Abstract
Viruses are crucial for regulating deep-sea microbial communities and biogeochemical cycles. However, their roles are still less characterized in deep-sea holobionts. Bathymodioline mussels are endemic species inhabiting cold seeps and harboring endosymbionts in gill epithelial cells for nutrition. This study unveiled a diverse array of viruses in the gill tissues of Gigantidas platifrons mussels and analyzed the viral metagenome and transcriptome from the gill tissues of Gigantidas platifrons mussels collected from a cold seep in the South Sea. The mussel gills contained various viruses including Baculoviridae, Rountreeviridae, Myoviridae and Siphovirdae, but the active viromes were Myoviridae, Siphoviridae, and Podoviridae belonging to the order Caudovirales. The overall viral community structure showed significant variation among environments with different methane concentrations. Transcriptome analysis indicated high expression of viral structural genes, integrase, and restriction endonuclease genes in a high methane concentration environment, suggesting frequent virus infection and replication. Furthermore, two viruses (GP-phage-contig14 and GP-phage-contig72) interacted with Gigantidas platifrons methanotrophic gill symbionts (bathymodiolin mussels host intracellular methanotrophic Gammaproteobacteria in their gills), showing high expression levels, and have huge different expression in different methane concentrations. Additionally, single-stranded DNA viruses may play a potential auxiliary role in the virus–host interaction using indirect bioinformatics methods. Moreover, the Cro and DNA methylase genes had phylogenetic similarity between the virus and Gigantidas platifrons methanotrophic gill symbionts. This study also explored a variety of viruses in the gill tissues of Gigantidas platifrons and revealed that bacteria interacted with the viruses during the symbiosis with Gigantidas platifrons. This study provides fundamental insights into the interplay of microorganisms within Gigantidas platifrons mussels in deep sea.
Similar content being viewed by others
Introduction
Deep sea represents an extreme and inhospitable environment characterized by darkness, hypoxia, high hydrostatic pressure, and low temperatures, resulting in generally low species1,2,3. However, chemosynthetic environments, including hydrothermal vents and cold seeps, serve as energy hotspots on the seafloor, supporting some of the most unique ecosystems on Earth4. These environments are created by the discharge of reduced fluids from the seafloor, often enriched with methane, sulfide, hydrogen, and iron II5. These chemicals drive primary production by chemosynthetic microorganisms. Additionally, macrofauna in hydrothermal vents and cold seeps must adapt to the highly toxic chemical environment by acquiring chemosynthetic microbes as symbionts6. For example, sponges from the Campos Basin in Southeastern Brazil have endosymbiotic Nitrosopumilaceae7. Thus far, more than 600 macrofaunal species have been identified in these extremely chemosynthetic environments8, with bivalve species, especially mussels, dominating the biomass in these ecosystems9. Bathymodiolin mussels harbor endosymbiotic methanotrophs in their gills and derive the vast majority of their nutrition from these symbionts10,11. Recent evidence suggests that deep-sea animal holobionts contain a diversity of phages, which could potentially influence animal-bacterial symbioses12,13. These intimate symbiotic relationships, common and crucial in vent and seep ecosystems, contribute to the formation of holobionts as adaptations to extreme environments.
Chemosynthetic ecosystems harbor a wide diversity of archaea and bacteria, which play crucial roles in hydrocarbon metabolism14,15. These microbial populations drive various biological processes including sulfate reduction, sulfur oxidation, denitrification, metal reduction, and methanogenesis within the seabed16,17. Viruses have also been observed in water, sediment, and invertebrates of chemosynthetic ecosystems18,19,20,21. Investigations from seven cold seep sediment sites have reported a broad range of archaeal and bacterial viruses, such as those for Bathyarchaeota, Methanomicrobia, Thaumarchaeota, Bipolaricaulota, Coatesbacteria, and Sumerlaeota14. Additionally, viruses of the Myoviridae, Siphoviridae, Podoviridae, and Microviridae families have been observed in various invertebrates, such as tube worms, sponges, and mussels13,22,23,24. These viruses serve as key agents in natural ecosystems through a range of interactions with their microbial hosts. These interactions manifest in lytic and lysogenic infections of viruses. Lytic infection leads to the reproduction of viral progeny, the lysis of host cells, and the release of cellular dissolved organic matter, thereby influencing microbial community diversity and organic carbon and nutrient turnover25. By contrast, temperate viruses replicate alongside their hosts, forming a mutualistic relationship during lysogenic infection25. They can reprogram host metabolism by encoding a number of putative auxiliary metabolic genes, affecting processes like sulfur oxidation26, phosphate metabolism27, pentose phosphate pathway28, nitrogen metabolism29,30, purine and pyrimidine metabolism pathways31, and others. With successive investigations into marine viruses, it has become clear that virus–host interactions are more complex than previously thought, and viruses play essential roles in microbe-driven ocean biogeochemical processes.
Bathymodiolin mussels represent one of the dominant megafaunal taxa found at hydrothermal vents and cold seeps, which provides the microorganisms with optimal growth conditions, necessary metabolites, and shelter32,33. They host endosymbiotic bacteria within gill epithelial cells, known as bacteriocytes, and derive energy and nutrients through the oxidation of reducing substances, including methane, hydrogen sulfide, thiosulfate, and hydrogen34. Previous studies have primarily focused on the roles of chemoautotrophic bacteria in symbiotic systems, overlooking the presence of viruses. Recent studies, however, have revealed the presence and abundance of viruses, and their interactions with hosts in chemosynthetic ecosystems14. Knowledge of the ecological roles of viruses in the deep sea has been constrained by challenges in sampling and extracting viral particles (virions)35. In recent years, advancements in sequencing and bioinformatics have enabled the analysis of viruses obtained from metagenomes sequenced without prior virion separation. These methods have significantly advanced viral ecology, from the discovery of novel viruses to understanding their global distribution. In this study, we utilized high-throughput metagenomic and transcriptome sequencing to characterize the viral community composition (only bacterial viruses) and diversity in Gigantidas platifrons. We focused on analyzing viral expression, as well as the interactions between viruses and bacteria, to explore the microbial interactions of G. platifrons mussels in the deep sea.
Results and discussion
Relative abundance and composition of viruses of G. platifrons
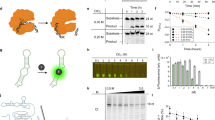
Metagenome and transcriptome data from nine G. platifrons samples collected from a cold seep in the South China Sea, in the western Pacific Ocean, were used to identify viromes and active viromes. Kraken 2 identified a total of 86 viral families (Supplementary Table S1). The main viral communities in G. platifrons were Baculoviridae, Rountreeviridae, Myoviridae, and Siphovirdae in both DNA and RNA sequencing results, but the active viromes were Myoviridae, Siphovirdae, and Podoviridae belonging to the order Caudovirales (Fig. 1a and b). Although the abundance of viruses varied greatly among different individuals, the species composition of the active viral community remained consistent. This diversity in viral abundance was primarily attributed to samples collected from different environments, suggesting that viral abundance exhibits specificity in various environments and among individuals, while the species composition of the viral community remains relatively uniform (environmental parameters of the samples listed in Table 1). In our study, we specifically discuss the uniformity of the species composition of the activity viral community in G. platifrons. Within the Caudovirales order, which includes the families Siphoviridae, Myoviridae, and Podoviridae, Siphoviridae, and Myoviridae were the dominant viral communities in seawater, with variations observed in different seawater areas. Notably, the relative abundance of Podoviridae was consistently low in all samples. Conversely, sponge metagenome analysis revealed a higher relative abundance of Podoviridae in the Crane site and Swan site36. The second-largest viral communities across all samples belonged to the Rountreeviridae family within the Caudovirales order and the Baculoviridae family within the Lefavirales order, ranging from 1.72 to 21.04% (Supplementary Table S1). The differences in viral communities and diversity in G. platifrons may be attributed to variations in deep-sea regions and the bacterial host species within organisms.
Viral contigs extracted from the metagenome of G. platifrons
Metagenomic data were used to assemble reads and identify viral contigs in our study. In total, 1459 contigs were identified from the metagenome using the VirSorter2 program, with a viral hit max_score of ≥ 50 (Fig. 1)37. The most contigs range from 1000 to 5000 bp, and 5000–10,000 bp and longer than 10,000 bp are 118 and 39, respectively (Fig. 2a). The longest contig measured 98,882 bp, while the average contig length was 12,905 bp. The viral contigs were categorized into two types of DNA: double-stranded DNA (dsDNA) and single-stranded DNA (ssDNA), constituting 45.23 and 54.76% of the total contigs, respectively (Fig. 2b). Among the 1459 viral contigs, 89 exceeded 5000 bp (Supplementary S2) according to the screening criteria. Additionally, 35 provirus contigs were identified, flanked by host genes on one or both sides (Supplementary Table S2). The viral contigs were classified into two categories: lysis and lysogeny, with lysogenetic viral contigs containing lysogenic genes such as integrase, excision enzyme, and Cro/CI repressor. Our study identified 29 lysogenetic viruses and 60 lysis viruses (Supplementary Table S2).
To investigate the community classification of viral contigs in G. platifrons from the cold seep, we constructed a gene-sharing network using vConTACT2. A total of 435 viral clusters (VCs) with family-level taxonomy were identified (Supplementary Table S3). Among the 89 viral contigs we studied, 12 clustered with RefSeq virus genomes, implying that these viral contigs can be assumed to belong to the same virus genera as that of the corresponding RefSeq genome. The majority of viral contigs showed no similarity to RefSeq virus genomes, suggesting they may cluster together as novel VCs or as classified as outliers and singletons38,39. Compared with the prokaryotic viruses in the RefSeq viral database (V85) (The reference database of vConTACT2 including ProkaryoticViralRefSeq85-ICTV, ProkaryoticViralRefSeq85, ProkaryoticViralRefSeq88, ProkaryoticViralRefSeq94, ProkaryoticViralRefSeq97, ProkaryoticViralRefSeq201, ArchaeaViralRefSeq85, ArchaeaViralRefSeq94, ArchaeaViralRefSeq97, and ArchaeaViralRefSeq201). We selected the viral RefSeq85 as the reference database following Li’s study14), the viral contigs were highly novel and variant. The network of viral contigs clustered with Myoviridae (3 sequences), Siphoviridae (4 sequences), Podoviridae (2 sequences), and Chaseviridae (3 sequences; Fig. 3). Myoviridae, Siphoviridae, and Podoviridae were common and prevalent virus populations in the ocean, existing widely in seawater, sediment, and marine invertebrates40,41,42. Myoviridae and Siphoviridae were observed in mussels, clams, tubeworms, snails, and sponges from deep-sea hydrothermal vents and cold seeps13. GP-phage-contig14 belongs to Myoviridae and shares similar genes with Enterobacteria phage, which is a virus associated with gut bacteria (Supplementary Table S3). The viral contigs filtered from snail holobionts in hydrothermal vents also connect to Enterobacteria phages13. Similarly, there also found phage contigs identified as the Microvirus genus of Enterobacteria phages in methane seeps43. This suggests that similar bacterial families may be associated with the same phage families shared by different animal species or environments13.
Gene expression of virus in G. platifrons
Transcripts from 50 genes of 47 different viruses, including both dsDNA and ssDNA types, were detected (Supplementary Table S4, Fig. 4). Among the predicted transcriptional genes were viral structural genes, integrase, restriction endonuclease, hydrolases, reductases, and other metabolically important enzymes. Interestingly, the integrase and restriction endonuclease genes were all from ssDNA, while most of the viral structural genes were from dsDNA (RNA phages were not considered). We analyzed the differential expression of viral transcriptomes in different environments; individuals from different environments showed significant variations in active viral expression (Fig. 4). In G. platifrons-1–3, which collected from an environment from low methane concentrations (up to 407 ppm), three genes involved in cell division were highly expressed: Fic/DOC family, FtsK/SpoIIIE family, and peptidase family M23 genes. In G. platifrons-7–9, which was collected from an environment with high methane concentrations (up to 4580 ppm), viral structural genes, integrase, and restriction endonuclease genes were highly expressed, indicating that the virus was more active in high methane concentrations. Additionally, aldose 1-epimerase, a key enzyme in carbohydrate metabolism, and enoyl-CoA hydratase/isomerase, an enzyme crucial in the fatty acid β-oxidative metabolic pathway, were highly expressed. Integrase and restriction endonuclease genes also exhibited high expression, suggesting that these two genes may serve as signature genes for virus integration into the bacterial genome. The results suggest the potential for gene exchange between viruses and bacteria in G. platifrons. Because of the nine individuals collected from different methane concentrations, the activity of viral composition had huge difference. It indicated that methane concentration is also critical for virus expression in G. platifrons.
Expression analyses of viruses of G. platifrons in nine different individuals. mRNA data were obtained from the transcriptome of gill tissues for the generation of heatmaps. The color scale in the heatmap indicates expression values; blue color indicates low transcript abundance, and red color indicates high transcript abundance. The red circles represent viral structural genes, and green triangles represent integrase and restriction endonuclease genes. The highlighted squares represent highly expressed genes.
Virus–host interaction
To investigate virus–host interactions in G. platifrons and understand the role of viruses in controlling host population dynamics and metabolic function, especially in relation to endosymbiotic bacteria, we employed CHERRY for species-level host prediction. We identified four interactions between viral contigs and prokaryotic genomes resulting from various types of interactions (e.g., protein organization information between viruses, sequence similarity between viruses and prokaryotes, and CRISPR signals). Four viral contigs (GP-phage-contig14, GP-phage-contig22, GP-phage-contig30, and GP-phage-contig67) were predicted to infect specific hosts, while four were linked to hosts from different prokaryotic bacteria. This result aligns with the common perception and previous findings that viral infections tend to be host-specific with a narrow host range14,44,45. The predicted hosts include Methyloprofundus sediment, Bathymodiolus platifrons methanotrophic gill symbiont, Helicobacter pylori, Campylobacter jejuni, Planktothrix agardhii, and Leuconostoc pseudomesenteroides (Fig. 5). Most viruses were linked to the methanotrophic symbiont in G. platifrons, which is a methane-oxidizing bacterium belonging to Gammaproteobacteria and plays a major role in the removal of the greenhouse gas methane from the biosphere. This result can be attributed to two main factors: the predominance of methanotrophic symbionts in the gill of G. platifrons and the similarity in virus community structure within the same tissue across different environments. GP-phage-contig14 and GP-phage-contig30, which specifically infected B. platifrons methanotrophic gill symbiont, indicate that virus–host interactions may be influenced by the environment.
Predicted virus–host interaction. (a) Predicted virus–host network in G. platifrons. MOB is shown in red ovals, G. platifrons methanotrophic gill symbiont, which is a methane-oxidizing bacterium. The number in pink ovals represents other viral hosts listed beside. (b) Genome domains of the G. platifrons symbiont and the B. platifrons symbiont (GCF_002189065.1); (c) phylogenetic tree of viral contigs. There were 1000 bootstrap sub-replicates, and bootstrap values < 50 are not shown. The viral genome contigs of G. platifrons are shown in bold.
In conjunction with the analysis of viral transcripts, we observed high expression of GP-phage-contig14 and GP-phage-contig72 (k141_1921954||full and k141_6385074||full) based on our data that found in G. platifrons and B. platifrons methanotrophic gill symbiont genomes (Fig. 5b) shown in Fig. 4. To investigate the interaction between the methanotrophic symbiont in G. platifrons and viruses, we annotated the genome sequences of G. platifrons and B. platifrons methanotrophic gill symbiont (GCF_002189065.1; Fig. 5b) and revealed the two virus (GP-phage-contig14 and GP-phage-contig72) activity in the two hosts. We found that viral genes had inserted into the host genomes, and these insertions were random and unstable. Additionally, the CRISPR/Cas and AbiEii domains (AbiE is a type IV toxin-antitoxin system, and AbiEii toxin is a putative nucleotidyltransferase containing a C-terminal domain) found adjacent to the insertion sites in the genome of G. platifrons methanotrophic gill symbiont, indicating the presence of multiple strategies in bacterial resist phage infection. In viral contigs that interacted with the host and exhibited transcript expression, the significant presence of ssDNA viruses was observed (G. platifrons methanotrophic gill symbiont contigs shown in Fig. 5b). Based on all the findings using the indirect bioinformatics methods, it is mere surmise that the integrase and restriction enzyme genes belonging to ssDNA may assist the insertion of viral genes (GP-phage-contig14 and GP-phage-contig72), which belong to dsDNA, into the host genome. As expected, more evidence is needed to confirm this surmise. Furthermore, there is substantial evidence indicating that interactions between viruses themselves are widespread46,47,48,49. When multiple viruses co-infect a single host, they can establish indirect interactions that alter host susceptibility, modify or suppress interferon response, or affect immune cell activation50, thereby avoiding superinfection51. A single virus infecting two or more hosts open the possibility of contact-based transfer of viral particles and/or genomes between or among syntrophic microbes, even those in different domains52. This could drive the rate of adaptive evolution of symbionts in different environments. The viral insertion element in the host genome serves two functions: assisting metabolism or defunctionalization.
Phylogenetic analysis was employed to investigate the homologous relationships among the viruses interacting with hosts, as shown in Fig. 5c. The results suggest that these viruses belong to the Gammaproteobacteria phage group. They can be clustered into two groups based on their host differences. Specifically, four viruses (GP-phage-contig14, GP-phage-contig30, GP-phage-contig72, and GP-phage-contig76) whose hosts were methane-oxidizing bacteria clustered with Vibrio and Enterobacteria phage, while one (GP-phage-contig78) clustered with Mannheimia phage belonging to the Pasteurellales phage. GP-phage-contig14, belonging to Myoviridae, showed a greater similarity to Enterobacteria phage during amino acid blast analysis with the NCBI database. This suggests that the host specificity is a distinctive feature of these viruses. However, the viral contigs were incomplete, preventing a comprehensive investigation of host functions. The identified function domains mainly involved infecting and regulating DNA transcription and replication. Previous studies have defined such incomplete viruses as defective interfering particles (DIPs)53. These particles exhibit critical absences in the viral genome, which can interfere with standard virus replication during coinfection54. However, it is important to note that we cannot conclusively confirm that the incomplete viral contigs in our study are indeed DIPs, as further evidence is required.
Phylogenetic similarity between the virus and G. platifrons methanotrophic gill symbionts
Viruses selectively integrate certain bacterial genes when infecting phages (bacterial viruses) lyse the bacterial chromosome, a process akin to horizontal gene transfer55. These viruses carry virulence genes as an integral part of their genomes, and these transferred genes may be highly expressed when the viruses infect other bacterial hosts and initiate replication during a lytic cycle. In our study, both Cro and DNA methylase genes were found in viral contigs and bacterial genomes, indicating a potential horizontal gene transfer. To validate our hypothesis, a phylogenetic tree was constructed incorporating both viral and bacterial genes. The analysis revealed that the genes for Cro and DNA methylase had indeed phylogenetic similarity between GP-phage-contig72 and the B. platifrons methanotrophic gill symbiont (Fig. 6). Notably, these genes play a pivotal role in the lytic/lysogenic system. Transcriptomic analysis further demonstrated the expression of the DNA methylase gene in GP-phage-contig72 (Supplementary Table S2), affirming the utilization of the lytic/lysogenic system by the virus GP-Phage-contig72.
Phylogenetic analysis of Cro and DNA methylase genes of viruses and bacteria. (a) Phylogenetic tree of the Cro gene. (b) Phylogenetic tree of the DNA methylase gene. There were 1000 bootstrap sub-replicates, and bootstrap values < 50 are not shown. The viral contigs of G. platifrons are shown in bold.
Conclusions
In the deep-sea cold seep environment, viruses, bacteria, and G. platifrons must collaborate to adapt to the extreme conditions. We conducted an analysis of the viral metagenome and transcriptome extracted from G. platifrons, revealing a diverse viral community within the organism, including Baculoviridae, Rountreeviridae, Myoviridae, and Siphovirdae, but the active viromes were Myoviridae, Siphoviridae, and Podoviridae belonging to the Caudovirales order, with significant variations in abundance among individuals. The high expression levels of integrase, restriction endonuclease, and viral structural genes suggest an active virus lysis process. The activity of virus was affected by different methane concentrations. This led to an interesting surmise: ssDNA and dsDNA virus appeared to assist each other in infecting or lysing the host. Our findings provide additional evidence for the study of interactions between viruses and bacteria within G. platifrons, shedding light on the evolution of gene transfers between viruses and hosts. This serves as a foundational basis for further investigations into the role of microorganisms in G. platifrons.
Methods
Sample collection
G. platifrons specimens were collected from a cold seep in the South China Sea, located in the western Pacific Ocean (22°06.919′N and 119°17.140′E), at a depth of 1119 m, using the remotely operated vehicle Faxian aboard the research vessel Kexue in 2019. Gill tissues from nine individuals were stored at − 80 °C for subsequent DNA/RNA extraction. Of these nine mussels, three were collected from a seepage region with a high methane concentration, three were collected from a mussel bed with a moderate methane concentration, and the remaining three from regions with low methane levels. The mussels were promptly fixed with the RNAsafer stabilizer reagent (Omega Bio-Tek, Norcross, GA, USA) in situ and transferred to the deck within 1 h for use for extracting total DNA from the gill tissues.
Metagenome sequencing and data from G. platifrons analysis
Total DNA was extracted from gill tissue samples of G. platifrons using the cetyl trimethylammonium bromide method56. The degree of DNA degradation and potential contamination was monitored on 1% agarose gels. DNA concentration was measured using the Qubit® dsDNA Assay Kit in a Qubit® 2.0 Fluorometer (Life Technologies, CA, USA). DNA samples with optical density values between 1.8 and 2.0 and a concentration above 1 µg were used to construct the library.
For DNA sample preparations, 1 µg of DNA per sample was used as input material. Sequencing libraries were generated using the NEBNext® Ultra DNA Library Prep Kit for Illumina (NEB, USA) following the manufacturer's recommendations. Index codes were added to attribute sequences to each sample. Briefly, the DNA sample was sonicated to a size of 350 bp, followed by end-polishing, A-tailing, and ligation with full-length adaptors for Illumina sequencing, followed by polymerase chain reaction (PCR) amplification. PCR products were then purified using the AMPure XP system, and libraries were analyzed for size distribution using an Agilent 2100 Bioanalyzer and quantified using real-time PCR.
The index-coded samples were clustered using a cBot Cluster Generation System according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina HiSeq platform, generating paired-end reads.
The original raw data from the nine individuals obtained from the Illumina HiSeq platform were cleaned using FastQC (v0.12.1) and Trimmomatic (v0.39) with custom parameters (ILLUMINACLTP: TruSeq3-PE.fa:2:30:10 SLIDINGWINDOW:4:15 LEADING:3 TRAILING:3 MINLEN:40 HEADCROP:12)57 for quality filtering (read length of 150 bp). Bowtie2 and samtools were then used to remove G. platifrons sequences58,59, and the G. platifrons genome was used to build the index database. The microbial sequence was obtained after removing the alignment of G. platifrons sequences. The trimmed reads of microorganisms in G. platifrons were assembled using Megahit with custom parameters (--k-min 21 --k-max 141 --k-step 10)60,61. The coverage of the assembled contigs was calculated with BBmap and Samtools59,62. The quality of the assembled contigs was evaluated using Quast63. Viral contigs were identified using VirSorter2 with default parameters, and the statistics of the virus sequences and viral type was performed using Excel (Fig. 2). The VirSorter2 database stores genetic information from viruses, which includes NCBI RefSeq, GenBank, PFAM, KEGG64,65, UniProt, EggNOG, and son on. The key criterion was the presence of viral hallmark genes (Table 2)33. The criteria for screening viral contigs were as follows: (1) viral genes > 0; (2) viral genes = 0 and (host gene = 0 or Virsorter2 viral hit max_score ≥ 0.95 or hallmark > 2); (3) manual check for those not falling under (1) and (2), with viral genes = 0 and host genes = 1 and length of ≥ 10 kb; (4) discarding the rest66. CheckV was used to evaluate the quality of viral metagenome contigs, including the identification of host contamination of proviruses, estimation of genomic fragment integrity, and identification of closed viral genomes65. Prodigal (version 2.6.3) was used to predict the coding sequences (CDS) of the viral genomes extracted from VirSorter231. Predicted CDS were then annotated using HMMER3 (version 3.3) against a custom comprehensive viral HMM database, which includes Xfams (as described in the “Methods” Section) and viral protein families from the JGI Earth’s virome project67,68.
RNA-Seq and data of virus of the transcriptome from G. platifrons analyses
Transcriptome sequencing was performed using Novogene (Tianjin, China) as described previously. Briefly, gill tissues from mussels were subjected to RNA extraction using TRIzol reagent (Invitrogen, MA, USA). RNA quantification, mRNA purification, cDNA synthesis, adaptor ligation, cDNA library construction, and sequencing using the Illumina HiSeq 2500 platform with paired-end reads were conducted using Novogene (Tianjin, China).
Abundance of viral transcriptome reads was computed using Bracken, which assigns taxonomy labels using Kraken 2 software with default parameters to estimate the number of viral reads present in each sample69,70. The viral taxonomy was based on alignment against the NCBI RefSeq database.
The trimmed reads of the G. platifrons transcriptome were obtained through a quality check using FastQC, followed by trimming using Trimmomatic. The G. platifrons sequences were then removed using Bowtie and samtools. The resulting trimmed reads were assembled de novo into a transcript using the paired-end method with Trinity71. Trinity consists of three software modules: Inchworm, Chrysalis, and Butterfly. In the first step in Trinity, the Inchworm module assembles the RNA-seq data into longer, gapless unique sequences called contigs. Next, chrysalis clusters the Inchworm contigs, representing the full transcriptional complexity for a given gene (or sets of genes that share common sequences). Chrysalis then partitions the full-read set among these clusters. Finally, Butterfly processes the individual graphs in parallel, tracing the paths that reads and pairs of reads take within the graph. It ultimately reports full-length transcripts for alternatively spliced isoforms and separates transcripts corresponding to paralogous genes. The resulting sequences are referred to as unigenes.
Abundance estimation was calculated using the expectation–maximization method72. The trimmed mean of M values method was employed to normalize the abundance73. Differentially expressed unigenes in the samples of G. platifrons virus from areas with different methane concentrations were identified using edgeR with a log2 fold change (Log2FC) > 0.5 (upregulated) or Log2FC < − 0.5 (downregulated) and false discovery rate (FDR) < 0.0574.
Phage protein clustering and taxonomic comparisons
Diamond was used to map the predicted ORFs against the NCBI Viral RefSeq database75. The Markov clustering algorithm (MCL) was used to group viral protein clusters (PCs), and ClusterONE was used to format the VCs76,77. The similarities between viral genomes were calculated using a one-tailed P value37. These steps were all conducted by vConTACT2 with default parameters (--rel-mode ‘Diamond’ --proteins-fp test_data/proteins.csv --db 'ProkaryoticViralRefSeq94-Merged' --pcs-mode MCL --vcs-mode ClusterONE)78. The viral protein network formed from vConTACT2 was visualized with Cytoscape software (v.3.6.0; http://cytoscape.org/), where nodes represent genomes and edges connect significantly similar viral genomes37,79.
Host prediction of viral contigs
Virus–host connections were predicted using the CHERRY method, considering various prediction types including protein organization, CRISPR, sequence similarity, and k-mer frequency79. MCL was used to construct PC, with reference viral genome proteins downloaded from NCBI RefSeq. Prodigal was used for gene finding and protein translation of query contigs67. Diamond was used to blastp the viral proteins against NCBI RefSeq with an e-value of 1e−5, followed by clustering proteins with an inflation value of 2.0 through MCL76. Prediction on the species level used pretrained parameters (--len 5000 –model pretrain –topk 1). The virus–host connection network was prediction in CHERRY78 also visualized in Cytoscape software (v.3.6.0; http://cytoscape.org/), with nodes representing host genomes and edges connecting significantly similar viral genomes80. The viral contigs and the G. platifrons methanotrophic gill symbiont genome annotated using TrinotateFFAM and KEGG database and visualized using IBS 1.0.3 software (Fig. 5b).
Phylogenetic analysis
To study the homologous relationship between viruses interacting with hosts, a neighbor-joining tree was created for viruses and bacteria contigs from the NCBI database using MEGA v1181. Bootstrap sub-replicates were set to 1000, and bootstrap values less than 50 were not obtained.
Data availability
All data needed to evaluate the conclusions in the paper are presented in the paper and/or the supplementary materials. All metagenome and transcriptome sequencing data have been deposited with links to BioProject accession number PRJNA1040436 in the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/).
References
Hourdez, S. & Lallier, F. H. Adaptations to hypoxia in hydrothermal-vent and cold-seep invertebrates. Rev. Environ. Sci. Biotechnol. 6(1–3), 143 (2007).
Dong, X. et al. Thermogenic hydrocarbon biodegradation by diverse depth-stratified microbial populations at a Scotian Basin cold seep. Nat. Commun. 11(1), 1–14 (2020).
Orphan, V. J., House, C. H., Hinrichs, K. U., McKeegan, K. D. & DeLong, E. F. Multiple archaeal groups mediate methane oxidation in anoxic cold seep sediments. Proc. Natl. Acad. Sci. 99(11), 7663–7668 (2002).
Govenar, B. Shaping vent and seep communities: habitat provision and modification by foundation species. In The Vent and Seep Biota: Aspects from Microbes to Ecosystems 403–432 (Springer, 2010).
Orcutt, B. N., Sylvan, J. B., Knab, N. J. & Edwards, K. J. Microbial Ecology of the Dark Ocean above, at, and below the Seafloor. Microbiol. Mol. Biol. Rev. 75, 361–422 (2011).
Mcmullin, E. R., Bergquist, D. C., & Fisher, C. R. Metazoans in extreme environments: Adaptations of hydrothermal vent and hydrocarbon seep Fauna. Gravit. Space Res. 13(2) (2007).
Garritano, A. N. et al. Species-specific relationships between deep sea sponges and their symbiotic Nitrosopumilaceae. ISME J. 1–3 (2023).
German, C. R. et al. Deep-water chemosynthetic ecosystem research during the census of marine life decade and beyond: A proposed deep-ocean road map. PLoS One 6(8), e23259 (2011).
Ward, M. E., Shields, J. D. & Van Dover, C. L. Parasitism in species of Bathymodiolus (Bivalvia: Mytilidae) mussels from deep-sea seep and hydrothermal vents. Dis. Aquat. Organ. 62, 1–16 (2004).
Feng, D. et al. Using Bathymodiolus tissue stable carbon, nitrogen and sulfur isotopes to infer biogeochemical process at a cold seep in the South China Sea. Deep Sea Res. Part I Oceanogr. Res. Pap. 104, 52–59 (2015).
Yu, J., Wang, M., Liu, B., Yue, X. & Li, C. Gill symbionts of the cold-seep mussel Bathymodiolus platifrons: Composition, environmental dependency and immune control. Fish Shellfish Immunol. 86, 246–252 (2019).
Weldon, S. R., Strand, M. R. & Oliver, K. M. Phage loss and the breakdown of a defensive symbiosis in aphids. Proc. Royal Soc. B 280(1751), 20122103 (2013).
Zhou, K., Xu, Y., Zhang, R. & Qian, P. Y. Phages associated with animal holobionts in deep-sea hydrothermal vents and cold seeps. Deep Sea Res. Part I Oceanogr. Res. Pap 190, 103900 (2022).
Li, Z. et al. Deep sea sediments associated with cold seeps are a subsurface reservoir of viral diversity. ISME J. 15, 2366–2378 (2021).
Takishita, K. Diversity of microbial eukaryotes in deep sea chemosynthetic ecosystems illuminated by molecular techniques. In Marine Protists: Diversity and Dynamics 47–61 (Springer, 2015).
Jing, H., Wang, R., Jiang, Q., Zhang, Y. & Peng, X. Anaerobic methane oxidation coupled to denitrification is an important potential methane sink in deep-sea cold seeps. Sci. Total Environ. 748, 142459 (2020).
Joye, S. B. The geology and biogeochemistry of hydrocarbon seeps. Annu. Rev. Earth Planet. Sci. 48, 205–231 (2020).
Cleary, D. F. R. et al. Habitat- and host-related variation in sponge bacterial symbiont communities in Indonesian waters. FEMS Microbiol. Ecol. 85, 465–482 (2013).
Kellogg, C. A. Enumeration of viruses and prokaryotes in deep-sea sediments and cold seeps of the Gulf of Mexico. Deep Sea Res. Part II Top. Stud. Oceanogr. 57(21–23), 2002–2007 (2010).
Sacristán-Soriano, O., Pérez Criado, N. & Avila, C. Host species determines symbiotic community composition in Antarctic sponges (Porifera: Demospongiae). Front. Mar. Sci. 7, 474 (2020).
Yoshida-Takashima, Y. et al. Spatial distribution of viruses associated with planktonic and attached microbial communities in hydrothermal environments. Appl. Environ. Microbiol. 78, 1311–1320 (2012).
Patra, A. K., Perez, M., Jang, S. J. & Won, Y. J. A regulatory hydrogenase gene cluster observed in the thioautotrophic symbiont of Bathymodiolus mussel in the East Pacific Rise. Sci. Rep. https://doi.org/10.1038/s41598-022-26669-y (2022).
Ponnudurai, R. et al. Genome sequence of the sulfur-oxidizing Bathymodiolus thermophilus gill endosymbiont. Stand. Genomic Sci. https://doi.org/10.1186/s40793-017-0266-y (2017).
Sibuet, M. & Olu, K. Biogeography, biodiversity and fluid dependence of deep-sea cold-seep communities at active and passive margins. Deep Sea Res. Part II Top. Stud. Oceanogr. 45(1–3), 517–567 (1998).
Chen, X., Weinbauer, M. G., Jiao, N. & Zhang, R. Revisiting marine lytic and lysogenic virus–host interactions: Kill-the-Winner and Piggyback-the-Winner. Sci. Bull. 66(9), 871–874 (2021).
Anantharaman, K. et al. Sulfur oxidation genes in diverse deep-sea viruses. Science 344(1979), 757–760 (2014).
Kathuria, S. & Martiny, A. C. Prevalence of a calcium-based alkaline phosphatase associated with the marine cyanobacterium Prochlorococcus and other ocean bacteria. Environ. Microbiol. 13(1), 74–83 (2011).
Thompson, L. R. et al. Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc. Natl. Acad. Sci. 108(39), E757–E764 (2011).
Sullivan, M. B. et al. Genomic analysis of oceanic cyanobacterial myoviruses compared with T4-like myoviruses from diverse hosts and environments. Environ. Microbiol. 12(11), 3035–3056 (2010).
Zhao, J. et al. Novel viral communities potentially assisting in carbon, nitrogen, and sulfur metabolism in the upper slope sediments of Mariana Trench. mSystems 7, e0135821 (2022).
Enav, H., Mandel-Gutfreund, Y. & Béjà, O. Comparative metagenomic analyses reveal viral-induced shifts of host metabolism towards nucleotide biosynthesis. Microbiome 2(1), 1–12 (2014).
Moya, A., Peretó, J., Gil, R. & Latorre, A. Learning how to live together: Genomic insights into prokaryote–animal symbioses. Nat. Rev. Genet 9(3), 218–229 (2008).
Dubilier, N., Claudia, B. & Christian, L. Symbiotic diversity in marine animals: The art of harnessing chemosynthesis. Nat. Rev. Microbiol 6(10), 725–740 (2008).
Lin, Y. T. et al. Interactions among deep-sea mussels and their epibiotic and endosymbiotic chemoautotrophic bacteria: Insights from multi-omics analysis. Zool. Res. 44(1), 106 (2023).
Pan, D., Morono, Y., Inagaki, F. & Takai, K. An improved method for extracting viruses from sediment: Detection of far more viruses in the subseafloor than previously reported. Front. Microbiol. 10, 878 (2019).
Zhou, K., Qian, P. Y., Zhang, T., Xu, Y. & Zhang, R. Unique phage–bacterium interplay in sponge holobionts from the southern Okinawa Trough hydrothermal vent. Environ. Microbiol. Rep. 13, 675–683 (2021).
Guo, J. et al. VirSorter2: A multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 1–13 (2021).
Gulino, K. et al. Initial mapping of the New York City wastewater virome. Msystems 5(3), e00876-e919 (2020).
Rusanova, A., Fedorchuk, V., Toshchakov, S., Dubiley, S. & Sutormin, D. An interplay between viruses and bacteria associated with the white sea sponges revealed by metagenomics. Life 12(1), 25 (2022).
Gong, Z. et al. Viral diversity and its relationship with environmental factors at the surface and deep sea of Prydz Bay, Antarctica. Front. Microbiol. 9, 2981 (2018).
Deng, Z. et al. Phage-prokaryote coexistence strategy mediates microbial community diversity in the intestine and sediment microhabitats of shrimp culture pond ecosystem. Front. Microbiol. 13, 1011342 (2022).
Zhou, K. et al. Viruses in marine invertebrate holobionts: Complex interactions between phages and bacterial symbionts. Annu. Rev. Mar. Sci. 16, 467–485 (2024).
Bryson, S. J., Thurber, A. R., Correa, A. M., Orphan, V. J. & Vega Thurber, R. A novel sister clade to the enterobacteria microviruses (family Microviridae) identified in methane seep sediments. Environ. Microbiol. 17, 3708–3721 (2015).
Thomas, E., Anderson, R. E., Li, V., Rogan, L. J. & Huber, J. A. Diverse viruses in deep-sea hydrothermal vent fluids have restricted dispersal across ocean basins. Msystems 6(3), 10–1128 (2021).
Zhang, R. et al. Viral control of biomass and diversity of bacterioplankton in the deep sea. Commun. Biol. 3(1), 256 (2020).
Andreu-Moreno, I., Bou, J. V. & Sanjuán, R. Cooperative nature of viral replication. Sci. Adv. 6(49), eabd4942 (2020).
de Paoli, P. & Carbone, A. Microenvironmental abnormalities induced by viral cooperation: Impact on lymphomagenesis. Semin. Cancer Biol. 34, 70–80 (2015).
Sanjuán, R. The social life of viruses. Annu. Rev. Virol. 8, 183–199 (2021).
Segredo-Otero, E. & Sanjuán, R. Cooperative virus-virus interactions: An evolutionary perspective. BioDesign Res. https://doi.org/10.34133/2022/9819272 (2022).
DaPalma, T., Doonan, B. P., Trager, N. M. & Kasman, L. M. A systematic approach to virus–virus interactions. Virus Res. 149(1), 1–9 (2010).
Bondy-Denomy, J. et al. Prophages mediate defense against phage infection through diverse mechanisms. ISME J. 10, 2854–2866 (2016).
Hwang, Y., Roux, S., Coclet, C., Krause, S. J. & Girguis, P. R. Viruses interact with hosts that span distantly related microbial domains in dense hydrothermal mats. Nat. Microbiol. 8(5), 946–957 (2023).
Bdeir, N. et al. A system for production of defective interfering particles in the absence of infectious influenza A virus. PLoS One 14(3), e0212757 (2019).
Yang, Y. et al. The antiviral and antitumor effects of defective interfering particles/genomes and their mechanisms. Front. Microbiol. 10, 1852 (2019).
Gyles, C. & Boerlin, P. Horizontally transferred genetic elements and their role in pathogenesis of bacterial disease. Vet. Pathol. 51(2), 328–340 (2014).
Wilson, K. Preparation of genomic DNA from bacteria. Curr. Protoc. Mol. Biol. 56(1), 2–4 (2001).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9(4), 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Cai, R., Zhang, J., Liu, R., & Sun, C. Metagenomic insights into the metabolism and ecologic functions of the widespread DPANN archaea from deep-sea hydrothermal vents. bioRxiv (2020).
Li, D. et al. MEGAHIT v1.0: A fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Bushnell, B. BBMap: A fast, accurate, splice-aware aligner (No. LBNL-7065E). Lawrence Berkeley National Lab. (LBNL) (2014).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7(10), e1002195 (2011).
Hyatt, D. et al. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 1–11 (2010).
Lu, J., Breitwieser, F. P., Thielen, P. & Salzberg, S. L. Bracken: Estimating species abundance in metagenomics data. PeerJ Comput. Sci. 3, e104 (2017).
Wood, D. E. & Salzberg, S. L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15(3), 1–12 (2014).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Li, B. & Dewey, C. N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. https://doi.org/10.1186/1471-2105-12-323 (2011).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11(3), 1–9 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1), 139–140 (2009).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12(1), 59–60 (2015).
Azad, A. et al. HipMCL: A high-performance parallel implementation of the Markov clustering algorithm for large-scale networks. Nucleic Acids Res. 46(6), e33 (2018).
Nepusz, T., Yu, H. & Paccanaro, A. Detecting overlapping protein complexes in protein-protein interaction networks. Nat. Methods 9, 471–472 (2012).
Hryckowian, A. J. et al. Bacteroides thetaiotaomicron-infecting bacteriophage isolates inform sequence-based host range predictions. Cell Host Microbe 28(3), 371–379 (2020).
Shang, J. & Sun, Y. CHERRY: A Computational metHod for accuratE pRediction of virus–pRokarYotic interactions using a graph encoder–decoder model. Brief. Bioinform. 23(5), bbac182 (2022).
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P. L. & Ideker, T. Cytoscape 2.8: New features for data integration and network visualization. Bioinformatics 27, 431–432 (2011).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Acknowledgements
We thank the crew members of the R/V Kexue for their assistance in sample collection and the laboratory staff for continuous technical advice and helpful discussions. The present study was supported by the Laoshan Laboratory (No. 2022QNLM030004), National Natural Science Foundation of China (42030407, 42106134, 42076091 and 42221005), the Key Research Program of Frontier Sciences (ZDBS-LY-DQC032), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA22050303 and XDB42020401).
Author information
Authors and Affiliations
Contributions
Y.Z. and H.C. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Chen, H., Lian, C. et al. Insights into phage-bacteria interaction in cold seep Gigantidas platifrons through metagenomics and transcriptome analyses. Sci Rep 14, 10540 (2024). https://doi.org/10.1038/s41598-024-61272-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-61272-3
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.