Featured
-
-
Article
| Open Access A comprehensive, mechanistically detailed, and executable model of the cell division cycle in Saccharomyces cerevisiae
A comprehensive, mechanistically detailed, and executable model of the cell division cycle in Saccharomyces cerevisiae
Whole-cell models hold great promise for fundamental and translational biology, but genome-scale modelling of signalling networks has been a challenge. Here, the authors present a genome-scale, mechanistic and executable model of the network controlling and executing the S. cerevisiae cell cycle.
- Ulrike Münzner
- , Edda Klipp
- & Marcus Krantz
-
Article
| Open Access Growth strategy of microbes on mixed carbon sources
Growth strategy of microbes on mixed carbon sources
Bacteria grown on two carbon sources either consume both sources simultaneously or consume them sequentially. Here the authors use a metabolic network model of E. coli to show that optimal protein resource allocation and topological features of the network can explain the choice of carbon acquisition.
- Xin Wang
- , Kang Xia
- & Chao Tang
-
Article
| Open Access Network-based prediction of drug combinations
Network-based prediction of drug combinations
Combination therapy holds great promise, but discovery remains challenging. Here, the authors propose a method to identify efficacious drug combinations for specific diseases, and find that successful combinations tend to target separate neighbourhoods of the disease module in the human interactome.
- Feixiong Cheng
- , István A. Kovács
- & Albert-László Barabási
-
Article
| Open Access Topological scoring of protein interaction networks
Topological scoring of protein interaction networks
Inferring direct protein−protein interactions (PPIs) and modules in PPI networks remains a challenge. Here, the authors introduce an algorithm to infer potential direct PPIs from quantitative proteomic AP-MS data by identifying enriched interactions of each bait relative to the other baits.
- Mihaela E. Sardiu
- , Joshua M. Gilmore
- & Michael P. Washburn
-
Article
| Open Access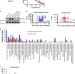 Serine synthesis through PHGDH coordinates nucleotide levels by maintaining central carbon metabolism
Serine synthesis through PHGDH coordinates nucleotide levels by maintaining central carbon metabolism
Serine synthesis from glucose is required even when serine is available from the environment. Here, the authors explain this paradox by showing that the enzyme PHGDH enables nucleotide synthesis by coordinating anabolic fluxes related to central carbon metabolism, independent of its role in serine production.
- Michael A. Reid
- , Annamarie E. Allen
- & Jason W. Locasale
-
Article
| Open Access Modeling genome-wide enzyme evolution predicts strong epistasis underlying catalytic turnover rates
Modeling genome-wide enzyme evolution predicts strong epistasis underlying catalytic turnover rates
The catalytic efficiency of many enzymes is lower than the theoretical maximum. Here, the authors combine genome-scale metabolic modeling with population genetics models to simulate enzyme evolution, and find that strong epistasis limits turnover numbers due to diminishing returns of fitness gains.
- David Heckmann
- , Daniel C. Zielinski
- & Bernhard O. Palsson
-
Article
| Open Access Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models
Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models
Experimental data on enzyme turnover numbers is sparse and noisy. Here, the authors use machine learning to successfully predict enzyme turnover numbers for E. coli, and show that using these to parameterize mechanistic genome-scale models enhances their predictive accuracy.
- David Heckmann
- , Colton J. Lloyd
- & Bernhard O. Palsson
-
Article
| Open Access Genome-scale metabolic reconstructions of multiple Salmonella strains reveal serovar-specific metabolic traits
Genome-scale metabolic reconstructions of multiple Salmonella strains reveal serovar-specific metabolic traits
Salmonella serovars colonize a wide range of hosts but the underlying genetic determinants remain poorly understood. Here, Seif et al. use a network-based computational analysis to link specific metabolic capabilities with host range and nutritional niche.
- Yara Seif
- , Erol Kavvas
- & Jonathan M. Monk
-
Article
| Open Access Systems analysis of intracellular pH vulnerabilities for cancer therapy
Systems analysis of intracellular pH vulnerabilities for cancer therapy
Tumors often exhibit a pH gradient, with an acidic extracellular space and alkaline cytoplasm. Here the authors develop a computational model to show how alkaline pHi supports changes inherent to cancer cell metabolism and acidification disables these adaptations, and demonstrate the effect of acidic pHi on breast cancer cell survival.
- Erez Persi
- , Miquel Duran-Frigola
- & Eytan Ruppin
-
Article
| Open Access Statistical mechanics for metabolic networks during steady state growth
Statistical mechanics for metabolic networks during steady state growth
Single cell growth rate variability has been difficult to understand. Here, the authors apply a generalization of flux balance analysis to single cells based on maximum entropy modeling, and find that growth rate fluctuations of E. coli reflect metabolic flux variability and growth sub-optimality, in turn highlighting information costs for growth optimization.
- Daniele De Martino
- , Anna MC Andersson
- & Gašper Tkačik
-
Article
| Open Access The topological requirements for robust perfect adaptation in networks of any size
The topological requirements for robust perfect adaptation in networks of any size
Robust perfect adaptation (RPA), the ability of a system to return to its pre-stimulus state in the presence of a new signal, enables organisms to respond to further changes in stimuli. Here, the authors identify the modular structure of the full set of network topologies that can confer RPA on complex networks.
- Robyn P. Araujo
- & Lance A. Liotta
-
Article
| Open Access Sign epistasis caused by hierarchy within signalling cascades
Sign epistasis caused by hierarchy within signalling cascades
Sign epistasis clearly constrains evolution, but its causes are difficult to decipher. Here, the authors study epistasis in a signalling cascade, and arrive at a general criterion and understanding of sign epistasis as arising from the inherent hierarchy between signalling cascade components.
- Philippe Nghe
- , Manjunatha Kogenaru
- & Sander J. Tans
-
Article
| Open Access Experimental noise cutoff boosts inferability of transcriptional networks in large-scale gene-deletion studies
Experimental noise cutoff boosts inferability of transcriptional networks in large-scale gene-deletion studies
Reliable inference of gene interactions from perturbation experiments remains a challenge. Here, the authors quantify the upper limits of transcriptional network inference from knockout screens, identify the key determinants of accuracy, and introduce an unbiased and scalable inference algorithm.
- C. F. Blum
- , N. Heramvand
- & M. Kollmann
-
Article
| Open Access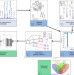 Network inference from glycoproteomics data reveals new reactions in the IgG glycosylation pathway
Network inference from glycoproteomics data reveals new reactions in the IgG glycosylation pathway
IgG glycosylation is an important factor in immune function, yet the molecular details of protein glycosylation remain poorly understood. The data-driven approach presented here uses large-scale plasma IgG mass spectrometry measurements to infer new biochemical reactions in the glycosylation pathway.
- Elisa Benedetti
- , Maja Pučić-Baković
- & Jan Krumsiek
-
Article
| Open Access The interdependent network of gene regulation and metabolism is robust where it needs to be
The interdependent network of gene regulation and metabolism is robust where it needs to be
Although networks of interacting genes and metabolic reactions are interdependent, they have largely been treated as separate systems. Here the authors apply a statistical framework for interdependent networks to E. coli, and show that it is sensitive to gene and protein perturbations but robust against metabolic changes.
- David F. Klosik
- , Anne Grimbs
- & Marc-Thorsten Hütt
-
Article
| Open Access The self-inhibitory nature of metabolic networks and its alleviation through compartmentalization
The self-inhibitory nature of metabolic networks and its alleviation through compartmentalization
Metabolites act as enzyme inhibitors, but their global impact on metabolism has scarcely been considered. Here, the authors generate a human genome-wide metabolite-enzyme inhibition network, and find that inhibition occurs largely due to limited structural diversity of metabolites, leading to a global constraint on metabolism which subcellular compartmentalization minimizes.
- Mohammad Tauqeer Alam
- , Viridiana Olin-Sandoval
- & Markus Ralser
-
Article
| Open Access An analytic approximation of the feasible space of metabolic networks
An analytic approximation of the feasible space of metabolic networks
Large-scale metabolic models of organisms from microbes to mammals can provide great insight into cellular function, but their analysis remains challenging. Here, the authors provide an approximate analytic method to estimate the feasible solution space for the flux vectors of metabolic networks, enabling more accurate analysis under a wide range of conditions of interest.
- Alfredo Braunstein
- , Anna Paola Muntoni
- & Andrea Pagnani
-
Article
| Open Access Reconstruction of the metabolic network of Pseudomonas aeruginosa to interrogate virulence factor synthesis
Reconstruction of the metabolic network of Pseudomonas aeruginosa to interrogate virulence factor synthesis
Targeting virulence rather than bacterial growth is less likely to select for antibiotic resistance, but many possible targets function in both processes. Here, the authors reconstruct a genome-scale metabolic network ofP. aeruginosastrain PA14 and update that of strain PAO1, which, together with mutant screens, enable them to identify genes uniquely critical for virulence factor production.
- Jennifer A. Bartell
- , Anna S. Blazier
- & Jason A. Papin
-
Article
| Open Access The OncoPPi network of cancer-focused protein–protein interactions to inform biological insights and therapeutic strategies
The OncoPPi network of cancer-focused protein–protein interactions to inform biological insights and therapeutic strategies
Understanding of dysregulation in cancers requires knowledge, beyond cancer genomes, of the interactions of cancer-associated proteins. Here, the authors use high-throughput, time-resolved FRET to map protein–protein interactions to establish a lung cancer protein network, and demonstrate its utility in revealing new oncogenic pathways and connectivity of tumour suppressors with druggable targets.
- Zenggang Li
- , Andrei A. Ivanov
- & Haian Fu
-
Article
| Open Access Boosting functionality of synthetic DNA circuits with tailored deactivation
Boosting functionality of synthetic DNA circuits with tailored deactivation
Nonlinearity in synthetic molecular circuits is usually achieved by manipulation of network topology or of production kinetics. Here, the authors achieve bistability and other nonlinear behaviours by manipulating the individual degradation rate laws of circuit components using saturable pathways.
- Kevin Montagne
- , Guillaume Gines
- & Yannick Rondelez
-
Article
| Open Access Multi-omic data integration enables discovery of hidden biological regularities
Multi-omic data integration enables discovery of hidden biological regularities
Translating omics data sets into biological insight is one of the great challenges of our time. Here, the authors make headway by synchronising pairs of omics data types via invariants across conditions and by integrating datasets into a genome-scale model of E. coli metabolism and gene expression.
- Ali Ebrahim
- , Elizabeth Brunk
- & Bernhard O. Palsson
-
Article
| Open Access A network property necessary for concentration robustness
A network property necessary for concentration robustness
Absolute concentration robustness (ACR), independence of the steady-state concentration of a molecule from the environment, is difficult to predict. Here, the authors derive a network structure-based necessary condition for ACR, and suggest that metabolites satisfying the condition are prevalent.
- Jeanne M. O. Eloundou-Mbebi
- , Anika Küken
- & Zoran Nikoloski
-
Article
| Open Access Proteome-wide association studies identify biochemical modules associated with a wing-size phenotype in Drosophila melanogaster
Proteome-wide association studies identify biochemical modules associated with a wing-size phenotype in Drosophila melanogaster
How genetic diversity generates complex phenotypes along a continuum remains a fundamental question of biology. Here—applying the emerging SWATH proteomics technology—the authors describe a proteome wide association study (PWAS) of Drosophila wing size and identify functional protein clusters associated with this trait.
- Hirokazu Okada
- , H. Alexander Ebhardt
- & Ernst Hafen
-
Article
| Open Access A rheostat mechanism governs the bifurcation of carbon flux in mycobacteria
A rheostat mechanism governs the bifurcation of carbon flux in mycobacteria
Microbes survive in dynamic environments by modulating their intracellular metabolism. Here, the authors reveal that mycobacteria employ a rheostat-like mechanism to regulate carbon flux between the oxidative TCA cycle and the glyoxylate shunt during glucose-acetate diauxic shift.
- Paul Murima
- , Michael Zimmermann
- & John D. McKinney
-
Article
| Open Access A geometrical approach to control and controllability of nonlinear dynamical networks
A geometrical approach to control and controllability of nonlinear dynamical networks
Complex networks, including physical, biological and social systems are ubiquitous, but understanding of how to control them is elusive. Here Wang et al. develop a framework based on the concept of attractor networks to facilitate the study of controllability of nonlinear dynamics in complex systems.
- Le-Zhi Wang
- , Ri-Qi Su
- & Ying-Cheng Lai
-
Article
| Open Access Reconstruction and topological characterization of the sigma factor regulatory network of Mycobacterium tuberculosis
Reconstruction and topological characterization of the sigma factor regulatory network of Mycobacterium tuberculosis
Sigma factors are regulatory proteins that reprogram the bacterial RNA polymerase in response to stress conditions to transcribe certain genes, including those for other sigma factors. Here, Chauhan et al. describe the complete sigma factor regulatory network of the pathogen Mycobacterium tuberculosis.
- Rinki Chauhan
- , Janani Ravi
- & Maria Laura Gennaro
-
Article
| Open Access Network-based in silico drug efficacy screening
Network-based in silico drug efficacy screening
Attempts to predict novel use for existing drugs rarely consider information on the impact on the genes perturbed in a given disease. Here, the authors present a novel network-based drug-disease proximity measure that provides insight on gene specific therapeutic effect of drugs and may facilitate drug repurposing.
- Emre Guney
- , Jörg Menche
- & Albert-László Barábasi
-
Article
| Open Access An exact arithmetic toolbox for a consistent and reproducible structural analysis of metabolic network models
An exact arithmetic toolbox for a consistent and reproducible structural analysis of metabolic network models
Current tools to analyse constraint-based models of metabolic networks have limited accuracy due to their use of floating-point arithmetic. Here the authors present MONGOOSE, a new computational tool that analyses such models in exact arithmetic, providing improved accuracy and reproducibility.
- Leonid Chindelevitch
- , Jason Trigg
- & Bonnie Berger
-
Article
| Open Access Identification of a human neonatal immune-metabolic network associated with bacterial infection
Identification of a human neonatal immune-metabolic network associated with bacterial infection
Infection remains a leading cause of morbidity and mortality in neonates worldwide. Here the authors report disproportionate immune stimulatory, co-inhibitory and metabolic pathway responses that specifically mark bacterial infection and can be used to predict sepsis in neonatal patients at the first clinical signs of infection.
- Claire L. Smith
- , Paul Dickinson
- & Peter Ghazal
-
Article |
 Predicting network functions with nested patterns
Predicting network functions with nested patterns
Identifying functionally important features of complex biological networks is computationally challenging. Ganter et al.develop a probabilistic framework that uses recurrent metabolite patterns to predict the properties and existence of reactions within a genome-scale metabolic network.
- Mathias Ganter
- , Hans-Michael Kaltenbach
- & Jörg Stelling
-
Article
| Open Access Existence of long-lasting experience-dependent plasticity in endocrine cell networks
Existence of long-lasting experience-dependent plasticity in endocrine cell networks
Experience-dependent plasticity and functional adaptation are thought to be restricted to the central nervous and immune systems. This study shows that long-lasting experience-dependent plasticity is a key feature of endocrine cell networks, allowing improved tissue function and hormone output following repeat demand.
- David J. Hodson
- , Marie Schaeffer
- & Patrice Mollard

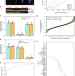 Spatial-fluxomics provides a subcellular-compartmentalized view of reductive glutamine metabolism in cancer cells
Spatial-fluxomics provides a subcellular-compartmentalized view of reductive glutamine metabolism in cancer cells