Abstract
Resistance to nirmatrelvir (Paxlovid) has been shown by multiple groups and may already exist in clinical SARS-CoV-2 isolates. Here a robust cell-based assay is used to determine the relative potencies of nirmatrelvir, ensitrelvir, and FB2001 against a panel of SARS-CoV-2 main protease (Mpro) variants. The results reveal that these three drugs have at least partly distinct resistance mutation profiles and raise the possibility that the latter compounds may be effective in some instances of Paxlovid resistance and vice versa.
Similar content being viewed by others
Antiviral drugs are necessary to combat SARS-CoV-2/COVID-19, particularly with waning interest in the repeated vaccination boosts necessary to keep-up with virus evolution. The main protease (Mpro) of SARS-CoV-2 is essential for virus replication and, accordingly, a proven therapeutic target as evidenced by Paxlovid, an orally administered antiviral drug (active component: nirmatrelvir; Fig. 1a). However, as for drugs developed to treat other viruses1 and for first-generation SARS-CoV-2 vaccines, there is a high probability that variants will emerge that resist nirmatrelvir. Indeed, a flurry of recent studies has described a variety of candidate nirmatrelvir-resistance mutations2,3,4,5,6,7,8,9. Thus, considerable urgency exists to develop next-generation Mpro inhibitors with different resistance mechanisms and, in parallel, robust systems to rapidly assess the potential impact of candidate resistance mutations. Two additional Mpro inhibitors, Ensitrelvir (Xocova) and FB2001 (Bofutrelvir), which are orally and intravenously administered, respectively, are currently being evaluated in clinical trials10,11 (Fig. 1a). Ensitrelvir has also received EUA in Japan (https://www.shionogi.com/global/en/news/2022/11/e20221122.html). However, the potency of these drug candidates against nirmatrelvir-resistant Mpro variants has yet to be fully assessed.
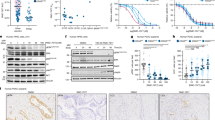
a Co-crystal structures of SARS-CoV-2 Mpro in complex with nirmatrelvir (PDB:7SI9), ensitrelvir (PDB:7VU6), or FB2001 (PDB:6LZE) (chemical structures depicted below each image). Labeled residues, as well as T304I, are tested in panel B. b Fold-change in IC50 relative to WT for the indicated mutants using the live cell gain-of-signal assay in 293T cells.
We recently developed a gain-of-signal system for facile quantification of Mpro inhibition12, and used it together with evolution- and structure-guided approaches to characterize candidate nirmatrelvir- and ensitrelvir-resistance mutations2. This system is comprised of a single plasmid encoding SARS-CoV-2 Mpro flanked by its native cleavage sites and an N-terminal myristoylation domain from Src kinase and a C-terminal HIV-1 Tat fused to firefly luciferase12 (schematic in Supplementary Fig. 1a). Overexpression of catalytically active Mpro in this system results in cleavage of multiple cellular substrates13,14 including at least one required for RNA Polymerase II-dependent gene expression12, which results in low luciferase signal that can be restored by either genetic inactivation or chemical inhibition2,12.
Here, this system is used to examine an expanded panel of Mpro single and double mutants based on recent studies by our group and others2,3,4,5,6,7,8,9 and determine their impact on the potency of nirmatrelvir, ensitrelvir, and FB2001 (heatmap of results in Fig. 1b; specific references and quantification summary in Table 1; representative dose responses in Supplementary Fig. 1b). Several single amino acid substitution mutants selected during serial passaging3,5,6,7 including T21I, L50F, P252L, and T304I show minimal resistance to nirmatrelvir, ensitrelvir, or FB2001 in our assay. In contrast, selective resistance to ensitrelvir is conferred by M49I and M49L, and selective resistance to nirmatrelvir by A173V (highlighted in gray in Table 1). ∆P168 elicits similar resistance to all inhibitors, and synergistic resistance to nirmatrelvir when combined with A173V. S144A and L167F show the greatest resistance to ensitrelvir, intermediate resistance to nirmatrelvir, and lower resistance toward FB2001. M49L in combination with S144A elicits very high resistance to ensitrelvir, intermediate resistance to nirmatrelvir, and essentially no resistance to FB2001. In contrast to E166A and L50F/E166A, which cause a similar broad-spectrum resistance, E166V and L50F/E166V elicit very high resistance to nirmatrelvir, intermediate resistance to ensitrelvir, and substantially lower resistance to FB2001.
Genetic mutants can also exhibit phenotypes in our system in the absence of drug, with some showing wildtype Mpro activity (background luminescence) and others compromising activity weakly or strongly depending on the nature of the mutation (low to high luminescence, respectively). For example, in comparison to wildtype Mpro, catalytic mutants such as C145A yield 50- to 100-fold higher luminescence2,12. The Mpro variant constructs used here display a range of luminescence levels in the absence of drug indicative of near-normal Mpro activity (notably, M49I and M49L), weakly compromised Mpro activity (notably, A173V), and strongly compromised Mpro activity (notably, E166V) (Supplementary Fig. 2). These results suggest that several variants can confer at least partial drug resistance with little loss in Mpro functionality (and likely also viral fitness), whereas others such as E166V require suppressor mutations such as L50F to restore Mpro function to a level that enables virus replication (evidenced by recent resistance studies with pathogenic SARS-CoV-2 in culture and in vivo in animal models3,5).
The results here demonstrate that nirmatrelvir, ensitrelvir, and FB2001 have distinct susceptibilities to several resistance-associated substitutions, even without knowledge of the precise details of each molecular mechanism. Importantly, in cases where bona fide resistance develops against one drug, the others may still prove effective depending on the identity of the selected mutations. Although FB2001 has structural similarity to nirmatrelvir, it appears less susceptible to several resistance-associated mutations, which suggests that it and potentially other peptidomimetic inhibitors in development may be able to achieve greater durability. In support of this possibility, FB2001 also exhibits efficacy against the distantly related coronaviruses 229E and NL63 compared to nirmatrelvir and ensitrelvir (Supplementary Fig. 3a, b). However, the antiviral activity of each drug against large panels of diverse viruses, as well as dedicated resistance studies, will be required to fully evaluate broader spectrum utility.
Importantly, the relative fold-resistance as determined by the gain-of-signal assay associates positively with published findings using infectious viruses2,3,5,6,7 (Supplementary Fig. 4). This live cell assay therefore provides an accurate, safe, and rapid system for assessing resistance. As the SARS-CoV-2 variant pool deepens, this assay and variant panel can be expanded in lock-step to provide early resistance “fingerprints” of candidate next-generation Mpro inhibitors. Such an early profiling strategy has the potential to minimize the risks of developing drugs prone to cross-resistance and to help identify inhibitors with the highest barriers to resistance and broadest spectrum of utility.
Methods
DNA constructs and cell culture
The live cell gain-of-signal assay for Mpro inhibition was performed using the pcDNA5/TO-Src-Mpro-Tat-fLuc reporter construct which encodes the N-myristoylation domain of Src kinase followed by SARS-CoV-2 WA1 Mpro flanked by its cognate cleavage sites, HIV-1 Tat and firefly luciferase12. All SARS-CoV-2 Mpro single and double mutants selected for analysis here were based on recent reports of candidate resistant mutants2,3,4,5,6,7,8,9 generated by site-directed mutagenesis (primers in Supplementary Table 1) and verified by Sanger sequencing. Transfections were done using 293T cells (ATCC CRL-3216) maintained at 37 °C and 5% CO2 in DMEM (Gibco catalog number 11875093) supplemented with 10% fetal bovine serum (ThermoFisher catalog number 11965084) and penicillin-streptomycin (Gibco catalog number 15140122).
Mpro resistance experiments
For each individual SARS-CoV-2 Mpro variant, 3 × 106 293T (ATCC CRL-3216) cells were plated in a 10 cm dish and transfected 24 h later with 2 µg of the corresponding variant plasmid using TransIT-LT1 (Mirus catalog number MIR 2304). Transfected cells were incubated at 37 °C and 5% CO2 for 4 h, washed once with phosphate buffered saline (PBS), trypsinized, resuspended in fresh media, and diluted to a concentration of 4 × 105 cells/ml. 50 µL of each cell suspension was added to a 96-well white clear bottom cell culture plate (ThermoFisher #165306) containing pre-aliquoted inhibitor-supplemented media for a final concentration of 20,000 cells per well and inhibitor dose response range of 10 µM to 2.4 nM. Inhibitors were purchased from commercial vendors (nirmatrelvir, MedChemExpress catalog number HY-138687; ensitrelvir, MedChemExpress catalog number HY-143216; FB2001, Sigma-Aldrich catalog number SML2877) and purity was confirmed by HPLC and NMR. After an additional 44 h incubation (48 h total post-transfection), luciferase activity was quantified by removing growth medium and adding 50 µL of Bright-Glo reagent (Promega catalog number E2610) to each well and incubating at room temperature in the dark for 2 m before measuring luminescence on a Biotek Synergy H1 plate reader.
Percent Mpro inhibition was calculated at each concentration of inhibitor using the formula below using the relative luminescence of an inhibitor (RLi) treated sample to the untreated control for each individual mutant.
Results were plotted using GraphPad Prism 9 and fit using a four-parameter non-linear regression to calculate IC50 values (Fig. S1; Table 1). Resistance of mutants was calculated by the fold change in IC50 of the mutant relative to WT Mpro, and these values were used to generate a heatmap in GraphPad Prism 9 (Fig. 1b).
As an increase in luminescence in the absence of any inhibitor treatment is indicative of decreased Mpro catalytic activity, the relative activity of each mutant was calculated by the formula below using the relative luminescence of a mutant (RLm) to the WT enzyme in the absence of inhibitor (Fig. S2).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All results are presented in the main display items or supplementary figures. The SARS-CoV-2 Mpro gain-of-signal plasmid and mutant derivatives are available upon email request to rsh@uthscsa.edu and completion of a MTA (U.S. Provisional Application Serial No. 63/108,611, filed on November 2, 2020).
References
Majerova, T. & Konvalinka, J. Viral proteases as therapeutic targets. Mol. Aspects Med. 88, 101159 (2022).
Moghadasi, S. A. et al. Transmissible SARS-CoV-2 variants with resistance to clinical protease inhibitors. Sci. Adv. 9, eade8778 (2023).
Zhou, Y. et al. Nirmatrelvir-resistant SARS-CoV-2 variants with high fitness in an infectious cell culture system. Sci. Adv. 8, eadd7197 (2022).
Iketani, S. et al. Functional map of SARS-CoV-2 3CL protease reveals tolerant and immutable sites. Cell Host Microbe 30, 1354–1362.e1356 (2022).
Iketani, S. et al. Multiple pathways for SARS-CoV-2 resistance to nirmatrelvir. Nature 613, 558–564 (2023).
Jochmans, D. et al. The substitutions L50F, E166A, and L167F in SARS-CoV-2 3CLpro are selected by a protease inhibitor in vitro and confer resistance to nirmatrelvir. mBio 14, e0281522 (2023).
Heilmann, E. et al. SARS-CoV-2 3CL(pro) mutations selected in a VSV-based system confer resistance to nirmatrelvir, ensitrelvir, and GC376. Sci. Transl. Med. 15, eabq7360 (2023).
Noske, G. D. et al. Structural basis of nirmatrelvir and ensitrelvir activity against naturally occurring polymorphisms of the SARS-CoV-2 Main Protease. J. Biol. Chem. 299, 103004 (2023).
Lan, S. et al. Nirmatrelvir resistance in SARS-CoV-2 Omicron_BA.1 and WA1 replicons and escape strategies. BioRxiv https://doi.org/10.1101/2022.12.31.522389 (2023).
Dai, W. et al. Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science 368, 1331–1335 (2020).
Unoh, Y. et al. Discovery of S-217622, a noncovalent oral SARS-CoV-2 3CL protease inhibitor clinical candidate for treating COVID-19. J. Med. Chem. 65, 6499–6512 (2022).
Moghadasi, S. A. et al. Gain-of-signal assays for probing inhibition of SARS-CoV-2 M(pro)/3CL(pro) in living cells. mBio 13, e0078422 (2022).
Meyers, J. M. et al. The proximal proteome of 17 SARS-CoV-2 proteins links to disrupted antiviral signaling and host translation. PLoS Pathog. 17, e1009412 (2021).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Acknowledgements
This work was supported by National Institute of Allergy and Infectious Disease grant U19-AI171954. RSH is an Investigator of the Howard Hughes Medical Institute and the Ewing Halsell President’s Council Distinguished Chair at University of Texas Health San Antonio.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.A.M., R.S.H.; Methodology: S.A.M., R.G.B., D.A.H.; Investigation: S.A.M., R.G.B.; Visualization: S.A.M.; Funding acquisition: R.S.H.; Project administration: R.S.H.; Supervision: R.S.H., D.A.H.; Writing—original draft: S.A.M., R.S.H.; Writing—review and editing: All authors.
Corresponding author
Ethics declarations
Competing interests
The Mpro gain-of-signal system is the subject of U.S. Provisional Application Serial No. 63/108,611, filed on November 2, 2020, with R.S.H. and S.A.M. as inventors. The authors declare no additional competing interests.
Ethics
Studies here were performed under University of Minnesota IBC protocol 1902-36822H to R.S.H., University of Minnesota IBC protocol 2111-39591H to D.A.H., and University of Texas Health. San Antonio IBC B-00000013853 to R.S.H.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moghadasi, S.A., Biswas, R.G., Harki, D.A. et al. Rapid resistance profiling of SARS-CoV-2 protease inhibitors. npj Antimicrob Resist 1, 9 (2023). https://doi.org/10.1038/s44259-023-00009-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s44259-023-00009-0