Featured
-
-
Article
| Open Access Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models
Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models
Experimental data on enzyme turnover numbers is sparse and noisy. Here, the authors use machine learning to successfully predict enzyme turnover numbers for E. coli, and show that using these to parameterize mechanistic genome-scale models enhances their predictive accuracy.
- David Heckmann
- , Colton J. Lloyd
- & Bernhard O. Palsson
-
Article
| Open Access Neural network analysis of sleep stages enables efficient diagnosis of narcolepsy
Neural network analysis of sleep stages enables efficient diagnosis of narcolepsy
The diagnosis of sleep disorders such as narcolepsy and insomnia currently requires experts to interpret sleep recordings (polysomnography). Here, the authors introduce a neural network analysis method for polysomnography that could reduce time spent in sleep clinics and automate narcolepsy diagnosis.
- Jens B. Stephansen
- , Alexander N. Olesen
- & Emmanuel Mignot
-
Article
| Open Access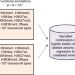 A semi-supervised approach for predicting cell-type specific functional consequences of non-coding variation using MPRAs
A semi-supervised approach for predicting cell-type specific functional consequences of non-coding variation using MPRAs
Predicting the functional consequences of non-coding genetic variants is a challenge. Here, He et al. present GenoNet, a semi-supervised method that combines information from experimentally confirmed regulatory variants with cell type- and tissue specific annotation for function prediction.
- Zihuai He
- , Linxi Liu
- & Iuliana Ionita-Laza
-
Article
| Open Access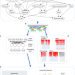 A web server for comparative analysis of single-cell RNA-seq data
A web server for comparative analysis of single-cell RNA-seq data
Publicly available single cell RNA-seq datasets represent valuable resources for comparative and meta-analysis. Here, the authors develop scQuery, a web server integrating over 500 different studies with over 300 unique cell types for comparative analysis of existing and new scRNA-seq data.
- Amir Alavi
- , Matthew Ruffalo
- & Ziv Bar-Joseph
-
Article
| Open Access Gene synthesis allows biologists to source genes from farther away in the tree of life
Gene synthesis allows biologists to source genes from farther away in the tree of life
Gene synthesis has expanded the ability to modify and create DNA sequences, with implications for biosurveillance. The authors use machine learning and codon theory to identify synthetic genes in Addgene data, and show that synthesis accelerates human-directed gene transfer across the tree of life.
- Aditya M. Kunjapur
- , Philipp Pfingstag
- & Neil C. Thompson
-
Article
| Open Access Machine learning and structural analysis of Mycobacterium tuberculosis pan-genome identifies genetic signatures of antibiotic resistance
Machine learning and structural analysis of Mycobacterium tuberculosis pan-genome identifies genetic signatures of antibiotic resistance
Mycobacterium tuberculosis exhibits complex evolution of antimicrobial resistance (AMR). Here, the authors perform machine learning and structural analysis to identify signatures of AMR evolution to 13 antibiotics.
- Erol S. Kavvas
- , Edward Catoiu
- & Bernhard O. Palsson
-
Article
| Open Access Genetic signature to provide robust risk assessment of psoriatic arthritis development in psoriasis patients
Genetic signature to provide robust risk assessment of psoriatic arthritis development in psoriasis patients
Approximately 30% of psoriasis patients develop psoriatic arthritis (PsA) and early diagnosis is crucial for the management of PsA. Here, Patrick et al. develop a computational pipeline involving statistical and machine-learning methods that can assess the risk of progression to PsA based on genetic markers.
- Matthew T. Patrick
- , Philip E. Stuart
- & Lam C. Tsoi
-
Article
| Open Access Refined efficacy estimates of the Sanofi Pasteur dengue vaccine CYD-TDV using machine learning
Refined efficacy estimates of the Sanofi Pasteur dengue vaccine CYD-TDV using machine learning
Clinical trials for the CYD-TDV dengue vaccine showed that vaccine efficacy varies with prior dengue exposure, but baseline serostatus is only known for 12% of subjects. Here, Dorigatti et al. use machine learning to impute baseline serostatus and determine vaccine efficacy by baseline serostatus, age and dengue serotype.
- I. Dorigatti
- , C. A. Donnelly
- & N. M. Ferguson
-
Article
| Open Access microCLIP super learning framework uncovers functional transcriptome-wide miRNA interactions
microCLIP super learning framework uncovers functional transcriptome-wide miRNA interactions
AGO-PAR-CLIP is widely used for high-throughput miRNA target characterization. Here, the authors show that the previously neglected non-T-to-C clusters denote functional miRNA binding events, and develop microCLIP, a super learning framework that accurately detects miRNA interactions.
- Maria D. Paraskevopoulou
- , Dimitra Karagkouni
- & Artemis G. Hatzigeorgiou
-
Article
| Open Access Systematic identification of non-coding pharmacogenomic landscape in cancer
Systematic identification of non-coding pharmacogenomic landscape in cancer
In this study the authors build lncRNA-drug response models for 265 anti-cancer agents across 27 cancer types. They report their cancer cell line based lncRNA EN-models are able to effectively predict therapeutic outcome for breast cancer, ovarian cancer, endometrial cancer and stomach cancer patients.
- Yue Wang
- , Zehua Wang
- & Da Yang
-
Article
| Open Access Deep learning to predict the lab-of-origin of engineered DNA
Deep learning to predict the lab-of-origin of engineered DNA
The synthetic biology era has seen a rapidly growing number of engineered DNA sequences. Here, the authors develop a deep learning method to predict the lab-of-origin of a DNA sequence based on hidden design signatures.
- Alec A. K. Nielsen
- & Christopher A. Voigt
-
Article
| Open Access Network enhancement as a general method to denoise weighted biological networks
Network enhancement as a general method to denoise weighted biological networks
Technical noise in experiments is unavoidable, but it introduces inaccuracies into the biological networks we infer from the data. Here, the authors introduce a diffusion-based method for denoising undirected, weighted networks, and show that it improves the performances of downstream analyses.
- Bo Wang
- , Armin Pourshafeie
- & Jure Leskovec
-
Article
| Open Access An effect of serotonergic stimulation on learning rates for rewards apparent after long intertrial intervals
An effect of serotonergic stimulation on learning rates for rewards apparent after long intertrial intervals
Serotonin (5-HT) plays many important roles in reward, punishment, patience and beyond, and optogenetic stimulation of 5-HT neurons has not crisply parsed them. The authors report a novel analysis of a reward-based decision-making experiment, and show that 5-HT stimulation increases the learning rate, but only on a select subset of choices.
- Kiyohito Iigaya
- , Madalena S. Fonseca
- & Peter Dayan
-
Article
| Open Access Uncovering pseudotemporal trajectories with covariates from single cell and bulk expression data
Uncovering pseudotemporal trajectories with covariates from single cell and bulk expression data
Cross-sectional omic data often have non-homogeneous genetic, phenotypic, or environmental backgrounds. Here, the authors develop a statistical framework to infer pseudotime trajectories in the presence of such factors as well as their interactions in both single-cell and bulk gene expression analysis
- Kieran R Campbell
- & Christopher Yau
-
Article
| Open Access Scalable training of artificial neural networks with adaptive sparse connectivity inspired by network science
Scalable training of artificial neural networks with adaptive sparse connectivity inspired by network science
Artificial neural networks are artificial intelligence computing methods which are inspired by biological neural networks. Here the authors propose a method to design neural networks as sparse scale-free networks, which leads to a reduction in computational time required for training and inference.
- Decebal Constantin Mocanu
- , Elena Mocanu
- & Antonio Liotta
-
Article
| Open Access Exploring patterns enriched in a dataset with contrastive principal component analysis
Exploring patterns enriched in a dataset with contrastive principal component analysis
Dimensionality reduction and visualization methods lack a principled way of comparing multiple datasets. Here, Abid et al. introduce contrastive PCA, which identifies low-dimensional structures enriched in one dataset compared to another and enables visualization of dataset-specific patterns.
- Abubakar Abid
- , Martin J. Zhang
- & James Zou
-
Article
| Open Access Interpretable dimensionality reduction of single cell transcriptome data with deep generative models
Interpretable dimensionality reduction of single cell transcriptome data with deep generative models
Although single-cell transcriptome data are increasingly available, their interpretation remains a challenge. Here, the authors present a dimensionality reduction approach that preserves both the local and global neighbourhood structures in the data thus enhancing its interpretability.
- Jiarui Ding
- , Anne Condon
- & Sohrab P. Shah
-
Article
| Open Access Deconvolution of subcellular protrusion heterogeneity and the underlying actin regulator dynamics from live cell imaging
Deconvolution of subcellular protrusion heterogeneity and the underlying actin regulator dynamics from live cell imaging
Cell protrusion dynamics are heterogeneous at the subcellular level, but current analyses operate at the cellular or ensemble level. Here the authors develop a computational framework to quantify subcellular protrusion phenotypes and reveal the underlying actin regulator dynamics at the leading edge.
- Chuangqi Wang
- , Hee June Choi
- & Kwonmoo Lee
-
Article
| Open Access A geometric approach to characterize the functional identity of single cells
A geometric approach to characterize the functional identity of single cells
Functional characterisation of single cells is crucial for uncovering the true extent of cellular heterogeneity. Here the authors offer an approach to infer functional identities of cells from their transcriptomes, identify their dominant function, and reconstruct the underlying regulatory networks.
- Shahin Mohammadi
- , Vikram Ravindra
- & Ananth Grama
-
Article
| Open Access PREDICTD PaRallel Epigenomics Data Imputation with Cloud-based Tensor Decomposition
PREDICTD PaRallel Epigenomics Data Imputation with Cloud-based Tensor Decomposition
Assays to characterize the epigenome and interrogate chromatin state genome wide have so far been performed in a selected set of conditions. Here, Durham et al. develop a computational method based on tensor decomposition to impute missing experiments in collections of epigenomics experiments.
- Timothy J. Durham
- , Maxwell W. Libbrecht
- & William Stafford Noble
-
Article
| Open Access The effects of death and post-mortem cold ischemia on human tissue transcriptomes
The effects of death and post-mortem cold ischemia on human tissue transcriptomes
RNA levels in post-mortem tissue can differ greatly from those before death. Studying the effect of post-mortem interval on the transcriptome in 36 human tissues, Ferreira et al. find that the response to death is largely tissue-specific and develop a model to predict time since death based on RNA data.
- Pedro G. Ferreira
- , Manuel Muñoz-Aguirre
- & Roderic Guigó
-
Article
| Open Access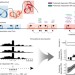 Transcriptional decomposition reveals active chromatin architectures and cell specific regulatory interactions
Transcriptional decomposition reveals active chromatin architectures and cell specific regulatory interactions
Transcriptional regulation is coupled with chromosomal positioning and chromatin architecture. Here the authors develop a transcriptional decomposition approach to separate expression associated with genome structure from independent effects not directly associated with genomic positioning.
- Sarah Rennie
- , Maria Dalby
- & Robin Andersson
-
Article
| Open Access Intelligent image-based in situ single-cell isolation
Intelligent image-based in situ single-cell isolation
The isolation of single cells while retaining context is important for quantifying cellular heterogeneity but technically challenging. Here, the authors develop a high-throughput, scalable workflow for microscopy-based single cell isolation using machine-learning, high-throughput microscopy and laser capture microdissection.
- Csilla Brasko
- , Kevin Smith
- & Peter Horvath
-
Article
| Open Access A machine learning approach to integrate big data for precision medicine in acute myeloid leukemia
A machine learning approach to integrate big data for precision medicine in acute myeloid leukemia
Identification of markers of drug response is essential for precision therapy. Here the authors introduce an algorithm that uses prior information about each gene’s importance in AML to identify the most predictive gene-drug associations from transcriptome and drug response data from 30 AML samples.
- Su-In Lee
- , Safiye Celik
- & Pamela S. Becker
-
Article
| Open Access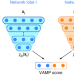 VAMPnets for deep learning of molecular kinetics
VAMPnets for deep learning of molecular kinetics
Extracting kinetic models from high-throughput molecular dynamics (MD) simulations is laborious and prone to human error. Here the authors introduce a deep learning framework that automates construction of Markov state models from MD simulation data.
- Andreas Mardt
- , Luca Pasquali
- & Frank Noé
-
Article
| Open Access Model-free inference of direct network interactions from nonlinear collective dynamics
Model-free inference of direct network interactions from nonlinear collective dynamics
Network dynamical systems can represent the interactions involved in the collective dynamics of gene regulatory networks or metabolic circuits. Here Casadiego et al. present a method for inferring these types of interactions directly from observed time series without relying on their model.
- Jose Casadiego
- , Mor Nitzan
- & Marc Timme
-
Article
| Open Access Identifying host regulators and inhibitors of liver stage malaria infection using kinase activity profiles
Identifying host regulators and inhibitors of liver stage malaria infection using kinase activity profiles
Host kinases facilitate Plasmodium liver stage (LS) infection, but systematic accounting of important players is lacking. Here, the authors use a computational approach and kinase activity profiles to identify host kinase regulators of LS infection and drugs that could eliminate parasite burden.
- Nadia Arang
- , Heather S. Kain
- & Alexis Kaushansky
-
Article
| Open Access A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
Network-based data integration for drug–target prediction is a promising avenue for drug repositioning, but performance is wanting. Here, the authors introduce DTINet, whose performance is enhanced in the face of noisy, incomplete and high-dimensional biological data by learning low-dimensional vector representations.
- Yunan Luo
- , Xinbin Zhao
- & Jianyang Zeng
-
Article
| Open Access Reconstructing cell cycle and disease progression using deep learning
Reconstructing cell cycle and disease progression using deep learning
The interpretation of information-rich, high-throughput single-cell data is a challenge requiring sophisticated computational tools. Here the authors demonstrate a deep convolutional neural network that can classify cell cycle status on-the-fly.
- Philipp Eulenberg
- , Niklas Köhler
- & F. Alexander Wolf
-
Article
| Open Access Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Global patterns of gene transcription can be represented with reduced dimensionality. Here, the authors devise a method called Tradict that learns and uses 100 marker genes to predict transcriptome-wide pathway expression levels and patterns that reflect cell activity and state.
- Surojit Biswas
- , Konstantin Kerner
- & Philip A. Wigge
-
Article
| Open Access Sensitive detection of rare disease-associated cell subsets via representation learning
Sensitive detection of rare disease-associated cell subsets via representation learning
While rare cell subpopulations frequently make the difference between health and disease, their detection remains a challenge. Here, the authors devise CellCnn, a representation learning approach to detecting such rare cell populations from high-dimensional single cell data, and, among other examples, demonstrate its capacity for detecting rare leukaemic blasts in minimal residual disease.
- Eirini Arvaniti
- & Manfred Claassen
-
Article
| Open Access Crowdsourced assessment of common genetic contribution to predicting anti-TNF treatment response in rheumatoid arthritis
Crowdsourced assessment of common genetic contribution to predicting anti-TNF treatment response in rheumatoid arthritis
Rheumatoid arthritis patients respond differently to anti-TNF treatment. Using community-based challenge, the authors show that currently available data does not reveal meaningful genetic predictors of response to anti-TNF therapy, thus confirming clinical observations.
- Solveig K. Sieberts
- , Fan Zhu
- & Lara M. Mangravite
-
Article
| Open Access Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features
Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features
Diagnosis of lung cancer through manual histopathology evaluation is insufficient to predict patient survival. Here, the authors use computerized image processing to identify diagnostically relevant image features and use these features to distinguish lung cancer patients with different prognoses.
- Kun-Hsing Yu
- , Ce Zhang
- & Michael Snyder
-
Article
| Open Access The ligand binding mechanism to purine nucleoside phosphorylase elucidated via molecular dynamics and machine learning
The ligand binding mechanism to purine nucleoside phosphorylase elucidated via molecular dynamics and machine learning
Understanding the dynamics of enzyme-substrate complexation provides an insight into potential drugs, but intermediate states are difficult to observe experimentally. Here, the authors use simulations and machine learning to analyse the binding of transition state inhibitors to purine nucleoside phosphorylase.
- Sergio Decherchi
- , Anna Berteotti
- & Andrea Cavalli
-
Article |
 microTSS: accurate microRNA transcription start site identification reveals a significant number of divergent pri-miRNAs
microTSS: accurate microRNA transcription start site identification reveals a significant number of divergent pri-miRNAs
microRNAs are short non-coding RNAs that post-transcriptionally regulate gene expression for which the identification of promoter and primary transcripts (pri-miRNAs) has been difficult. Here the authors describe microTSS, an algorithm that supports the precise identification of intergenic pri-miRNA transcription start sites.
- Georgios Georgakilas
- , Ioannis S. Vlachos
- & Artemis G. Hatzigeorgiou

 A machine learning approach for online automated optimization of super-resolution optical microscopy
A machine learning approach for online automated optimization of super-resolution optical microscopy