Abstract
Immune checkpoint inhibitors (ICIs) targeting programmed cell death protein 1 or its ligand (PD-1/L1) have expanded the treatment landscape against cancers but are effective in only a subset of patients. Tumor mutation burden (TMB) is postulated to be a generic determinant of ICI-dependent tumor rejection. Here we describe the association between TMB and survival outcomes among microsatellite-stable cancers in a real-world clinicogenomic cohort consisting of 70,698 patients distributed across 27 histologies. TMB was associated with survival benefit or detriment depending on tissue and treatment context, with eight cancer types demonstrating a specific association between TMB and improved outcomes upon treatment with anti-PD-1/L1 therapies. Survival benefits were noted over a broad range of TMB cutoffs across cancer types, and a dose-dependent relationship between TMB and outcomes was observed in a subset of cancers. These results have implications for the use of cancer-agnostic and universal TMB cutoffs to guide the use of anti-PD-1/L1 therapies, and they underline the importance of tissue context in the development of ICI biomarkers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data will be made available upon reasonable request with the permission of Caris Life Sciences. Raw sequencing data are owned by Caris Life Sciences and cannot be shared due to patient privacy and protected proprietary information. Access to aggregated data can be requested by contacting the corresponding author, including a brief description of the requirements and intended use. Requests will be discussed with the Caris data access team and a response given within 4 weeks. External datasets used in this study are available from the following public resources: gnomAD, gnomad.broadinstitute.org; International Genome Sample Resource (1000 Genomes Project), www.internationalgenome.org; dbSNP, www.ncbi.nlm.nih.gov/snp. Source data are provided with this paper.
References
Bagchi, S., Yuan, R. & Engleman, E. G. Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu. Rev. Pathol. 16, 223–249 (2021).
Robert, C. A decade of immune-checkpoint inhibitors in cancer therapy. Nat. Commun. 11, 3801 (2020).
Vaddepally, R. K., Kharel, P., Pandey, R., Garje, R. & Chandra, A. B. Review of indications of FDA-approved immune checkpoint inhibitors per NCCN guidelines with the level of evidence. Cancers 12, 738 (2020).
Lei, Y., Li, X., Huang, Q., Zheng, X. & Liu, M. Progress and challenges of predictive biomarkers for immune checkpoint blockade. Front. Oncol. 11, 617335 (2021).
Wang, D. R., Wu, X. L. & Sun, Y. L. Therapeutic targets and biomarkers of tumor immunotherapy: response versus non-response. Signal Transduct. Target. Ther. 7, 331 (2022).
Gunjur, A. et al. ‘Know thyself’—host factors influencing cancer response to immune checkpoint inhibitors. J. Pathol. 257, 513–525 (2022).
Lybaert, L. et al. Challenges in neoantigen-directed therapeutics. Cancer Cell 41, 15–40 (2023).
Ganesan, S. & Mehnert, J. Biomarkers for response to immune checkpoint blockade. Annu. Rev. Cancer Biol. 4, 331–351 (2020).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Segal, N. H. et al. Epitope landscape in breast and colorectal cancer. Cancer Res. 68, 889–892 (2008).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Marabelle, A. et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 21, 1353–1365 (2020).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Valero, C. et al. The association between tumor mutational burden and prognosis is dependent on treatment context. Nat. Genet. 53, 11–15 (2021).
Valero, C. et al. Response rates to anti-PD-1 immunotherapy in microsatellite-stable solid tumors with 10 or more mutations per megabase. JAMA Oncol. 7, 739–743 (2021).
McGrail, D. J. et al. High tumor mutation burden fails to predict immune checkpoint blockade response across all cancer types. Ann. Oncol. 32, 661–672 (2021).
Merino, D. M. et al. Establishing guidelines to harmonize tumor mutational burden (TMB): in silico assessment of variation in TMB quantification across diagnostic platforms: phase I of the Friends of Cancer Research TMB Harmonization Project. J. Immunother. Cancer 8, e000147 (2020).
Offin, M. et al. Tumor mutation burden and efficacy of EGFR-tyrosine kinase inhibitors in patients with EGFR-mutant lung cancers. Clin. Cancer Res. 25, 1063–1069 (2019).
McGranahan, N. & Swanton, C. Neoantigen quality, not quantity. Sci. Transl. Med. 11, eaax7918 (2019).
Sha, D. et al. Tumor mutational burden as a predictive biomarker in solid tumors. Cancer Discov. 10, 1808–1825 (2020).
Chowell, D. et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 359, 582–587 (2018).
Havel, J. J., Chowell, D. & Chan, T. A. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat. Rev. Cancer 19, 133–150 (2019).
Doroshow, D. B. et al. PD-L1 as a biomarker of response to immune-checkpoint inhibitors. Nat. Rev. Clin. Oncol. 18, 345–362 (2021).
Davis, A. A. & Patel, V. G. The role of PD-L1 expression as a predictive biomarker: an analysis of all US Food and Drug Administration (FDA) approvals of immune checkpoint inhibitors. J. Immunother. Cancer 7, 278 (2019).
Li, F. et al. The association between CD8+ tumor-infiltrating lymphocytes and the clinical outcome of cancer immunotherapy: a systematic review and meta-analysis. EClinicalMedicine 41, 101134 (2021).
Ott, P. A. et al. T-cell-inflamed gene-expression profile, programmed death ligand 1 expression, and tumor mutational burden predict efficacy in patients treated with pembrolizumab across 20 cancers: KEYNOTE-028. J. Clin. Oncol. 37, 318–327 (2019).
Finotello, F. et al. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 11, 34 (2019).
Bao, R., Stapor, D. & Luke, J. J. Molecular correlates and therapeutic targets in T cell-inflamed versus non-T cell-inflamed tumors across cancer types. Genome Med. 12, 90 (2020).
Marcus, L. et al. FDA approval summary: pembrolizumab for the treatment of tumor mutational burden-high solid tumors. Clin. Cancer Res. 27, 4685–4689 (2021).
Subbiah, V., Solit, D. B., Chan, T. A. & Kurzrock, R. The FDA approval of pembrolizumab for adult and pediatric patients with tumor mutational burden (TMB) ≥10: a decision centered on empowering patients and their physicians. Ann. Oncol. 31, 1115–1118 (2020).
Prasad, V. & Addeo, A. The FDA approval of pembrolizumab for patients with TMB > 10 mut/Mb: was it a wise decision? No. Ann. Oncol. 31, 1112–1114 (2020).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Jardim, D. L., Goodman, A., de Melo Gagliato, D. & Kurzrock, R. The challenges of tumor mutational burden as an immunotherapy biomarker. Cancer Cell 39, 154–173 (2021).
Gupta, R., Mehta, A. & Wajapeyee, N. Transcriptional determinants of cancer immunotherapy response and resistance. Trends Cancer 8, 404–415 (2022).
Nowicki, T. S., Hu-Lieskovan, S. & Ribas, A. Mechanisms of resistance to PD-1 and PD-L1 blockade. Cancer J. 24, 47–53 (2018).
Richman, L. P., Vonderheide, R. H. & Rech, A. J. Neoantigen dissimilarity to the self-proteome predicts immunogenicity and response to immune checkpoint blockade. Cell Syst. 9, 375–382 (2019).
Luksza, M. et al. A neoantigen fitness model predicts tumour response to checkpoint blockade immunotherapy. Nature 551, 517–520 (2017).
Wolf, Y. & Sameuls, Y. Neoantigens in cancer immunotherapy: quantity vs. quality. Mol. Oncol. 17, 1457–1459 (2023).
Kieffer, Y. et al. Single-cell analysis reveals fibroblast clusters linked to immunotherapy resistance in cancer. Cancer Discov. 10, 1330–1351 (2020).
Niknafs, N. et al. Persistent mutation burden drives sustained anti-tumor immune responses. Nat. Med. 29, 440–449 (2023).
Acknowledgements
D.H. is supported by a Cancer Prevention and Research Institute of Texas Early Clinical Investigator Award (RP200549) and the Josephine Hughes Sterling Foundation. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the paper. D.S.B.H. is supported by the Dr. Miriam and Sheldon Adelson Medical Research Foundation. The remaining authors received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
D.H. and H.Z. conceptualized the study. M.M., M.E., A.E., J.X. and D.H. performed data analyses. A.S., W.E.-D., E.S.A., S.L.G., M.J.H., H.B., D.S.B.H., S.V.L., P.C.M., R.R.M., T.W.-D., J.M., G.W.S., D.S., H.Z. and D.H. contributed to the assembly of the CARIS cohort. M.M. and D.H. drafted the paper, and all authors participated in the review and editing of the paper.
Corresponding author
Ethics declarations
Competing interests
A.E., J.X., G.W.S. and D.S. are employees of Caris Life Sciences. S.L.G. serves a paid consultant and advisor to Pfizer, Daiichi Sankyo, Eli Lilly, AstraZeneca, Genentech, SeaGen, Novartis and Menarini and has stock ownership in HCA Healthcare. E.S.A. serves as a paid consultant and advisor to Janssen, Astellas, Sanofi, Dendreon, Bayer, BMS, Amgen, Constellation, Blue Earth, Exact Sciences, Invitae, Curium, Pfizer, Merck, AstraZeneca, Clovis and Eli Lilly; has received research support (to his institution) from Janssen, J&J, Sanofi, BMS, Pfizer, AstraZeneca, Novartis, Curium, Constellation, Celgene, Merck, Bayer and Clovis; and is the co-inventor of a patented AR-V7 biomarker technology that has been licensed to Qiagen. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Samra Turajlic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 TMB cutoffs associated with ICI benefit retain predictive value across demographics.
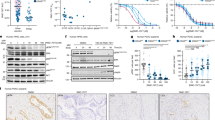
Subset analyses of patients stratified by age groups and self-reported sex show that the association between TMB cutoffs and outcomes are similar across demographics. Forest plots depict hazard ratios (squares) and error bars indicate 95% confidence intervals.
Extended Data Fig. 2 Associations between overall survival and TMB at the 75th percentile for individual cancer types are independent of the sequencing platform used.
Hazard ratios of overall survival in the ICI cohort using a TMB threshold at the 75th percentile for individual cancer types were separately calculated for cases analyzed by the 592-gene panel or exome sequencing. Forest plots depict hazard ratios (squares) and error bars indicate 95% confidence intervals.
Extended Data Fig. 3 Immune correlates between TMB-high and TMB-low cancers using the earliest cutoff at which ICIs are predictive.
PD-L1 positive cell frequency, T-cell inflammatory score, and CD8 + T cell frequency are compared between TMB-high cancers and TMB-low cancers using the earliest cutoff at which ICIs are associated with OS benefit. Biomarkers enriched in TMB-high and TMB-low cancers using a false discovery rate of 0.05 are highlighted.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Muquith, M., Espinoza, M., Elliott, A. et al. Tissue-specific thresholds of mutation burden associated with anti-PD-1/L1 therapy benefit and prognosis in microsatellite-stable cancers. Nat Cancer (2024). https://doi.org/10.1038/s43018-024-00752-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43018-024-00752-x