the PLLp, and the defective phenotype of the lateral line organ in Smpx-deficient larvae, was suggestive that the lower number of HCs positive neuromasts could be the direct consequence of the PLLp shrinkage that takes place at earlier developmental stages. By visualizing the PLLp with the prox1a riboprobe and with the tg (cldnb:GFP) line, we observed that these structures of most Smpx-deficient embryos were largely deprived of the HCs precursors. Indeed, the classical rosette-like structures (with the precursors at their center) that form into the trailing zone of the PLLp and, in turn, assemble themselves into protoneuromasts under the control of the Fgf signaling38,39, were mostly absent or barely distinguishable. Although we reported a defective phenotype in both PLLp- and primII/primD-derived neuromasts, the alteration of the overall structure of the PLLp in Smpx-deficient embryos might account only for those neuromasts that are deposited from the PLLp. Therefore, further investigations will be required to assess not only whether Smpx localizes in the secondary primordia, but also to comprehend the upstream event underlying the aberrant morphology that we observed also in the accessory/secondary neuromasts. Interestingly though, the PLLp and primII/primD are formed from their respective primary and secondary placodes upon the same retinoic acid signaling during late gastrulation, suggesting a common origin and a strict interconnection24. Due to the prominent presence of Smpx in the neuromast primary cilia, we searched for possible structural kinocilium alteration following Smpx ablation. As expected, the HC kinocilia of Smpx-deficient neuromasts were shorter compared to those of the controls. Moreover, although it was not possible to precisely quantify the number of primary cilia because they overlapped each other, kinocilia of Smpx-deficient embryos appeared visually less dense than controls, thus suggesting a reduction in number, as occurred in the inner ear HCs11. The mechanism by which Smpx-deficiency causes primary cilia degeneration/developmental impairment is still unknown and remains highly speculative, thus requiring additional investigations. Nevertheless, our analysis pointed out that the lack of Smpx in the neuromast HCs caused a reduction in the mechanotransduction activity of such cells, as demonstrated by the lower FM 4–64 dye fluorescence, in line to what we observed in the inner ear HCs11.
In conclusion, our study provides further evidence of the role played by smpx in neuromast HC development and functioning. This work highlights novel territories of smpx expression, namely the mechanosensory HCs of the neuromasts and the PLLp and demonstrates the central role of Smpx in controlling the shaping of the PLL sensory organ and its mechanosensitivity (Fig. 7). This will be of great interest for hearing research (zebrafish lateral line represents a useful tool to study human hearing loss40) as in that the neuromasts are much more accessible in terms of imaging assays and drugs administration. Because of the lack of definitive therapeutic options for genetic deafness, the combination of the knockout models under generation, the expression of smpx in the mechanosensory ciliated cells of the neuromasts and the functional, genetical and evolutionarily comparability of such cells with those of the human inner ear28 will eventually lead to the establishment of an in-vivo platform for fast and unbiased pharmacological screenings of novel (or repositioned) molecules capable of reverting the pathological condition.
smpx shapes the lateral line organ and affects its mechanosensitivity. Smpx-deficiency affects the number of intact neuromasts in the PLL, possibly because of the role of the gene in directing the development of the HCs in the PLLp. Furthermore, Smpx is essential in proper kinocilium modeling and HCs mechanotransduction.
Materials and methods
Zebrafish husbandry and maintenance
Fish were maintained in the facility of the University of Milan, Via Celoria 26, 20,133 Milan, at 28 °C on a 14-h light/10-h dark cycle. The temperature of the water in the aquarium was maintained between 28 °C and 28.5 °C, with the pH spanning between 6.8 and 7.5. Fish in the aquarium were fed three times per day with granular food of different dimension according to growth phases. Our facility strictly complies with the relevant European (EU Directive 2010/63/EU for animal experiments) and Italian (Legislative Decree No. 26/2014) laws, rules, and regulations (Auth. Min. 24/2019-UT), as also confirmed by the authorization issued by the municipality of Milan (PG 198,283/2019). All procedures were carried out in accordance with the relevant Italian and European guidelines and regulations, and the ARRIVE guidelines (https://arriveguidelines.org/).
Zebrafish lines used in this work and egg collection
In the experiments involving embryos, the zebrafish lines used were as follows: wild-type of the AB and ABTL strains, tg (sod1:sod1; hsp70:DsRed) transgenic line26, and tg (cldnb:GFP)30. Embryos were collected by means of natural spawning, picked up with a plastic Pasteur pipette and staged in embryo water-containing Petri dishes for downstream analysis, according to Kimmel and co-workers41. To avoid pigmentation, embryos used in whole mount in-situ hybridization (WISH) and immunofluorescence experiments were raised in embryo water supplemented with 0.003% 1-phenyl-2 thiourea (PTU), until the desired developmental stage. For WISH experiments, embryos were fixed at least overnight in 4% paraformaldehyde (PFA) pH 7.2 in 1 × PBS at 4 °C. For immunofluorescence experiments, embryos were fixed 1.5/2 h at room temperature in 4% PFA pH 7.2 in 1X PBS, to avoid background fluorescence. Following fixation, they were dehydrated with increasing concentrations (30%, 50%, 70%, 100% two times) of methanol (MetOH) in PBT (1X PBS, 0.1% Tween20®) for 5 min. Dehydrated embryos were stored in absolute methanol at -20 °C for at least 1 h or until required. For immunofluorescence experiments requiring phalloidin staining, dehydration in methanol was avoided and embryos were stored in PBT at 4 °C, following PFA fixation.
Whole mount in-situ hybridization (WISH)
Dig-tagged smpx antisense riboprobe was synthesized with the T7 RNA polymerase (Roche) as previously described17 from a pre-existent template31. In-situ hybridization was carried out by using standard procedures on embryos rehydrated with decreasing concentrations (75%, 50%, 25%) of MetOH/PBT for 5 min each, then washed twice with PBT. Minor adjustments to the incubation time in 10 µg proteinase K (pK), for the permeabilization step, were made as follows to avoid potential damages to the neuromasts and PLLp: no pK for 24 hpf embryos, 2.5 min for 29 and 30 hpf embryos, 20 min for 48 hpf embryos, 30 min for 72 hpf larvae and 40 min for 110/120 hpf larvae. PBT washes during the first day were performed with a Tween20® concentration of 0.02% to avoid excessive permeabilization.
Whole mount immunofluorescence
Dehydrated embryos stored in absolute MetOH at -20 °C were rehydrated with decreasing concentrations (75%, 50%, 25%) of MetOH/PBT for 5 min each, then washed twice with PBT. Embryos were incubated with 10 µg/ml pK in PBS-TT (1X PBS, 0.1% Tween20®, 0.1% Triton X-100 (Sigma-Aldrich)) for a variable time, according to their developmental stage, as described in the previous section. Then, the embryos were rinsed twice for 5 min in 1X PBS, post-fixed with 4% PFA in 1X PBS for 20 min at RT, washed three times with PBS-TT and incubated for 2 h at room temperature in a blocking Solution consisting of 5% bovine serum albumin (Sigma-Aldrich) in PBS-TD (1% DMSO in PBT). Embryos were incubated overnight at 4 °C with primary antibody in Blocking Solution. Primary antibodies and their dilutions used in this work were as follows: rabbit polyclonal anti-SMPX IgG antibody (PA3-070) (ThermoFisher Scientific) 1:100, mouse anti-acetylated α-tubulin (T7451, Merck) 1:200, rabbit anti-GFP antibody (ab290, Abcam). The next day, several washes with PBS-TD were performed, then embryos were incubated overnight at 4 °C with secondary antibody in PBS-TD. The following secondary antibodies and dilutions were used: Alexa Fluor 488 F(ab’)2-Goat anti-rabbit IgG (H + L) Secondary Antibody (A-11070) (ThermoFisher Scientific) 1:200, Alexa Fluor 555 F(ab’)2-Goat anti-mouse IgG (H + L) Secondary Antibody (ThermoFisher Scientific) 1:200, Alexa Fluor 555 F(ab’)2-Goat anti-rabbit IgG (H + L) Secondary Antibody (ThermoFisher Scientific) 1:200. Embryos were counterstained, when required, with 4’,6-diamidino-2-phenylindole (DAPI), 1:2000, to visualize cell nuclei. The last day, several washes in 1X PBS were performed.
FM 4–64 dye staining of neuromast hair cells
Neuromast hair cells were labelled as described by Erickson and colleagues29. Briefly, larvae were incubated in 3 µM FM 4–64 (ThermoFisher Scientific), diluted in embryo water for 60 s, then rinsed at least four time in embryo water.
Preparation of crRNA:tracrRNA duplex and Cas9 enzyme
Target sites in the zebrafish smpx genomic locus were found using the ChopChop online tool (https://chopchop.cbu.uib.no), by selecting the crRNAs with the best predicted efficiency and the lowest off-target activity (Supp. Fig. S5A). For the generation of F0 smpx knockout embryos, we used the protocol described by Hoshijima and colleagues42 with slight modifications. Chemically synthesized smpx-specific Alt-R crRNAs (smpx_exon2: 5’-AGGGCTTTGACGTTGGACGA-3’, smpx_prom: 5’-AATAGTCGTGTGATACACAG-3’, smpx_3’UTR: 5’-CAATTCGAATCTCTGAACCA-3’) and universal Alt-R tracrRNA were purchased from Integrated DNA Technologies (IDT). Duplex buffer (IDT) was used to resuspend the two RNA oligos into 100 μM stock solutions that were then aliquoted and stored at -20 °C. The 25 μM duplex cRNA:tracrRNA (dgRNA) was generated by mixing equal volumes of the two 100 μM RNA oligos with duplex buffer under the following annealing protocol: 95 °C for 5 min, cool at 0.5 °C/sec to 25 °C, 25 °C for 10 min, cool to 4 °C rapidly. The 25 μM dgRNA was stored at -20 °C. Alt-R® S.p. Cas9 nuclease, v.3 (IDT) stored as small aliquots at -20 °C. dgRNAs:Cas9 ribonucleoprotein complex (RNP) for microinjection was generated by gently mixing equal volumes of the three 25 μM dgRNAs (to obtain 5 μM of each dgRNA) with nuclease-free water. The Cas9 was used at 15 μM concentration to obtain a 1:1 dgRNAs/Cas9 ratio. These 3 smpx-specific dgRNAs (smpx_exon2, smpx_prom, and smpx_3’UTR) were coinjected to generate the crispant larvae used in all the experiments reported in the result section of this paper. Indeed, three synthetic gRNAs were reported to achieve more than 90% biallelic gene knockouts in F043 and for the purpose of this work are believed to increase the efficiency in generating RNA-less embryos.
Microinjections
Microinjections were performed at the 1-cell stage embryos (for CRISPR/Cas9 experiments we typically used 1-cell stage embryos before cell inflation). The following ATG-targeting morpholino (MO) was used to achieve the repression of smpx mRNA translation: 5′-AAGGAAAGTGCTGTTCCCTGGTGTC-3′. MO was diluted in Ringer’s solution (116 mM NaCl, 2.9 mM KCl, 1.8 mM CaCl2, 5 mM HEPES, pH 7.3) and injected at a final concentration of 0.3 pmol/embryo in a 4 nL droplet as previously described11, using rhodamine or FITC dextran (Sigma-Aldrich) as dye tracer. As a control for each experiment involving smpx downregulation, a standard control morpholino (ctrlMO) was injected at the same concentration. For the CRISPR/Cas9 experiments, we typically prepared the injection solution as follows: 1 µL of each of the three 25 µM duplex, 1.23 µL 61 µM Cas9 enzyme, 0.5 µL 5% rhodamine or FITC dextran dye tracer and 0.27 µL nuclease-free water42. To test the efficiency of the CRISPR/Cas9 approach, we singularly injected the RNP targeting exon2, or co-injected the 2 RNPs targeted against the promoter and 3’ region (see section “efficiency of the CRISPR/Cas9 approach”) to delete the entire smpx locus, maintaining the 1:1 dgRNA/Cas9 ratio. Before microinjecting, the solutions were incubated at 37 °C for 10 min, then left at room temperature; 1 nL of this solution was injected into the yolk. At 24 hpf, the injected embryos were screened and only those exhibiting either the rhodamine- or the FITC-specific fluorescence were used for the experiments. For the CRISPR/Cas9 experiments, uninjected or Cas9-injected embryos were used interchangeably as controls.
Genomic DNA extraction for indels identification
Genomic DNA extraction was carried out with the zebrafish hotshot method44. Briefly, single 48 hpf embryos (at least 5 crispants and 3 controls) were incubated with 50 mM NaOH at 95 °C for 30 min. After cooling down to 4 °C, 1M Tris HCl (pH 8.2) was added to buffer the solution. PCR was performed in 25 µL by using 1 μL of the previously extracted DNA and the following primers: gen_smpx_T7E1_ff (5’-TAGCGTGTGTGTATCCCAGG-3’) and gen_smpx_RevA (5’-CAAGTGTCGACTGTTGGTTAGG-3’) for the RNP targeting exon 2; smpx_DltProm_FF (5’-TCTCTCTTTTCTGAGACCCCAC-3’) and smpx_Dlt3UTR_rr (5’-TCATTTTAAGGTGAACCATCCC-3’) primers were used for the CRISPR/Cas9 experiment aimed at deleting the whole smpx locus.
Efficiency of the CRISPR/Cas9 approach
To assess the mutagenesis capacity of the RNP complex targeting exon 2, an aliquot of the above-described PCR (gen_smpx_T7E1_ff/gen_smpx_RevA) was sequenced (EUROFINS SRL) using the primer gen_smpx_ffA (5’-TAAATACAGAGGCGCAGACAAA-3’), located upstream of the target site. The sanger sequencing results of control and five injected embryos (AB1 files) were uploaded in the free online CRISPR analysis tool Inference of CRISPR Edits (ICE; https://www.synthego.com/products/bioinformatics/crispr-analysis) as previously described45. The results of the smpx_DltProm_FF/ smpx_Dlt3UTR_rr PCRs were loaded onto agarose gel and electrophoresis was performed to assess the presence/absence of deletion fragments. In supplementary Fig. 5 is depicted the efficiency of mutagenesis obtained using either the smpx_exon2 dgRNA alone, generating INDEL errors (Supp. Fig. S5B), or the smpx_prom/smpx_3’UTR dgRNAs pair inducing the deletion of the whole smpx locus (Supp. Fig. S5C). The ability of the coinjection of the three dgRNAs in generating RNA-less smpxcrispant embryos was assessed by means of RT-qPCR (Supp. Fig. S5D).
RNA extraction, cDNA synthesis and qPCR
A minimum of 15 embryos at 24 hpf were pooled and total RNA extracted using the ReliaPrep™ RNA Tissue Miniprep System (Promega) according to the manufacturer’s instructions. cDNA synthesis was carried out by the M-MLV Reverse Transcriptase kit (GeneSpin Srl) starting from 1 μg of total RNA. qPCR was run with the CFX Connect Real-Time PCR detection machine (Bio-Rad Laboratories, Inc.). For each biological replicate, three technical replicates were performed. Briefly, 1 μL of cDNA was amplified in a 20 μL total volume reaction containing a half volume of SYBR Green Master Mix (GeneSpin Srl) and 0.2 μM of each primer. Primer sequences were as follows: RT_smpx_FF (5’-TTTAACACCAGCAGCAGGAAC-3’), RT_smpx_RR (5’-CATCGGGATATTAAGGTTGGCC-3’), rna18s_FF (5’-AGGAATTCCCAGTAAGCGCA-3’), rna18s_RR (5’-ACCTCACTAAACCATCCAATC-3’). Normalization was carried out by analyzing the expression levels of RNA18s in parallel with smpx. The comparative ΔΔCt method was used to analyze data.
Imaging
Embryos subjected to WISH, immunohistochemistry, and live larvae subjected to FM 4–64 labeling for neuromast counting/fluorescence quantification were mounted laterally on a glass slide with 80% glycerol (WISH and immunohistochemistry), and 1% low gelling agarose (live larvae). They were imaged and counted singularly by digital camera DFC310FX (Leica) and a Leica Application Suite V4.7 software on a MZFLIII Fluorescence Stereomicroscope (Leica). For embryos hybridized with the prox1a RNA probe, PLLp area was measured either by manually drawing the region of interest (ROI) with ImageJ v1.51 or by using the thresholding method. For FM 4–64 dye fluorescence intensity quantification, ImageJ v1.51 was used as previously described29. Confocal images were obtained on a Nikon A1 laser-scanning confocal microscope (Nikon Instruments Inc.) and a NIS-Elements C software. Optical sections (Z-stacks) were merged into one maximum projection with Image J. Final panels of images were set up and, when required, adjusted for both contrast and brightness using Adobe Photoshop CS6 (Adobe Systems Inc.).
Statistical analyses
All the experiments were performed at least twice. For each CRISPR/Cas9 experiment, at least 10 embryos were genotyped at 24 hpf to assess the goodness of smpx deletion. For statistical analysis, data were taken from a single experiment and reported as means + / − sd. Unless specified, biological replicates are the individual embryos analyzed in each experiment. Graphs were generated with GraphPad Prims.
Data availability
All data generated during this study are included in the manuscript.
References
Patzak, D., Zhuchenko, O., Lee, C.-C. & Wehnert, M. Identification, mapping, and genomic structure of a novel X-chromosomal human gene (SMPX) encoding a small muscular protein. Hum. Genet. 105, 506–512 (1999).
Kemp, T. J. et al. Identification of a novel stretch-responsive skeletal muscle gene (Smpx). Genomics 72, 260–271 (2001).
Reverte, C. G., Ahearn, M. D. & Hake, L. E. CPEB degradation during Xenopus oocyte maturation requires a PEST domain and the 26S proteasome. Dev. Biol. 231, 447–458 (2001).
Schraders, M. et al. Next-generation sequencing identifies mutations of SMPX, which encodes the small muscle protein, X-linked, as a cause of progressive hearing impairment. Am. J. Hum. Genet. 88, 628–634 (2011).
Abdelfatah, N. et al. A novel deletion in SMPX causes a rare form of X-linked progressive hearing loss in two families due to a founder effect. Hum. Mutat. 34, 66–69 (2013).
Deng, Y. et al. A novel mutation in the SMPX gene associated with X-linked nonsyndromic sensorineural hearing loss in a Chinese family. J. Hum. Genet. 63, 723–730 (2018).
Gao, S. et al. Skewed X-chromosome inactivation and next-generation sequencing to identify a novel SMPX variants associated with X-linked hearing loss in a Chinese family. Int. J. Pediatr. Otorhinolaryngol. 113, 88–93 (2018).
Huebner, A. K. et al. Nonsense mutations in SMPX, encoding a protein responsive to physical force, result in X-chromosomal hearing loss. Am. J. Hum. Genet. 88, 621–627 (2011).
Niu, Z. et al. A novel splicing mutation in SMPX is linked to nonsyndromic progressive hearing loss. Int. J. Pediatr. Otorhinolaryngol. 104, 47–50 (2018).
Niu, Z. et al. A novel frameshift mutation of SMPX causes a rare form of X-linked nonsyndromic hearing loss in a Chinese family. PLOS One 12, e0178384 (2017).
Ghilardi, A. et al. Inner ear and muscle developmental defects in Smpx-deficient zebrafish embryos. Int. J. Mol. Sci. 22, 6497 (2021).
Lv, Y. et al. Whole-exome sequencing identifies a donor splice-site variant in SMPX that causes rare X-linked congenital deafness. Mol. Genet. Genomic Med. 7, e967 (2019).
del Castillo, I. et al. A novel locus for non-syndromic sensorineural deafness (DFN6) maps to chromosome Xp22. Hum. Mol. Genet. 5, 1383–1387 (1996).
Guo, Y. et al. A novel missense mutation in SMPX causes a rare form of X-linked postlingual sensorineural hearing loss in a Chinese family. Transl. Pediatr. 10, 378–387 (2021).
Yoon, H., Lee, D. J., Kim, M. H. & Bok, J. Identification of genes concordantly expressed with Atoh1 during inner ear development. Anat. Cell Biol. 44, 69–78 (2011).
Zhang, Y. et al. Rbm24a is necessary for hair cell development through regulating mRNA stability in zebrafish. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2020.604026 (2020).
Ghilardi, A., Diana, A., Prosperi, L. & Del Giacco, L. Expression pattern of the small muscle protein, X-linked (smpx) gene during zebrafish embryonic and larval developmental stages. Gene Expr. Patterns GEP 36, 119110 (2020).
Tu, H. et al. SMPX deficiency causes stereocilia degeneration and progressive hearing loss in CBA/CaJ mice. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2021.750023 (2021).
Thomas, E. D., Cruz, I. A., Hailey, D. W. & Raible, D. W. There and back again: Development and regeneration of the zebrafish lateral line system. Wiley Interdiscip. Rev. Dev. Biol. 4, 1–16 (2015).
López-Schier, H. & Hudspeth, A. J. Supernumerary neuromasts in the posterior lateral line of zebrafish lacking peripheral glia. Proc. Natl. Acad. Sci. 102, 1496–1501 (2005).
Nicolson, T. The genetics of hearing and balance in zebrafish. Annu. Rev. Genet. 39, 9–22 (2005).
Whitfield, T. T. Zebrafish as a model for hearing and deafness. J. Neurobiol. 53, 157–171 (2002).
Metcalfe, W. K., Kimmel, C. B. & Schabtach, E. Anatomy of the posterior lateral line system in young larvae of the zebrafish. J. Comp. Neurol. 233, 377–389 (1985).
Sarrazin, A. F. et al. Origin and early development of the posterior lateral line system of zebrafish. J. Neurosci. 30, 8234–8244 (2010).
Gompel, N. et al. Pattern formation in the lateral line of zebrafish. Mech. Dev. 105, 69–77 (2001).
Ramesh, T. et al. A genetic model of amyotrophic lateral sclerosis in zebrafish displays phenotypic hallmarks of motoneuron disease. Dis. Model. Mech. 3, 652–662 (2010).
Sarrazin, A. F. et al. Proneural gene requirement for hair cell differentiation in the zebrafish lateral line. Dev. Biol. 295, 534–545 (2006).
Lukasz, D., Beirl, A. & Kindt, K. Chronic neurotransmission increases the susceptibility of lateral-line hair cells to ototoxic insults. eLife 11, e77775 (2022).
Erickson, T. et al. Integration of Tmc1/2 into the mechanotransduction complex in zebrafish hair cells is regulated by Transmembrane O-methyltransferase (Tomt). eLife 6, e28474 (2017).
Haas, P. & Gilmour, D. Chemokine signaling mediates self-organizing tissue migration in the zebrafish lateral line. Dev. Cell 10, 673–680 (2006).
Pistocchi, A. et al. The zebrafish prospero homolog prox1 is required for mechanosensory hair cell differentiation and functionality in the lateral line. BMC Dev. Biol. 9, 58 (2009).
Kindt, K. S., Finch, G. & Nicolson, T. Kinocilia mediate mechanosensitivity in developing zebrafish hair cells. Dev. Cell 23, 329–341 (2012).
Kindt, K. S. & Sheets, L. Transmission disrupted: Modeling auditory synaptopathy in zebrafish. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2018.00114 (2018).
DalleNogare, D. & Chitnis, A. B. A framework for understanding morphogenesis and migration of the zebrafish posterior Lateral Line primordium. Mech. Dev. 148, 69–78 (2017).
Lush, M. E. et al. scRNA-Seq reveals distinct stem cell populations that drive hair cell regeneration after loss of Fgf and Notch signaling. eLife 8, e44431 (2019).
Olt, J., Johnson, S. L. & Marcotti, W. In vivo and in vitro biophysical properties of hair cells from the lateral line and inner ear of developing and adult zebrafish. J. Physiol. 592, 2041–2058 (2014).
Morthorst, S. K., Christensen, S. T. & Pedersen, L. B. Regulation of ciliary membrane protein trafficking and signalling by kinesin motor proteins. FEBS J. 285, 4535–4564 (2018).
Aman, A. & Piotrowski, T. Wnt/β-catenin and Fgf signaling control collective cell migration by restricting chemokine receptor expression. Dev. Cell 15, 749–761 (2008).
Nechiporuk, A. & Raible, D. W. FGF-dependent mechanosensory organ patterning in zebrafish. Science 320, 1774–1777 (2008).
Holmgren, M. & Sheets, L. Using the zebrafish lateral line to understand the roles of mitochondria in sensorineural hearing loss. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2020.628712 (2021).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Hoshijima, K. et al. Highly efficient methods for generating deletion mutations and F0 embryos that lack gene function in zebrafish. Dev. Cell 51, 645-657.e4 (2019).
Kroll, F. et al. A simple and effective F0 knockout method for rapid screening of behaviour and other complex phenotypes. eLife 10, e59683 (2021).
Meeker, N. D., Hutchinson, S. A., Ho, L. & Trede, N. S. Method for isolation of PCR-ready genomic DNA from zebrafish tissues. BioTechniques 43, 610–614 (2007).
Zhao, D. et al. Rapid and efficient cataract gene evaluation in F0 zebrafish using CRISPR-Cas9 ribonucleoprotein complexes. Methods 194, 37–47 (2021).
Acknowledgements
Work in L.D.G. lab and A.D. post-doc fellowship are supported by PNRR CN00000041-PNRR_CN3RNA_SPOKE1/SPOKE9 and Linea 2 intramural funds. We thank Dr. Miriam “Mrs.” Ascagni from Unitech NoLimits imaging facility for her priceless assistance and Darren Gilmour for the tg (cldnb:GFP) line. Figure 7 was generated with BioRender and modified by L.D.G. using Photoshop image-editing software.
Author information
Authors and Affiliations
Contributions
L.D.G., A.G., and A.D. conceived, designed, contributed, and analyzed data. A.D. performed the experiments. The manuscript was written by L.D.G. and A.D. with input from A.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diana, A., Ghilardi, A. & Del Giacco, L. Differentiation and functioning of the lateral line organ in zebrafish require Smpx activity. Sci Rep 14, 7862 (2024). https://doi.org/10.1038/s41598-024-58138-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-58138-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.