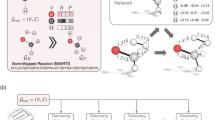
Abstract
Transition state search is key in chemistry for elucidating reaction mechanisms and exploring reaction networks. The search for accurate 3D transition state structures, however, requires numerous computationally intensive quantum chemistry calculations due to the complexity of potential energy surfaces. Here we developed an object-aware SE(3) equivariant diffusion model that satisfies all physical symmetries and constraints for generating sets of structures—reactant, transition state and product—in an elementary reaction. Provided reactant and product, this model generates a transition state structure in seconds instead of hours, which is typically required when performing quantum-chemistry-based optimizations. The generated transition state structures achieve a median of 0.08 Å root mean square deviation compared to the true transition state. With a confidence scoring model for uncertainty quantification, we approach an accuracy required for reaction barrier estimation (2.6 kcal mol–1) by only performing quantum chemistry-based optimizations on 14% of the most challenging reactions. We envision usefulness for our approach in constructing large reaction networks with unknown mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The Transition1x dataset55 used in this work can be found at GitLab, https://gitlab.com/matschreiner/Transition1x, with https://doi.org/10.6084/m9.figshare.19614657.v4. Source data are provided with this paper.
Code availability
Codebase for OA-ReactDiff is available as an open-source repository on GitHub for contiguous development, https://github.com/chenruduan/OAReactDiff. A stable version of the code56 used in this work is available at Zenodo, https://doi.org/10.5281/zenodo.10054963.
References
Dewyer, A. L., Argüelles, A. J. & Zimmerman, P. M. Methods for exploring reaction space in molecular systems. WIREs Comput. Mol. Sci. 8, e1354 (2018).
Unsleber, J. P. & Reiher, M. The exploration of chemical reaction networks. Annu. Rev. Phys. Chem. 71, 121–142 (2020).
Truhlar, D. G., Garrett, B. C. & Klippenstein, S. J. Current status of transition-state theory. J. Phys. Chem. 100, 12771–12800 (1996).
Mardirossian, N. & Head-Gordon, M. Thirty years of density functional theory in computational chemistry: an overview and extensive assessment of 200 density functionals. Mol. Phys. 115, 2315–2372 (2017).
Durant, J. L. Evaluation of transition state properties by density functional theory. Chem. Phys. Lett. 256, 595–602 (1996).
Simm, G. N., Vaucher, A. C. & Reiher, M. Exploration of reaction pathways and chemical transformation networks. J. Phys. Chem. A 123, 385–399 (2019).
Wang, L.-P. et al. Discovering chemistry with an ab initio nanoreactor. Nat. Chem. 6, 1044–1048 (2014).
Pieri, E. et al. The non-adiabatic nanoreactor: towards the automated discovery of photochemistry. Chem. Sci. 12, 7294–7307 (2021).
Zeng, J., Cao, L., Xu, M., Zhu, T. & Zhang, J. Z. H. Complex reaction processes in combustion unraveled by neural network-based molecular dynamics simulation. Nat. Commun. 11, 5713 (2020).
Van de Vijver, R. & Zádor, J. Kinbot: automated stationary point search on potential energy surfaces. Comput. Phys. Commun. 248, 106947 (2020).
von Lilienfeld, O. A., Müller, K.-R. & Tkatchenko, A. Exploring chemical compound space with quantum-based machine learning. Nat. Rev. Chem. 4, 347–358 (2020).
Margraf, J. T., Jung, H., Scheurer, C. & Reuter, K. Exploring catalytic reaction networks with machine learning. Nat. Catal. 6, 112–121 (2023).
Sheppard, D., Terrell, R. & Henkelman, G. Optimization methods for finding minimum energy paths. J. Chem. Phys. 128, 134106 (2008).
Schreiner, M., Bhowmik, A., Vegge, T., Busk, J. & Winther, O. Transition1x - a dataset for building generalizable reactive machine learning potentials. Sci. Data 9, 779 (2022).
Zhao, Q. & Savoie, B. M. Simultaneously improving reaction coverage and computational cost in automated reaction prediction tasks. Nat. Comput. Sci. 1, 479–490 (2021).
Lemm, D., von Rudorff, G. F. & von Lilienfeld, O. A. Machine learning based energy-free structure predictions of molecules, transition states, and solids. Nat. Commun. 12, 4468 (2021).
Zhang, J. et al. Deep reinforcement learning of transition states. Phys. Chem. Chem. Phys. 23, 6888–6895 (2021).
Pattanaik, L., Ingraham, J. B., Grambow, C. A. & Green, W. H. Generating transition states of isomerization reactions with deep learning. Phys. Chem. Chem. Phys. 22, 23618–23626 (2020).
Makoś, M. Z., Verma, N., Larson, E. C., Freindorf, M. & Kraka, E. Generative adversarial networks for transition state geometry prediction. J. Chem. Phys. 155, 024116 (2021).
Choi, S. Prediction of transition state structures of gas-phase chemical reactions via machine learning. Nat. Commun. 14, 1168 (2023).
Schreiner, M., Bhowmik, A., Vegge, T., Jørgensen, P. B. & Winther, O. NeuralNEB—neural networks can find reaction paths fast. Mach. Learn. Sci. Technol. 3, 045022 (2022).
Ho, J., Jain, A. & Abbeel, P. in (eds Larochelle, H. et al.) Advances in Neural Information Processing Systems Vol. 33, 6840–6851 (Curran Associates, 2020).
Sohl-Dickstein, J., Weiss, E., Maheswaranathan, N. & Ganguli, S. Deep unsupervised learning using nonequilibrium thermodynamics. In International Conference on Machine Learning 2256–2265 (2015).
Song, Y. et al. Score-based generative modeling through stochastic differential equations. In Int. Conference on Learning Representations (2020).
Hoogeboom, E., Satorras, V. G., Vignac, C. & Welling, M. Equivariant diffusion for molecule generation in 3D. In Proc. 39th International Conference on Machine Learning 8867–8887 (2022).
Corso, G., Stärk, H., Jing, B., Barzilay, R. & Jaakkola, T. DiffDock: Diffusion steps, twists, and turns for molecular docking. In Int. Conference on Learning Representations (2023).
Schneuing, A. et al. Structure-based drug design with equivariant diffusion models. Preprint at https://doi.org/10.48550/arXiv.2210.13695 (2022).
Thomas, N. et al. Tensor field networks: rotation- and translation-equivariant neural networks for 3D point clouds. Preprint at https://doi.org/10.48550/arXiv.1802.08219 (2018).
Satorras, V. G., Hoogeboom, E. & Welling, M. E(n) equivariant graph neural networks. In Proc. 38th International Conference on Machine Learning 9323–9332 (2021).
Batzner, S. et al. E(3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials. Nat. Commun. 13, 2453 (2022).
Zhou, H.-C., Long, J. R. & Yaghi, O. M. Introduction to metal-organic frameworks. Chem. Rev. 112, 673–674 (2012).
Du, W. et al. A new perspective on building efficient and expressive 3D equivariant graph neural networks. Preprint at https://doi.org/10.48550/arXiv.2304.04757 (2023).
Lugmayr, A. et al. Repaint: inpainting using denoising diffusion probabilistic models. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) (2022); https://doi.org/10.1109/CVPR52688.2022.01117
Henkelman, G., Uberuaga, B. P. & Jónsson, H. A climbing image nudged elastic band method for finding saddle points and minimum energy paths. J. Phys. Chem. 113, 9901–9904 (2000).
Chai, J.-D. & Head-Gordon, M. Systematic optimization of long-range corrected hybrid density functionals. J. Phys. Chem. 128, 084106 (2008).
Ditchfield, R., Hehre, W. J. & Pople, J. A. Self-consistent molecular-orbital methods. ix. an extended gaussian-type basis for molecular-orbital studies of organic molecules. J. Phys. Chem. 54, 724–728 (1971).
Grambow, C. A., Pattanaik, L. & Green, W. H. Reactants, products, and transition states of elementary chemical reactions based on quantum chemistry. Sci. Data 7, 137 (2020).
Grambow, C. A., Pattanaik, L. & Green, W. H. Deep learning of activation energies. J. Phys. Chem. Lett. 11, 2992–2997 (2020).
Ruddigkeit, L., van Deursen, R., Blum, L. C. & Reymond, J.-L. Enumeration of 166 billion organic small molecules in the chemical universe database GDB-17. J. Chem. Inf. Model. 52, 2864–2875 (2012).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589 (2021).
Duan, C., Nandy, A., Meyer, R., Arunachalam, N. & Kulik, H. J. A transferable recommender approach for selecting the best density functional approximations in chemical discovery. Nat. Comput. Sci. 3, 38–47 (2023).
Seifert, G. & Joswig, J.-O. Density-functional tight binding—an approximate density-functional theory method. WIREs Comput. Mol. Sci. 2, 456–465 (2012).
Liu, W.-G. & Goddard, W. A. I. First-principles study of the role of interconversion between NO2, N2O4, cis-ONO-NO2, and trans-ONO-NO2 in chemical processes. J. Am. Chem. Soc. 134, 12970–12978 (2012).
Duan, C., Chu, D. B. K., Nandy, A. & Kulik, H. J. Detection of multi-reference character imbalances enables a transfer learning approach for virtual high throughput screening with coupled cluster accuracy at dft cost. Chem. Sci. 13, 4962–4971 (2022).
Lipman, Y., Chen, R. T. Q., Ben-Hamu, H., Nickel, M. & Le, M. Flow matching for generative modeling. In 11th International Conference on Learning Representations (2023).
Liu, G.-H. et al. I2SB: Image-to-image schrödinger bridge. Preprint at https://doi.org/10.48550/arXiv.2302.05872 (2023).
Kim, S., Woo, J. & Kim, W. Y. Diffusion-based generative AI for exploring transition states from 2D molecular graphs. Preprint at https://doi.org/10.48550/arXiv.2304.12233 (2023).
Zhao, Q. et al. Comprehensive exploration of graphically defined reaction spaces. Sci. Data 10, 145 (2023).
Serre, J.-P. et al. Linear Representations of Finite Groups (Springer, 1977).
Bronstein, M. M., Bruna, J., Cohen, T. & Veličković, P. Geometric deep learning: grids, groups, graphs, geodesics, and gauges. Preprint at https://doi.org/10.48550/arXiv.2104.13478 (2021).
Köhler, J., Klein, L. & Noé, F. Equivariant flows: exact likelihood generative learning for symmetric densities. In International Conference on Machine Learning 5361–5370 (2020).
Nichol, A. Q. & Dhariwal, P. Improved denoising diffusion probabilistic models. In Proc. 38th International Conference on Machine Learning 8162–8171 (2021).
Du, W. et al. SE (3) equivariant graph neural networks with complete local frames. In International Conference on Machine Learning 5583–5608 (2022).
Gilmer, J., Schoenholz, S. S., Riley, P. F., Vinyals, O. & Dahl, G. E. Neural message passing for quantum chemistry. In International Conference on Machine Learning 1263–1272 (2017).
Transition1x data release. figshare https://doi.org/10.6084/m9.figshare.19614657.v4 (2023).
OA-ReactDiff stable code release. Zenodo https://doi.org/10.5281/zenodo.10054963 (2023).
Acknowledgements
This work was supported by the US Office of Naval Research under grant no. N00014-20-1-2150 (C.D. and H.J.K.) and National Science Foundation grant CBET-1846426 (H.J. and H.J.K.). C.D. thanks the Molecular Sciences Software Institute for the fellowship support under NSF grant OAC-1547580. C.D. thanks Q. Zhao and M. Monkey for discussions about elementary reactions. C.D. thanks A. Nandy and W. Du for discussions about equivariant graph neural networks. C.D. and Y.D. thank G.-H. Liu and T. Chen for discussions about diffusion model and Schrödinger bridge. C.D. and H.J. thank Y. Zhao for his help on preparing a demo Jupyter notebook for this work. The authors thank S. Choi and M. Schreiner for communications and providing their raw data that makes the comparison in Table 1 possible.
Author information
Authors and Affiliations
Contributions
C.D. was responsible for conceptualization, methodology, software, validation, investigation, data curation, writing of the original draft, review, editing and visualization. Y.D. was responsible for methodology, software, writing of original draft, review and editing. H.J. was responsible for data curation, review and editing. H.J.K. was responsible for writing of the original draft, review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Sunghwan Choi, Hyunwook Jung and Matteo Maestri for their contribution to the peer review of this work. Primary Handling Editor: Kaitlin McCardle, in collaboration with the Nature Computational Science team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–4, Figs. 1–10 and Text 1.
Source data
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 5
Statistical source data for Fig. 5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duan, C., Du, Y., Jia, H. et al. Accurate transition state generation with an object-aware equivariant elementary reaction diffusion model. Nat Comput Sci 3, 1045–1055 (2023). https://doi.org/10.1038/s43588-023-00563-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00563-7
This article is cited by
-
Diffusion-based generative AI for exploring transition states from 2D molecular graphs
Nature Communications (2024)