Abstract
PANoptosis plays a crucial role in cancer initiation and progression. However, the roles of PANoptosis-related genes (PARGs) in the prognosis and immune landscape of head and neck squamous cell carcinoma (HNSCC) remain unclear. Integrated bioinformatics analyses based on the data of HNSCC patients in the TCGA database were conducted. We extracted 48 PARGs expression profile and then conducted differentially expressed analysis, following building a Cox model to predict the survival of HNSCC patients. Subsequently, the relationships between the risk score, immune landscape, chemo-, and immune-therapy responses were analyzed, respectively. Moreover, we investigated the prognostic value, and further predicted the pathways influenced by PARGs. Finally, we identified the biological function of crucial PARGs. A total of 18 differentially expressed PARGs were identified in HNSCC, and a Cox model including CASP8, FADD, NLRP1, TNF, and ZBP1 was constructed, which showed that the risk score was associated with the prognosis as well as immune infiltration of HNSCC patients, and the risk score could be regarded as an independent biomarker. Additionally, patients with high-risk score might be an indicator of lymph node metastasis and advanced clinical stage. High-risk scores also contributed to the chemotherapy resistance and immune escape of HNSCC patients. In addition, FADD and ZBP1 played a crucial role in various cancer-related pathways, such as the MAPK, WNT, and MTOR signaling pathways. On the other hand, we suggested that FADD facilitated the progression and 5-fluorouracil (5-FU) resistance of HNSCC cells. A signature based on PANoptosis showed great predictive power for lymph node metastasis and advanced stage, suggesting that the risk score might be an independent prognostic biomarker for HNSCC. Meanwhile, FADD, identified as a prognostic biomarker, may represent an effective therapeutic target for HNSCC.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) is the seventh most common cancer type worldwide, with more than 850,000 new cases and 400,000 deaths per year1. HNSCC is characterized by chemoresistance, recurrence, lymph node and distant metastasis, which lead to an unsatisfactory prognosis2. Despite advances in therapeutic strategies, the 5-year overall survival rate is also unsatisfying3, being lower than 65% and lower than 40% in advanced stage cases4,5. Therefore, it is urgent to clarify the mechanism of HNSCC progression and thereby identify effective therapeutic targets.
As a widespread phenomenon in organisms, cell death is crucial for maintaining homeostasis. The imbalance between death and growth is the direct reason for tumorigenesis6. Programmed cell death (PCD), including pyroptosis, apoptosis, and necroptosis, is the mode of active cell death determined by genetic information. Recently, PANoptosis was identified as a new PCD pathway7, which is defined as an inflammatory PCD pathway regulated by the PANoptosome complex. PANoptosis contains the vital molecules and key features of traditional PCD pathways, which can crosstalk and influence their mutual relationship8, while the typical features of PANoptosis cannot be accounted for by any of these PCD pathways alone. PANoptosis has been reported to play a crucial role in tumor initiation and progression. For instance, the PANoptosis-Related lncRNA SNHG7 is involved in chemoresistance and metastasis in colon adenocarcinoma9. The characterization of PANoptosis patterns can also provide a roadmap for predicting survival and immunotherapy response10. Recently, the majority of studies have focused on PANoptosis to identify candidate biomarkers for various cancers11,12. For instance, PANoptosis-relevant subgroups could evaluate the prognosis and immune landscape of liver hepatocellular carcinoma (LIHC)11, suggesting that understanding PANoptosis might provide strategies for the clinical therapy of LIHC. However, few studies have focused on the contribution of PANoptosis to HNSCC prognostic value and therapy resistance. Recently. Feng Gao et al. indicated that the PANoptosis pattern could predict the prognosis and immunotherapy response of HNSCC patients13. Targeting PANoptosis might facilitate the development of more effective treatment approaches for HNSCC. Therefore, the discovery of relevant biomarkers of PANoptosis in HNSCC is crucial. A better understanding of the role of PANoptosis in HNSCC might be helpful to provide new insights into diagnosis and targeted therapy.
In this study, prognostic PANoptosis-related genes (PARGs) in HNSCC patients were identified by analyzing datasets obtained from The cancer genome atlas (TCGA). Furthermore, we investigated the prognostic value of PANoptosis by building a risk model, and the predictive power of the risk model was validated. Moreover, alterations in the apoptotic program are associated with the cancer response to drugs14. Hence, we explored the correlation between PANoptosis-related biomarkers and drug susceptibility. On the other hand, we analyzed the role of PANoptosis in immune infiltration and the immune response. Subsequently, we predicted the potential mechanism of PANoptosis-related biomarkers via gene set enrichment analysis (GSEA). Finally, we investigated the biological function of PANoptosis-related biomarkers. Taken together, this research may provide new insights into the diagnosis and targeted therapy of HNSCC.
Material and methods
Transcriptome data preparation
The transcriptome profile and relevant clinical information of HNSCC patients were downloaded from TCGA (https://portal.gdc.cancer.gov/) database. Cases without survival data were excluded. A total of 44 normal controls and 519 tumor cases were obtained. 48 PARGs were obtained from previous studies (Table S1)11,15,16,17, following the selection of differentially expressed PARGs (DEPARGs) with the cut-off criteria |log2 (FoldChange (FC))|> 1 and p-value < 0.05.
To identify the prognostic value of PARGs in HNSCC
According to the identified DEPARGs, we merged the gene expression level and survival data of each patient. Furthermore, the selected genes were enrolled in multivariate and stepwise Cox regression analyses to construct a risk model. In accordance with the risk formula, we calculated the risk score of each patient according to risk score model, and divided patients into high- and low-risk groups based on the risk score median. Kaplan–Meier curves along with long rank tests were used to identify the predictive power of risk model. Moreover, we used univariate and multivariate Cox regression analyses to explore whether the risk score could be regarded as an independent prognostic biomarker for HNSCC. The confidence interval was given with HR. Meanwhile, we identified the prognostic value of PARGs enrolled in risk model.
Tumor microenvironment (TME) analysis
According to genome expression, we calculated the immune cell levels via single sample gene set enrichment analysis (ssGSEA) in the R software “CIBERSORT” package18,19. We further explored the correlation between prognosis-related PARGs and immune cell levels. In addition, we predicted the immune cell and function scores of HNSCC patients in the R software packages “GVSA” and “GSEAbase”, and then investigated the difference in immune infiltration between the high- and low-risk groups. Meanwhile, we downloaded the stemness score based on DNA methylation (DNAss) and RNA (RNAss) from UCSC Xena (http://xena.ucsc.edu/), following exploration of the correlation between stemness score and PARGs.
Immunotherapy and drug therapy analysis
Tumor mutation burden (TMB) is regarded as a predictor of immunotherapeutic response. We obtained HNSCC patients’ TMB data from the UCSC Xena database (https://xena.ucsc.edu/) and further explored the correlation between TMB and prognosis-related PARGs. Furthermore, we explored the co-expression of PARGs and immune checkpoints (ICPs). We also downloaded the immunotherapy data of HNSCC patients from the TCIA database (https://tcia.at/home) and investigated the role of PARGs in HNSCC immunotherapy. Moreover, we obtained drug susceptibility data from the CellMiner database (https://discover.nci.nih.gov/cellminer/home.do) and further investigated the role of PANoptosis in drug resistance.
Gene set enrichment analysis (GSEA)
GSEA was used to predict the KEGG pathways affected by PANoptosis. Generally, prognosis-related PARGs were divided into high- and low-expression groups in accordance with the expression median. Subsequently, the genome expression file and group file were imported into GSEA software to screen KEGG pathways influenced by PARGs. The top 5 KEGG pathways were listed.
Cell culture and transfection
HNSCC cell lines HN12, SCC9, SCC15, Cal27, and human oral keratinocyte (HOK) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin at 37 °C with 5% CO2. All cell lines were obtained from ATCC. Small interfering RNAs (siRNAs) targeting Fas associated via death domain (FADD) were designed and synthesized. Cells were seeded in 6-well plates, and when the density reached 80%, siRNAs were transfected following the instructions of the Lipofectamine 3000 Kit. Total mRNAs of transfected cells were collected for analysis after 2 days. The siRNA sequences were as follows: Sense-GAGUCACUGAGAAUCUGGAAGAACA; Antisense-UGUUCUUCCAGAUUCUCAGUGACUC.
RNA isolation and qRT-PCR
The total RNA of cells was isolated according to the TRIZOL manufacturer’s instructions and further reversed transcribed to cDNA with a reverse transcription kit. The relative expression of targeted genes to GAPDH was evaluated by RT-qPCR. The primer sequences of the targeted genes are as follows: GAPDH-F: GGAGCGAGATCCC TCCAA AAT; GAPDH-R: GGCTGTTGTCATACTTCTCATGG. FADD-F: GTGGCT GACCT GGTACAAGAG; FADD-R: GGTAGATGCGTCTGAGTTCCAT.
Cell proliferation assay
Approximately 2 × 104 transfected cells with or without 5 μM 5-Fluorouracil (5-FU) were seeded in 12-well plates at 0 h. Subsequently, the total cells were detected at 24 h, 48 h, 72 h, and 96 h. Generally, the cell proliferation activity was evaluated by the cell numbers.
Transwell assay
Approximately 5 × 104 transfected cells were seeded in the transwell upper chamber with 200 µl serum-free medium, and 600 µl DMEM with 10% FBS was placed on the bottom. After 24 h, migrating cells were analyzed by staining with crystal violet. The invasion assay was done with 100 µl of 1:8 diluted Matrigel in the bottom of the upper chamber.
Statistical analysis
Statistical analysis was carried out in SPSS software (p < 0.05*; p < 0.01**; p < 0.001***; p < 0.0001****). Student’s t test, one- or two-way ANOVA and Kaplan–Meier curves along with long rank tests were used in present study.
Results
PARGs were related to HNSCC prognosis
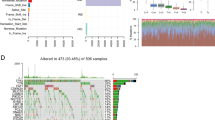
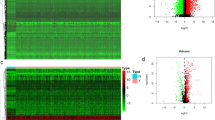
A total of 18 DEPARGs were identified in HNSCC (Fig. 1A,B, Table 1). To further investigate the prognostic value of these genes, we conducted Cox regression analysis and constructed a risk model including CASP8, FADD, NLRP1, TNF, and ZBP1 (Fig. 1C), suggesting that patients with high-risk levels had lower overall survival rate (Fig. 1D). Moreover, univariate Cox analysis indicated that the risk score according to PARGs might be a candidate prognostic factor for HNSCC (Fig. 1E). Furthermore, risk score was identified as an independent biomarker by multivariate Cox regression analysis (Fig. 1F). In addition, survival analysis combined with grade level could also predict HNSCC prognosis (Fig. 1G). Subsequently, we determined the prognostic value of variables in the risk model, and found that Z-DNA binding protein 1 (ZBP1) and FADD were candidate prognostic biomarkers for HNSCC (Fig. 1H).
PANoptosis played a crucial role in HNSCC prognosis. (A,B) Differentially expressed PANoptosis-related genes (DEPARGs) in HNSCC were visualized by volcano and heatmap, respectively. A total of 18 PARGs were differentially expressed in HNSCC. N: normal cases; T: tumor cases; blue: downregulated genes; red: upregulated genes. (C) A Cox model 5 DEPARGs including CAPS8, FADD, NLRP1, TNF and ZBP1 were built through regression analysis. (D) Risk score of each patient was calculated, following dividing patients into high- and low-risk group according to risk score median, which suggested that patients with high-risk score showed unfavorable overall survival. High and low means high-risk level and low-risk level, respectively. The number in the parenthesis is the size in each group. (E) Risk score according to DEPARGs was identified as a candidate prognostic biomarker for HNSCC by univariate Cox analysis. (F) Risk score based on DEPARGs was identified as an independent prognostic biomarker for HNSCC by multivariate Cox analysis. The confidence interval was given with HR. T: T classification in TNM system (tumor size); N: N classification in TNM system (lymph node metastasis); Stage: clinical stage for HNSCC samples in TCGA database; Grade: differentiation grade for HNSCC samples in TCGA database. (G) Survival analysis in line with stratification analysis of Grade level, suggesting that risk score might be a predictor for differentiation grade. High and low means high-risk level and low-risk level, respectively. (H) Survival analysis according to FADD and ZBP1 expression in TCGA database. The number in the parenthesis is the size in each group.
We further investigated the protein expression of ZBP1 and FADD in the Human Protein Atlas (HPA) database. FADD protein level was overexpressed (Fig. S1A), while ZBP1 protein level was nearly not expressed in tumor tissues (Fig. S1B). According to the mRNA expression level in TCGA, FADD was related to lymph node metastasis, and ZBP1 might play a crucial role in tumor size as well as clinical stage (Fig. S1C). The risk score in line with PARGs was also regarded as an unfavorable indicator for lymph node metastasis and advanced stage (Fig. S1D). Taken together, these results showed that PARGs might play a vital role in HNSCC progression and prognosis.
PARGs were related to immune infiltration of HNSCC
To further explore the role of PANoptosis in HNSCC, we analyzed the correlation between prognostic PARGs and HNSCC TME. We showed that the risk score according to the PARGs expression level was positively related to the stemness score (Fig. 2A), indicating that PARGs might play a critical role in the initiation of HNSCC. In addition, patients with high-risk scores tended to have lower immune cell scores, particularly NK cells and TILs (Fig. 2B, Table S2). Meanwhile, the immune response in high-risk level group was inhibited, in particular Type-I-IFN-response (Fig. 2B). FADD was identified to play a vital role in NK cell as well as TIL scores. Meanwhile, high expression of FADD might inhibit cell Cytolytic-activity (Fig. 2C). Although ZBP1 was overexpressed in tumor tissues, high expression of ZBP1 was positively related to the immune response (Fig. 2D). These results showed that PARGs were related to the immune infiltration of HNSCC.
PANoptosis related to immune infiltration of HNSCC. (A) Heatmap showing the relationship between risk score and cancer cell stemness score according to DNA methylation (DNAss) and RNA (RNAss) in HNSCC. (B) Box diagram displaying the correlation between risk score and immune cell scores, and between risk score and immune function scores in HNSCC. (C) Box diagram displaying the correlation between FADD expression and immune cell scores, and between FADD expression and immune function scores in HNSCC. (D) Difference analysis between ZBP1 expression and immune cell scores, and between ZBP1 expression and immune function scores in HNSCC.
ZBP1 enhanced the immunotherapy response of HNSCC
We further investigated the role of PARGs in HNSCC immunotherapy. The risk score was negatively associated with TMB, an indicator of immunotherapy response (Fig. 3A), suggesting that patients with high-risk score had an unsatisfactory immunotherapy response. The prognostic biomarker FADD was also positively, while ZBP1 was negatively related to TMB level (Fig. 3B,C), indicating that PANoptosis might play a crucial role in immunotherapy resistance. Furthermore, we showed that the expression of FADD and ZBP1 was correlated with majority of immune checkpoints (Fig. 3D). Moreover, high ZBP1 expression showed better response to immunotherapy (Fig. 3E), while FADD had no effect on HNSCC immunotherapy (Fig. 3F).
ZBP1 enhanced immunotherapy response. (A–C) Correlation analysis between risk score according to DEPARGs, FADD, ZBP1 and TMB, respectively. (D) Co-expression analysis of FADD, ZBP1 expression and immune checkpoints expression. (E,F) Violin plots displaying the effect of ZBP1 and FADD on the immunotherapy response of HNSCC patients. The immune cell proportion score (IPS) concerning immunotherapy data including ctla4-negative-pd1-neg, ctla4-negative-pd1-pos, ctla4-pos1-neg, and ctla4-pos1-pd1-pos were obtained from TCIA database. High expression of ZBP1 showed better immunotherapy response, while FADD had no effect on HNSCC immunotherapy on basis of TCIA database.
PARGs were related to drug therapy
To explore the role of PARGs in HNSCC drug therapy, we performed a correlation analysis between drug susceptibility and prognostic biomarkers. Patients with high expression of FADD tended to develop chemotherapy resistance, such as resistance to fluorouracil (Fig. 4A, Table 2). ZBP1 enhanced the response to drug therapy (Fig. 4B, Table 3). These results suggested that PANoptosis might play a significant role in the response of HNSCC to chemotherapy.
FADD and ZBP1 contributed to dysregulation of HNSCC-related pathways
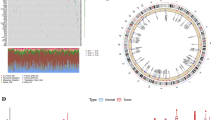
To predict the potential mechanism affected by PANoptosis, we further conducted GSEA to screen KEGG pathways influenced by FADD and ZBP1 (Fig. 5A). FADD overexpression was associated with the activation of several cancer-related pathways, such as the MAPK and MTOR signaling pathways. Meanwhile, FADD contributed to cell cycle, focal adhesion and inhibition of immune response (Fig. 5B, Fig. S2A), suggesting the vital role of FADD in HNSCC progression. ZBP1 might promote apoptosis and immune process. Upregulation of ZBP1 impaired the activity of cancer-related pathways, such as Hedgehog and WNT signaling pathways (Fig. 5C, Fig. S2B). These results suggested that FADD and ZBP1 might be the crucial regulators in these HNSCC-related pathways.
FADD and ZBP1 contributed to dysregulation of various cancer-associated signaling pathways. (A) The detailed information of GSEA results. (B) GSEA suggesting the role of FADD overexpression in various cancer-related pathways, such as focal adhesion, MAPK signaling pathways and cell cycle. (C) The correlation between ZBP1 expression and various KEGG pathways activities including apoptosis, immune response and Glutathione metabolism.
Silence of FADD inhibited HNSCC proliferation, migration, invasion and enhance susceptibility to 5-FU
We further explored the biological function of prognostic biomarkers. We found that FADD was overexpressed in HNSCC cell lines (Fig. S3), while ZBP1 was not detected (Data not shown). Furthermore, we showed that knockdown of FADD inhibited HNSCC cell proliferation, and inhibition of FADD might elevate the susceptibility of HNSCC cells to 5-FU (Fig. 6A,B). Moreover, silencing FADD impaired the migration and invasion abilities of cancer cells (Fig. 6C,D). These results suggested that FADD might promote HNSCC progression and showed promise as a candidate therapeutic target to enhance chemotherapy.
Knockdown of FADD inhibited the progression and enhanced susceptibility to 5-FU of HNSCC cell. (A,B) Line diagram displaying the role of FADD knockdown in HNSCC cells proliferation and susceptibility to 5-FU. Student’s T test (SCC15 p = 0.0059; SCC9 p = 0.0048) and Two-way ANOVA (SCC15 siNC vs. siFADD p < 0.0001, siNC vs. siNC + 5-FU p < 0.0001, siFADD vs. siFADD + 5-FU p < 0.0001, siNC + 5-FU vs. siFADD + 5-FU p = 0.001; SCC9 siNC vs. siFADD p < 0.0001, siNC vs. siNC + 5-FU p < 0.0001, siFADD vs. siFADD + 5-FU p < 0.0001, siNC + 5-FU vs. siFADD + 5-FU p < 0.0001) were used to statistically test the efficiency of FADD knockdown and cell proliferation assays, respectively. (C,D) Transwell assay showing the contribution of FADD in HNSCC cell migration and invasion ability after downregulation of FADD. (SCC15 Migration p = 0.0037, Invasion p = 0.0005; SCC9 Migration p = 0.0015, Invasion p = 0.0037). Student’s T test was used to conduct statistical tests for migration and invasion ability.
Discussion
As a newly identified PCD mode, PANoptosis contributes to anti-tumor immunity by regulating the enrichment of immune cells and then promoting the death of cancer cells20. Aberrant PANoptosis status played a crucial role in the therapy and prognosis of various malignancies. In esophageal cancer, PANoptosis mediated by sulconazole could induce oxidative stress and impair aerobic glycolysis to enhance radiosensitivity21. Xiong et al. identified that PANoptosis might be an important indicator for efficacy of immunotherapy and chemotherapy in hepatocellular carcinoma (HCC)22. Wei et al. also showed that PANoptosis was associated with HCC overall survival, immune infiltration, and immunotherapy response23, suggesting that PANoptosis showed promise to be a therapeutic target. The dysregulation of cell proliferation and death plays a crucial role in the tumorigenesis of HNSCC, which is characterized by therapy resistance and infiltration of immune cells, contributing to unfavorable prognosis24,25. Currently, the therapy response and immune status of HNSCC patients have been revealed in accordance with various forms of gene sets related to different forms of cell death26. For instance, Huang et al. suggested that a signature based on ferroptosis-associated gene showed great predictive power as an independent indicator of prognosis, and patients at low-risk may benefit from immunotherapy26. Recently, Gao et al. demonstrated the effectiveness of PANoptosis associated molecular clustering and prognostic features in predicting the prognosis and immune landscape of HNSCC13. Therefore, targeting PANoptosis might enhance the progress of more effective treatment strategies for HNSCC immune- and chemotherapy. A better understanding of the role of PANoptosis in HNSCC might represent an effective therapeutic target for HNSCC in clinics.
In this study, 18 PARGs were identified as differentially expressed genes in HNSCC, and five genes, including FADD, CASP8, NLRP1, TNF, and ZBP1, were enrolled in the prognostic model. The risk-score might be regarded as an independent biomarker for predicting the prognosis of HNSCC by univariate and multivariate Cox regression analysis, as well as a predictor of lymph node metastasis and advanced clinical stage. Risk-score was positively related to TMB level, which suggested that these five genes might be candidate immunotherapeutic targets for HNSCC. Moreover, we showed that risk score was negatively related to 14 immune cell scores, especially B cells, T cells, NK cells and TILs. Meanwhile, the dysregulation of PANoptosis might inhibit the immune process. Taken together, PANoptosis might play a crucial role in the prognosis and immune infiltration of HNSCC. We further found that FADD and ZBP1 were associated with the prognosis of HNSCC patients, and contributed to immune infiltration. ZBP1, as an innate sensor, was associated with inflammasome activation, inflammation, and cell death27. Recently, ZBP1 was identified to act as a crucial regulator in multiple PCD pathways28,29. Rajendra Karki et al. demonstrated that downregulation of ZBP1 could promote tumorigenesis by suppressing PANoptosis and impairing immune response30. In addition, ZBP1 plays a role in the immune response via mediating type-I interferon production27,31, suggesting that ZBP1 is fundamental to informing therapeutic strategy. Fu et al. demonstrated the critical role of ZBP1 in inducing tumor-associated proteins32. Upregulation of ZBP1 could enhance PANoptosis via inducing TNF and IFN-γ and then inhibit tumorigenesis33, while the immune response and cell death were impaired when ZBP1 was downregulated30. On the other hand, inhibition of ZBP1 expression might increase the migration of ovarian cancer, and impair fisetin-induced apoptosis34. Deletion of ZBP1 blocked tumor necroptosis during tumor development and inhibited breast cancer metastasis, suggesting that ZBP1 is the key regulator of tumor necroptosis and provides a potential therapeutic target for controlling tumor metastasis35. In HNSCC, we suggested that upregulation of ZBP1 enhanced the immunotherapy response of HNSCC patients in the TCGA database. However, ZBP1 protein expression was nearly not detected on basis of HPA database. Possibly, there are various post-transcriptional regulation of ZBP1 in HNSCC. Currently, ZBP1 could interact with various mRNAs, and play a key role in the post-transcriptional regulation of gene expression36. Generally, mRNAs could be directly interacted with the KH34 domain of ZBP1, leading to post-transcription and then regulating their corresponding protein expression37. Numerous studies have focused on the role of ZBP1 in regulating post-transcription. Nevertheless, fewer studies have discussed the post-transcription of ZBP1. Hence, elaborating on the post-transcription of ZBP1 might help reveal the potential mechanism of HNSCC tumorigenesis and progression. As an adaptor molecule of PANoptosis and apoptosis, FADD can interact with various cell surface receptors and mediate cell apoptotic signals. In addition, FADD could be recruited by various molecules through its C-terminal death domain, such as TNFRSF6/Fas-receptor, tumor necrosis factor receptor, TNFRSF25, thereby participating in the death signaling initiated by these receptors38. Moreover, FADD can regulate various signaling pathways related to cell death, such as caspase activation via the extrinsic apoptotic signaling pathway, ligand-dependent caspase activation and regulation of necroptotic cell death39,40. In addition, FADD could prevent the spontaneous expression of ZBP1, and then inhibit necroptosis of cancer cells41. Dysregulation of FADD contributes to the tumorigenesis and progression of various cancers42. For instance, FADD protected cancer cells from drug-induced apoptosis in pancreatic cancer, while impairment of FADD expression sensitized drug-resistant cells to Adriamycin®-mediated apoptosis43 suggesting that FADD was a crucial chemotherapeutic target. In penile squamous cell carcinoma (PSCC), overexpression of FADD was an adjunct biomarker with poor prognosis in PSCC, and might be regarded as a tumor immune environment regulator44. In lung cancer, Wei et al. suggested that FADD is one of the prominent risk factors. Inhibition of FADD could reduce cancer cell proliferation, and knockdown of FADD elevated the apoptosis and pyroptosis of cancer cells45. We also showed that FADD contributed to unfavorable overall survival and immune infiltration of HNSCC, suggesting that FADD might be a crucial oncogene in squamous cell carcinoma. Studies showed that amplification of 11q13.3 is related to increased metastasis in HNSCC46, while FADD was considered to be a driver gene in amplification of the chromosomal 11q13.3 region, thereby contributing to oncogene expression, such as cyclin D147. Pattje WJ et al. suggested that FADD was related to regional and distant metastases, and might be visualized as a therapy target to reduce the risk of distant metastases48. A recent Meta-Analysis also identified the crucial role of FADD in the prognosis of HNSCC49. We found that FADD was related to poor overall survival, and enhanced chemotherapy resistance. Wei et al. also suggested that FADD as the critical regulator of PANoptosis was related to therapy resistance in HCC23. On the other hand, we identified that high FADD expression might inhibit the immune process via regulating NK cells and TIL, which contributed to immune escape of cancer cells. Meanwhile, we conducted biological experiments to verify that FADD enhanced the proliferation, migration and invasion of HNSCC cancer cells, indicating that FADD showed promising diagnostic and prognostic significance in HNSCC. In conclusion, these results demonstrated that FADD was expected to be a therapeutic target.
In summary, we conducted a PANoptosis-based molecular signature, and identified that PANoptosis played a crucial role in predicting prognosis, TMB, and guiding drug and immunotherapy. Moreover, we suggested that FADD might be a potential therapeutic target for HNSCC. These findings in this study might enhance the understanding of PANoptosis and help to screen more effective treatment strategies for HNSCC.
Limitations
Our study may have some vital clinical significance, but there are some limitations. Firstly, we fail to investigate the mechanism of FADD-mediated PANoptosis in HNSCC. Then, the role of FADD in HNSCC should be explored in vivo. Thirdly, the roles of FADD in immune escape and therapy resistance need to be determined. Finally, the post-transcriptional regulation and the role of ZBP1 overexpression in HNSCC also need to be further explored.
Data availability
All the data in this study were obtained from public database. All of the data presented in this study could be obtained from corresponding author.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Vallianou, N. G. et al. Immunotherapy in head and neck cancer: Where do we stand?. Curr. Oncol. Rep. 25, 897–912. https://doi.org/10.1007/s11912-023-01425-1 (2023).
Svider, P. F. et al. Head and neck cancer. Otolaryngol. Head Neck Surg. 156, 10–13. https://doi.org/10.1177/0194599816674672 (2017).
Spoerl, S. et al. Lymphatic and vascular invasion in oral squamous cell carcinoma: Implications for recurrence and survival in a population-based cohort study. Oral Oncol. 111, 105009. https://doi.org/10.1016/j.oraloncology.2020.105009 (2020).
Arun, I. et al. Lymph node characteristics and their prognostic significance in oral squamous cell carcinoma. Head Neck 43, 520–533. https://doi.org/10.1002/hed.26499 (2021).
Mohammad, R. M. et al. Broad targeting of resistance to apoptosis in cancer. Semin. Cancer Biol. 35, S78–S103. https://doi.org/10.1016/j.semcancer.2015.03.001 (2015).
Samir, P., Malireddi, R. K. S. & Kanneganti, T. D. The PANoptosome: A deadly protein complex driving pyroptosis, apoptosis, and necroptosis (PANoptosis). Front. Cell. Infect. Microbiol. 10, 238. https://doi.org/10.3389/fcimb.2020.00238 (2020).
Wang, Y. et al. Single cell analysis of PANoptosome cell death complexes through an expansion microscopy method. Cell. Mol. Life Sci. 79, 531. https://doi.org/10.1007/s00018-022-04564-z (2022).
Huang, J. et al. Analysis of PANoptosis-related LncRNA-miRNA-mRNA network reveals LncRNA SNHG7 involved in chemo-resistance in colon adenocarcinoma. Front. Oncol. 12, 888105. https://doi.org/10.3389/fonc.2022.888105 (2022).
Pan, H., Pan, J., Li, P. & Gao, J. Characterization of PANoptosis patterns predicts survival and immunotherapy response in gastric cancer. Clin. Immunol. 238, 109019. https://doi.org/10.1016/j.clim.2022.109019 (2022).
Zhang, Z. et al. Identification of PANoptosis-relevant subgroups to evaluate the prognosis and immune landscape of patients with liver hepatocellular carcinoma. Front. Cell Dev. Biol. 11, 1210456. https://doi.org/10.3389/fcell.2023.1210456 (2023).
Wang, Y. et al. A novel defined PANoptosis-related miRNA signature for predicting the prognosis and immune characteristics in clear cell renal cell carcinoma: A miRNA signature for the prognosis of ccRCC. Int. J. Mol. Sci. https://doi.org/10.3390/ijms24119392 (2023).
Gao, F. et al. A PANoptosis pattern to predict prognosis and immunotherapy response in head and neck squamous cell carcinoma. Heliyon 10, e27162. https://doi.org/10.1016/j.heliyon.2024.e27162 (2024).
Jiang, M. et al. Caspase-8: A key protein of cross-talk signal way in “PANoptosis” in cancer. Int. J. Cancer 149, 1408–1420. https://doi.org/10.1002/ijc.33698 (2021).
Yi, X. et al. Construction of PANoptosis signature: Novel target discovery for prostate cancer immunotherapy. Mol. Ther. Nucleic Acids 33, 376–390. https://doi.org/10.1016/j.omtn.2023.07.010 (2023).
Zhang, C. et al. Identifying prognostic genes related PANoptosis in lung adenocarcinoma and developing prediction model based on bioinformatics analysis. Sci. Rep. 13, 17956. https://doi.org/10.1038/s41598-023-45005-6 (2023).
Zhuang, L., Sun, Q., Huang, S., Hu, L. & Chen, Q. A comprehensive analysis of PANoptosome to prognosis and immunotherapy response in pan-cancer. Sci. Rep. 13, 3877. https://doi.org/10.1038/s41598-023-30934-z (2023).
Liu, J., Liu, Y., Yang, C., Liu, J. & Hao, J. Comprehensive analysis for the immune related biomarkers of platinum-based chemotherapy in ovarian cancer. Transl. Oncol. 37, 101762. https://doi.org/10.1016/j.tranon.2023.101762 (2023).
Xu, S. et al. Cuproptosis-related signature for clinical prognosis and immunotherapy sensitivity in hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 149, 12249–12263. https://doi.org/10.1007/s00432-023-05099-x (2023).
Liu, J. et al. Programmed cell death tunes tumor immunity. Front. Immunol. 13, 847345. https://doi.org/10.3389/fimmu.2022.847345 (2022).
Liu, L. X. et al. Sulconazole induces PANoptosis by triggering oxidative stress and inhibiting glycolysis to increase radiosensitivity in esophageal cancer. Mol. Cell Proteomics 22, 100551. https://doi.org/10.1016/j.mcpro.2023.100551 (2023).
Xiong, X., Song, Q., Jing, M. & Yan, W. Identification of PANoptosis-based prognostic signature for predicting efficacy of immunotherapy and chemotherapy in hepatocellular carcinoma. Genet. Res. (Camb.) 2023, 6879022. https://doi.org/10.1155/2023/6879022 (2023).
Wei, Y. et al. Robust analysis of a novel PANoptosis-related prognostic gene signature model for hepatocellular carcinoma immune infiltration and therapeutic response. Sci. Rep. 13, 14519. https://doi.org/10.1038/s41598-023-41670-9 (2023).
Chaves, P. et al. Preclinical models in head and neck squamous cell carcinoma. Br. J. Cancer 128, 1819–1827. https://doi.org/10.1038/s41416-023-02186-1 (2023).
Sunga, G. M., Hartgerink, J., Sikora, A. G. & Young, S. Enhancement of immunotherapies in head and neck cancers using biomaterial-based treatment strategies. Tissue Eng. C Methods 29, 257–275. https://doi.org/10.1089/ten.TEC.2023.0090 (2023).
Huang, Z. et al. Identification of a ferroptosis-associated gene signature and the related therapeutic targets in head and neck squamous carcinoma. Int. Immunopharmacol. 102, 108431. https://doi.org/10.1016/j.intimp.2021.108431 (2022).
Karki, R. & Kanneganti, T. D. ADAR1 and ZBP1 in innate immunity, cell death, and disease. Trends Immunol. 44, 201–216. https://doi.org/10.1016/j.it.2023.01.001 (2023).
Thapa, R. J. et al. DAI senses influenza A virus genomic RNA and activates RIPK3-dependent cell death. Cell Host Microbe 20, 674–681. https://doi.org/10.1016/j.chom.2016.09.014 (2016).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aag2045 (2016).
Karki, R. et al. ADAR1 restricts ZBP1-mediated immune response and PANoptosis to promote tumorigenesis. Cell Rep. 37, 109858. https://doi.org/10.1016/j.celrep.2021.109858 (2021).
Chen, X. Y. et al. ZBP1-mediated necroptosis: Mechanisms and therapeutic implications. Molecules https://doi.org/10.3390/molecules28010052 (2022).
Fu, Y. et al. Cloning of DLM-1, a novel gene that is up-regulated in activated macrophages, using RNA differential display. Gene 240, 157–163. https://doi.org/10.1016/s0378-1119(99)00419-9 (1999).
Malireddi, R. K. S. et al. Inflammatory cell death, PANoptosis, mediated by cytokines in diverse cancer lineages inhibits tumor growth. Immunohorizons 5, 568–580. https://doi.org/10.4049/immunohorizons.2100059 (2021).
Liu, Y., Cao, H., Zhao, Y., Shan, L. & Lan, S. Fisetin-induced cell death in human ovarian cancer cell lines via zbp1-mediated necroptosis. J. Ovarian Res. 15, 57. https://doi.org/10.1186/s13048-022-00984-4 (2022).
Baik, J. Y. et al. ZBP1 not RIPK1 mediates tumor necroptosis in breast cancer. Nat. Commun. 12, 2666. https://doi.org/10.1038/s41467-021-23004-3 (2021).
Doyle, M. & Kiebler, M. A. A zipcode unzipped. Genes Dev. 26, 110–113. https://doi.org/10.1101/gad.184945.111 (2012).
Wang, G. et al. IMP1 suppresses breast tumor growth and metastasis through the regulation of its target mRNAs. Oncotarget 7, 15690–15702. https://doi.org/10.18632/oncotarget.7464 (2016).
Chinnaiyan, A. M., O’Rourke, K., Tewari, M. & Dixit, V. M. FADD, a novel death domain-containing protein, interacts with the death domain of Fas and initiates apoptosis. Cell 81, 505–512. https://doi.org/10.1016/0092-8674(95)90071-3 (1995).
Ashkenazi, A. & Dixit, V. M. Death receptors: Signaling and modulation. Science 281, 1305–1308. https://doi.org/10.1126/science.281.5381.1305 (1998).
Galluzzi, L., Kepp, O. & Kroemer, G. FADD: An endogenous inhibitor of RIP3-driven regulated necrosis. Cell Res. 21, 1383–1385. https://doi.org/10.1038/cr.2011.147 (2011).
Rodriguez, D. A. et al. Caspase-8 and FADD prevent spontaneous ZBP1 expression and necroptosis. Proc. Natl. Acad. Sci. U. S. A. 119, e2207240119. https://doi.org/10.1073/pnas.2207240119 (2022).
Liu, Y., Li, X., Zhou, X., Wang, J. & Ao, X. FADD as a key molecular player in cancer progression. Mol. Med. 28, 132. https://doi.org/10.1186/s10020-022-00560-y (2022).
Zhang, R. et al. The role of FADD in pancreatic cancer cell proliferation and drug resistance. Oncol. Lett. 13, 1899–1904. https://doi.org/10.3892/ol.2017.5636 (2017).
Xue, T. et al. Prognostic significance and immune correlates of FADD in penile squamous cell carcinoma. Virchows Arch. 482, 869–878. https://doi.org/10.1007/s00428-023-03514-9 (2023).
Wei, S., Chen, Z., Ling, X., Zhang, W. & Jiang, L. Comprehensive analysis illustrating the role of PANoptosis-related genes in lung cancer based on bioinformatic algorithms and experiments. Front. Pharmacol. 14, 1115221. https://doi.org/10.3389/fphar.2023.1115221 (2023).
Schuuring, E. The involvement of the chromosome 11q13 region in human malignancies: Cyclin D1 and EMS1 are two new candidate oncogenes–a review. Gene 159, 83–96. https://doi.org/10.1016/0378-1119(94)00562-7 (1995).
Gibcus, J. H. et al. Amplicon mapping and expression profiling identify the Fas-associated death domain gene as a new driver in the 11q13.3 amplicon in laryngeal/pharyngeal cancer. Clin. Cancer Res. 13, 6257–6266. https://doi.org/10.1158/1078-0432.Ccr-07-1247 (2007).
Pattje, W. J. et al. FADD expression is associated with regional and distant metastasis in squamous cell carcinoma of the head and neck. Histopathology 63, 263–270. https://doi.org/10.1111/his.12174 (2013).
González-Moles, M. et al. Prognostic and clinicopathological significance of FADD upregulation in head and neck squamous cell carcinoma: A systematic review and meta-analysis. Cancers (Basel) https://doi.org/10.3390/cancers12092393 (2020).
Author information
Authors and Affiliations
Contributions
Conceptualization: Li Q; methodology, Yang P; software, Huang GZ; validation, Yang P and Huang GZ; formal analysis, Li YL; investigation, Yu L; data curation, Yin ZL; writing—original draft preparation, Yang P and Huang GZ; writing—review and editing, Yang P, Huang GZ and Li Q; supervision, Li Q. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, P., Huang, G., Li, Y. et al. Identification of PANoptosis-related biomarkers and analysis of prognostic values in head and neck squamous cell carcinoma. Sci Rep 14, 9824 (2024). https://doi.org/10.1038/s41598-024-60441-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60441-8
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.