Abstract
White striping (WS) is a myopathy of growing concern to the turkey industry. It is rising in prevalence and has negative consequences for consumer acceptance and the functional properties of turkey meat. The objective of this study was to conduct a genome-wide association study (GWAS) and functional analysis on WS severity. Phenotypic data consisted of white striping scored on turkey breast fillets (N = 8422) by trained observers on a 0–3 scale (none to severe). Of the phenotyped birds, 4667 genotypic records were available using a proprietary 65 K single nucleotide polymorphism (SNP) chip. The SNP effects were estimated using a linear mixed model with a 30-SNP sliding window approach used to express the percentage genetic variance explained. Positional candidate genes were those located within 50 kb of the top 1% of SNP windows explaining the most genetic variance. Of the 95 positional candidate genes, seven were further classified as functional candidate genes because of their association with both a significant gene ontology and molecular function term. The results of the GWAS emphasize the polygenic nature of the trait with no specific genomic region contributing a large portion to the overall genetic variance. Significant pathways relating to growth, muscle development, collagen formation, circulatory system development, cell response to stimulus, and cytokine production were identified. These results help to support published biological associations between WS and hypoxia and oxidative stress and provide information that may be useful for future-omics studies in understanding the biological associations with WS development in turkeys.
Similar content being viewed by others
Introduction
Great improvements have been made in poultry growth, efficiency, and meat yield1. Strategic improvements in management, nutrition, and genetic selection, have led to turkey toms that can weigh over 20 kg at 20 weeks of age2. However, some negative consequences of this improvement in growth and production are becoming apparent. White striping (WS) is a growth-related myopathy that is of increasing interest to the poultry industry. This myopathy presents itself as varying degrees of white striations on the surface of the muscle running parallel to the muscle fibers3. White striping is highly prevalent and has recently been shown to be as high as 88% (including mild to severe cases) in a population of purebred turkeys and as much as 60% in other turkey populations4,5. This myopathy has known negative consequences for consumer acceptance, nutritional, and functional quality of the product3,6,7,8,9. These aspects make research into the biological mechanisms behind the condition and potential methods of prevention of importance to the turkey industry.
Although the amount of research conducted on WS in turkeys is limited, there is more known about the myopathy in broiler chickens. Several studies have been conducted in broilers that investigated WS microscopically and showed that affected breasts have necrotic muscle tissue and increased presence of inflammatory cells, connective tissue, and fat10,11,12. While the exact mechanism for development of WS is still unknown, one of the main mechanisms proposed is ischemia in the affected muscle13. With the magnitude and speed of growth in modern genotypes, the limits of supporting physiological systems such as the circulatory and cardiovascular might have been reached. A major consequence of selection for muscle growth is an increase in muscle fiber hypertrophy14,15,16. This hypertrophy can then lead to insufficient vascularization and reduced blood supply to the fast-growing muscles17,18,19. A restriction in the circulatory system can lead to changes in stem cell growth and the accumulation of metabolic byproducts, inducing oxidative stress likely leading to necrosis, and increases in hypoxic conditions potentially impairing muscle cell regeneration, ultimately leading to the development of WS.
This mechanism of WS development has been supported through various -omics studies in broiler chickens at the level of the transcriptome20,21,22, proteome23, and metabolome13. However, there is a lack of research in these areas for turkeys. Consequently, the objective of this study was to investigate the genomic architecture of WS in turkeys through the estimation of genomic heritability and execution of a genome-wide association study (GWAS) followed by functional analysis for detection of metabolic pathways and gene ontologies associated with the myopathy.
Results and discussion
Estimation of genetic parameters
Genetic and phenotypic correlations of WS with other economically important traits were published in Vanderhout et al.24. Heritability of WS was estimated to be 0.20 ± 0.022 and is the first published genomic heritability estimate for WS in turkeys. The addition of genomic data resulted in a 33% increase in estimated heritability compared to pedigree information alone24. The present estimate was found to be within the range of previously published estimates of heritability (observed scale) in broiler chickens of 0.18–0.5025,26,27. The moderate heritability estimated in the present study suggests that there is a presence of genetic factors influencing WS that could potentially be exploited in selecting birds for reduced WS severity, however, environmental factors can also influence most of the phenotypic variance observed in the population25. However, it is worth noting that the comparing the present heritability estimate with what is reported in the literature may be challenging due to the different species being studied (chickens vs. turkeys), different methods of scoring WS (e.g., different levels of scoring), different breeding goals, and different prevalences of WS in the given populations. Differences in trait prevalence are well known to influence heritability estimates when using linear models which estimate parameters on the observed scale. The prevalence of WS in the present study ranged from 84 to 92% which is substantially greater than what was observed for Bailey et al.25 (18.5–33.8%), Lake et al.27 (79%), and Alnahhas et al.26 (50%). Therefore, it is reasonable to expect variability among studies with regard to heritability estimates.
Significant SNP and positional candidate genes
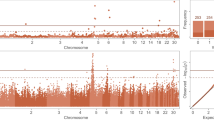
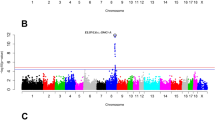
The percentage of genetic variance explained by each 30-SNP sliding window is presented in Fig. 1. Each window explained 0.05% of the genetic variance on average with no more than 1.00% of the variance being explained by any given window. This suggests that the inheritance of the trait is largely polygenic in nature. A total of 544 SNP windows were classified as significant (top 1% of variance explained) resulting in 95 positional candidate genes found within 50 kb upstream or downstream of these SNP. This distance has been suggested by Do et al.28 to be used when dealing with lower quality assemblies like that of the turkey. The positional candidate genes were located on Meleagris gallopavo autosomal chromosomes (MGA) 2 to 9, 11, 14, 19, 20, and 24. The 95 positional candidate genes were significantly associated (p < 0.05) with four KEGG metabolic pathways (Table 1) and 31 GO terms (21 BP, 3 CC, and 7 MF; Table 2). Positional candidate genes were further considered functional candidate genes (FCG) if they were associated with both a significant metabolic pathway and a significant GO term. Seven FCG were found and were involved mainly in the Wnt signaling pathway (NFATc1), RNA degradation (LSM6 and DHX36), and focal adhesion (COL6A3, FN1, VCL, and GRB2).
Due to WS being a growth-related myopathy, it is not surprising that several growth and muscle development related BP terms (GO:0003012 muscle system process, GO:0040007 growth, and GO:0061061 muscle structure development) were found to be significantly overrepresented (p < 0.05) by the 95 positional candidate genes. The large selection pressure placed on growth for economically significant muscle groups (i.e., pectoralis) has resulted in meat producing birds that are likely reaching the limit of supporting systems, such as the circulatory system. Thus, the proposed mechanism of WS development is primarily thought to be related to poor blood flow in the breast muscle leading to hypoxia, pressure on satellite cells, and oxidative stress13,21,29. The highly conserved Wnt signalling pathway, one of the four significant metabolic pathways, plays an important role in both embryonic development, where it regulates processes such as differentiation and cell proliferation, polarity, and migration, as well as post-natally, where it regulates tissue homeostasis and biological processes involved in many disorders and cancers30,31,32. The FCG associated with the Wnt signalling pathway, nuclear factor of activated T cells 1 (NFATc1), is found on MGA3 associated with the largest peak in variance explained. This gene has been shown to play a large role in cell cycle progression of human aortic smooth muscle cells33 and promoting the response to injury in arterial smooth muscle cells34. The effect of NFATc1 and the Wnt signalling pathway in the development and repair of the vascular system may be what leads to its significant relationship to WS. Some significant BP terms found in the present study were associated with FCG (including NFATc1, DHX36, and FN1), specifically GO:0009628 response to abiotic stimulus (p = 0.01) and GO:0072359 circulatory system development (p = 0.04), further supporting the relationship between hypoxia, oxidative stress, and WS.
Functional candidate genes collagen type VI alpha 3-chain (COL6A3) and fibronectin 1 (FN1) were also previously found to be significantly associated with WS in broiler chickens35 and differentially expressed between broiler chicken breasts affected versus not affected by WS36,37. The COL6A3 gene produces collagen found in the extracellular matrix of cells that make up skeletal muscles, and mutations in the gene are associated with muscle weakness, atrophy, and necrosis in humans38. The FN1 gene encodes a glycoprotein which plays a role in the creation of extracellular matrix structures during tissue repair and increases in the expression of this gene have been linked with Duchenne muscular dystrophy in humans39. Given the increase in fat and connective tissue that replaces damaged muscle tissue in affected breast muscles, the link between these two genes and WS is reasonable. Another gene of interest is cytosine and glycine rich protein 3 (CSRP3), a positional candidate gene found to be associated with several significant GO terms including 10 BP, one CC, and 1 MF. The CSRP3 gene has been previously shown to be upregulated in broiler chicken breasts affected with WS22,40. This gene encodes a muscle LIM protein and overexpression of such protein can promote muscle differentiation, regeneration, and structural repair of skeletal muscle41,42 further emphasizing the link between WS and muscle tissue damage.
The BP term, GO: 0001816 cytokine production, was found to be significantly overrepresented (p = 0.02) by the positional candidate genes in the present study, including three of the seven FCG (NFATc1, FN1, and DXH36). A microscopic characteristic consistently found in poultry breast tissue affected by WS is an elevated presence of inflammatory cells and cytokines10,43,44. Cytokines are small proteins that play a large role in immune response and inflammation and the elevated presence of these molecules in the muscle of affected breasts is symbolic of muscle cell injury45,46. Whether these genes, and subsequent production of cytokines, was upregulated in the affected breasts of the current study is unknown, however, the expression of inflammatory cytokine genes has been shown to increase with increasing severity of WS in broiler chickens44.
To the best of our knowledge, this study provides the first published estimate of genomic heritability of WS in turkeys and provides the first look into the genomic architecture of WS in turkeys by means of a GWAS and functional analysis. The heritability estimate of WS was found to be 0.20 ± 0.022, and results of the GWAS emphasize the polygenic nature of the trait with no specific genomic region contributing a large portion to the overall genetic variance. Results of the functional analysis identified four significant KEGG metabolic pathways, 31 significant GO terms (21 BP, 3 CC, and 7 MF) and seven functional candidate genes associated with WS. Overall, pathways relating to growth, muscle development, collagen formation, circulatory system development, cell response to stimulus, and cytokine production were highlighted. The results of the present study provide support for the oxidative stress and hypoxic theory of WS development. It should be noted that the WS phenotype was analyzed using a linear model which may reduce the statistical power when considering categorical traits (i.e., compared to a threshold model). Continued -omics research on the topic of WS in turkeys is recommended to further identify relationships between the myopathy and biological processes to identify improved prevention methods. For example, using a meta-GWAS approach to provide a comprehensive assessment of genetic factors influencing WS. Future research should also focus on developing methods of quantitatively scoring WS using technologies such as machine vision algorithms. Such measures would permit an increase in phenotypic measures increasing the power of future analyses.
Materials and methods
Animals
All protocols complied with the guidelines of the Canadian Council on Animal Care and were approved by the University of Guelph Animal Care Committee (AUP 3782). The study was conducted in accordance with relevant guidelines and regulations as well as the ARRIVE guidelines47. Adult male turkeys (20–24 weeks old) from three purebred genetic lines (A, B, and C) were processed over 44 weeks between July 2018 and November 2019. The genetic lines included a sire-line with selection focused on body weight, meat yield, and feed efficiency (line A), a dam-line that was selected primarily for body weight and reproductive traits (line B), and a dam-line selected mainly for reproductive traits (line C). Birds were reared under identical housing and management conditions as specified by the breeding company management guidelines (Hybrid Turkeys, 2020). During processing at a commercial poultry processing plant, birds were electrically stunned, exsanguinated, scalded, defeathered, and eviscerated before moving to the water chiller. Upon completion of the 24 h chilling period (40 min in 5 °C water, 1.5–2 h in 1–2 °C water, and remainder of time layered in ice), birds were deboned, and meat quality and breast muscle weights were measured.
Phenotype and genotype data
Summary statistics of the data are shown in Table 3. Deboned Pectoralis major muscles (N = 8422) were photographed (Hero 6, GoPro, San Mateo, CA, USA) approximately 24 h post-mortem. Photographs were taken using the normal focal length setting from approximately 40 cm above the surface of the breast. The photographs were randomly assigned to six observers who scored the breasts for WS using a 0–3 scoring scale adapted from a system developed in broiler chickens after testing the reliability of the system5,7. In brief, a score of 0 indicated no or minimal white striations whereas a score of 3 indicated the presence of thick white striations covering the breast. Genotypes were collected on 4667 birds using a proprietary 65 K single nucleotide polymorphism (SNP) array (65,000 SNP; Illumina, Inc.). PLINK software48 was used for quality control and SNP markers located on non-autosomal regions with minor allele frequency lower than 0.05, call rate lower than 90%, or significantly deviating from Hardy Weinberg proportions (p < 1 × 10–8) were removed. The quality control resulted in 54,407 markers retained for analysis.
Statistical analysis
A linear mixed model was used to estimate variance components through restricted maximum likelihood using the BLUPf90 family of programs49. The linear mixed model used can be described as follows:
where y is the vector of WS scores; b is a vector of fixed effects including genetic line (3 levels: A, B, and C), hatch week-year (58 levels), age at slaughter (7 levels; 141–163 days), and score observer (6 levels); a is a vector of additive genetic effects distributed as \(\mathbf{a}\sim N(0, \mathbf{H}{\sigma }_{a}^{2})\), where H is the combined pedigree-genomic relationship matrix as in Aguilar et al. (2010) constructed using the PREGSf90 program49. \({\sigma }_{a}^{2}\) is the additive genetic variance; e is the vector of residual effects which has a distribution of \(\mathbf{e}\sim N(0,{\sigma }_{e}^{2})\) where \({\sigma }_{e}^{2}\) is the residual variance; and X and Z are design matrices relating the observations to the fixed and random effects, respectively.
Estimates of SNP effects were derived from the estimated genomic breeding values (gEBV) following50, using a weighted genomic relationship matrix:
where \(\widehat{\mathbf{g}}\) is a vector of SNP marker effects; D is a diagonal matrix of weights for variances of SNPs; Z is a matrix relating genotype of each locus; and \({\widehat{\mathbf{u}}}_{{\text{g}}}\) is the vector of gEBV. Due to the proposed polygenic nature of WS and the relatively poor annotation of the turkey genome, a 30-SNP sliding window approach was utilized. This approach allows for accumulating the variance explained by each set of 30 adjacent SNP, which would lead to identify potential genomic regions associated with WS that may not be detected due to the low variance explained by single SNPs. These analyses were carried out using the BLUPf90 family of programs49.
Functional analysis
An arbitrary threshold for markers in the 99th percentile of variance explained were considered significant. Using the Turkey 5.1 assembly51, positional candidate genes within ± 50 kb of the significant SNP were retrieved using the Ensembl Genes database version 104 (https://useast.ensembl.org/Meleagris_gallopavo/Info/Index) implemented through the GALLO R package52. Gene ontology (GO) enrichment analysis including biological processes (BP), cellular components (CC), and molecular functions (MF) as well as metabolic pathway analysis using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database53 were performed on the positional candidate genes using the WebGestaltR R package54 and the Gallus gallus database.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Zuidhof, M. J., Schneider, B. L., Carney, V. L., Korver, D. R. & Robinson, F. E. Growth, efficiency, and yield of commercial broilers from 1957, 1978, and 2005. Poult. Sci. 93, 2970–2982 (2014).
Hiscock, H. M. et al. Describing the relationships among meat quality traits in domestic turkey (Meleagris gallopavo) populations. Poult. Sci. 101, 102055 (2022).
Barbut, S. Recent myopathies in broiler’s breast meat fillets. Worlds Poult. Sci. J. 75, 559–582 (2019).
Mudalal, S. Incidence of white striping and its effect on the quality traits of raw and processed Turkey breast meat. Food Sci. Anim. Resour. 39, 410–417 (2019).
Vanderhout, R. J. et al. Reliability of a white striping scoring system and description of white striping prevalence in purebred Turkey lines. Animals 12, 1–13 (2022).
Kuttappan, V. A. et al. Consumer acceptance of visual appearance of broiler breast meat with varying degrees of white striping. Poult. Sci. 91, 1240–1247 (2012).
Kuttappan, V. A., Hargis, B. M. & Owens, C. M. White striping and woody breast myopathies in the modern poultry industry: A review. Poult. Sci. 95, 2724–2733 (2016).
Soglia, F. et al. Effect of white striping on Turkey breast meat quality. Animal 12, 2198–2204 (2018).
de Carvalho, L. M., Ventanas, S., Olegario, L. S., Madruga, M. S. & Estévez, M. Consumers awareness of white-striping as a chicken breast myopathy affects their purchasing decision and emotional responses. Lwt 131, 109809 (2020).
Kuttappan, V. A. et al. Pathological changes associated with white striping in broiler breast muscles. Poult. Sci. 92, 331–338 (2013).
Russo, E. et al. Evaluation of White Striping prevalence and predisposing factors in broilers at slaughter. Poult. Sci. 94, 1843–1848 (2015).
Baldi, G. et al. Implications of white striping and spaghetti meat abnormalities on meat quality and histological features in broilers. Animal 12, 164–173 (2018).
Boerboom, G., Van Kempen, T., Navarro-Villa, A. & Pérez-Bonilla, A. Unraveling the cause of white striping in broilers using metabolomics. Poult. Sci. 97, 3977–3986 (2018).
Aberle, E. D. & Stewart, T. S. Growth of fiber types and apparent fiber number in skeletal muscle of broiler- and layer-type chickens. Growth 47, 135–144 (1983).
Remignon, H., Lefaucheur, L., Blum, J. C. & Ricard, F. H. Effects of divergent selection for body weight on three skeletal muscles characteristics in the chicken. Br. Poult. Sci. 35, 65–76 (1994).
MacRae, V. E., Mahon, M., Gilpin, S., Sandercock, D. A. & Mitchell, M. A. Skeletal muscle fibre growth and growth associated myopathy in the domestic chicken (Gallus domesticus). Br. Poult. Sci. 47, 264–272 (2006).
Sosnicki, A. A. & Wilson, B. W. Pathology of Turkey skeletal muscle: Implications for the poultry industry. Food Struct. 10, 317–326 (1991).
Velleman, S. G. Relationship of skeletal muscle development and growth to breast muscle myopathies: A review. Avian Dis. 59, 525–531. https://doi.org/10.1637/11223-063015-Review.1 (2015).
Kindlein, L. et al. Occurrence and severity of white striping in broilers until 50d of age fed with high and low-energy diets: Body weight, histopathological changes and meat quality. J. Vet. Sci. Technol. 8, 1–8 (2017).
Zambonelli, P. et al. Detection of differentially expressed genes in broiler pectoralis major muscle affected by White Striping—Wooden Breast myopathies. Poult. Sci. 95, 2771–2785 (2016).
Malila, Y. et al. Absolute expressions of hypoxia-inducible factor-1 alpha (HIF1A) transcript and the associated genes in chicken skeletal muscle with white striping and wooden breast myopathies. PLoS One 14, 1–23 (2019).
Marchesi, J. A. P. et al. Whole transcriptome analysis of the pectoralis major muscle reveals molecular mechanisms involved with white striping in broiler chickens. Poult. Sci. 98, 590–601 (2019).
Kuttappan, V. A. et al. Proteomic analysis reveals changes in carbohydrate and protein metabolism associated with broiler breast myopathy. Poult. Sci. 96, 2992–2999 (2017).
Vanderhout, R. J. et al. Genetic parameters of white striping and meat quality traits indicative of pale, soft, exudative meat in Turkeys (Meleagris gallopavo). Front. Genet. 13, 1–7 (2022).
Bailey, R. A., Watson, K. A., Bilgili, S. F. & Avendano, S. The genetic basis of pectoralis major myopathies in modern broiler chicken lines. Poult. Sci. 94, 2870–2879 (2015).
Alnahhas, N. et al. Genetic parameters of white striping in relation to body weight, carcass composition, and meat quality traits in two broiler lines divergently selected for the ultimate pH of the pectoralis major muscle. BMC Genet. 17(61), 1–9 (2016).
Lake, J. A., Dekkers, J. C. M. & Abasht, B. Genetic basis and identification of candidate genes for wooden breast and white striping in commercial broiler chickens. Sci. Rep. 11(6785), 1–13 (2021).
Do, D. N., Strathe, A. B., Ostersen, T., Pant, S. D. & Kadarmideen, H. N. Genome-wide association and pathway analysis of feed efficiency in pigs reveal candidate genes and pathways for residual feed intake. Front. Genet. 5, 1–10 (2014).
Velleman, S. G. Broiler breast muscle myopathies: Association with satellite cells. Poult. Sci. 102, 102917 (2023).
Clevers, H. Wnt/β-Catenin signaling in development and disease. Cell 127, 469–480 (2006).
Ackers, I. & Malgor, R. Interrelationship of canonical and non-canonical Wnt signalling pathways in chronic metabolic diseases. Diabetes Vasc. Dis. Res. 15, 3–13 (2018).
Noguchi, S., Saito, A. & Nagase, T. YAP/TAZ signaling as a molecular link between fibrosis and cancer. Int. J. Mol. Sci. 19, 1–23 (2018).
Karpurapu, M. et al. Cyclin D1 is a Bona fide target gene of NFATc1 and is sufficient in the mediation of injury-induced vascular wall remodeling. J. Biol. Chem. 285, 3510–3523 (2010).
Chow, W., Hou, G. & Bendeck, M. P. Glycogen synthase kinase 3β regulation of nuclear factor of activated T-cells isoform c1 in the vascular smooth muscle cell response to injury. Exp. Cell Res. 314, 2919–2929 (2008).
Pampouille, E. et al. Mapping QTL for white striping in relation to breast muscle yield and meat quality traits in broiler chickens. BMC Genom. 19, 1–14 (2018).
Pampouille, E. et al. Differential expression and co-expression gene network analyses reveal molecular mechanisms and candidate biomarkers involved in breast muscle myopathies in chicken. Sci. Rep. 9, 1–17 (2019).
Praud, C. et al. Molecular phenotyping of white striping and wooden breast myopathies in chicken. Front. Physiol. 11, 1–16 (2020).
Bertini, E. & Pepe, G. Collagen type VI and related disorders: Bethlem myopathy and Ullrich scleroatonic muscular dystrophy. Eur. J. Paediatr. Neurol. 6, 193–198 (2002).
Cynthia Martin, F. et al. Fibronectin is a serum biomarker for Duchenne muscular dystrophy. Proteom. Clin. Appl. 8, 269–278 (2014).
Marciano, C. M. M. et al. Differential expression of myogenic and calcium signaling-related genes in broilers affected with white striping. Front. Physiol. 12, 1–11 (2021).
Arber, S., Halder, G. & Caroni, P. Muscle LIM protein, a novel essential regulator of myogenesis, promotes myogenic differentiation. Cell 79, 221–231 (1994).
Barash, I. A., Mathew, L., Lahey, M., Greaser, M. L. & Lieber, R. L. Muscle LIM protein plays both structural and functional roles in skeletal muscle. Am. J. Physiol. Cell Physiol. 289, 1312–1320 (2005).
Carvalho, L. T. et al. Quality of turkeys breast meat affected by white striping myopathy. Poult. Sci. 100, 1–10 (2021).
Prisco, F. et al. Pathologic characterization of white striping myopathy in broiler chickens. Poult. Sci. 100, 1–14 (2021).
Prisk, V. & Huard, J. Muscle injuries and repair: The role of prostaglandins and inflammation. Histol. Histopathol. 18, 1243–1256 (2003).
Smith, C., Kruger, M. J., Smith, R. M. & Myburgh, K. H. The inflammatory response to skeletal muscle injury: Illuminating complexities. Sports Med. 38, 947–969 (2008).
Percie du Sert, N. et al. The ARRIVE guidelines 2.0: Updated guidelines for reporting animal research. J. Cereb. Blood Flow Metab. 40, 1769–1777 (2020).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Misztal, I. et al. Manual for BLUPF90 family of programs. Athens, USA 1–142 (2018).
Wang, H., Misztal, I., Aguilar, I., Legarra, A. & Muir, W. M. Genome-wide association mapping including phenotypes from relatives without genotypes. Genet. Res. 94, 73–83 (2012).
Dalloul, R. A., Zimin, A. V., Settlage, R. E., Kim, S. & Reed, K. M. Applying next-generation sequencing to solve poultry problems: Next-generation sequencing strategies for characterizing the turkey genome. Poult. Sci. 93, 479–484 (2014).
Fonseca, P. A. S., Suárez-Vega, A., Marras, G. & Cánovas, Á. GALLO: An R package for genomic annotation and integration of multiple data sources in livestock for positional candidate loci. GigaScience 9, 1–9 (2020).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Wang, J. & Liao, Y. WebGestaltR: Gene Set Analysis Toolkit WebGestaltR (2020).
Acknowledgements
This research was done in partial fulfillment of the requirements for completion of a Doctor of Philosophy degree by Ryley Vanderhout. The authors would like to gratefully acknowledge Jeff Mohr, Michelle Yahiro, Nienke van Staaveren, Heather Hiscock, Elizah McFarland, Jadelyn Appleby, and Bayode Makanjuola for assisting with data collection. The authors extend their gratitude to the managers and personnel of Hayter’s Farm (Dashwood, Ontario) and Hybrid Turkeys pedigree farm (Kitchener, Ontario) for collaborating on this study. This project was funded by the Government of Canada through Genome Canada and the Ontario Genomics Institute (OGI-133). This study was part of the project entitled “Application of genomic selection in turkeys for health, welfare, efficiency and production traits” funded by the government of Canada through the Genome Canada Genomic Application Partnership Program and administered by Ontario Genomics [recipients: B.J. Wood (Industry) and C.F. Baes (Academic)]. The authors would also like to acknowledge NSERC and Hybrid Turkeys for financial support.
Author information
Authors and Affiliations
Contributions
C.B., B.W., S.B., E.A., E.L., and R.V. conceived and designed the study. C.B. and B.W. secured funding for the study. E.L., E.A., and R.V. conducted the study. E.A. and R.V. analyzed the data. E.L. and R.V. wrote the original draft. C.B., B.W., S.B., E.L., E.A., and R.V. reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
BW was an employee of Hybrid Turkeys at the time of the study. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results. No other authors report a conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vanderhout, R.J., Abdalla, E.A., Leishman, E.M. et al. Genetic architecture of white striping in turkeys (Meleagris gallopavo). Sci Rep 14, 9007 (2024). https://doi.org/10.1038/s41598-024-59309-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-59309-8
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.