Featured
-
-
Article
| Open Access Diploid and tetraploid genomes of Acorus and the evolution of monocots
Diploid and tetraploid genomes of Acorus and the evolution of monocots
Acorales is sister to all other monocots and contains only one family with just one genus, Acorus. Here, the authors assemble the genome of the diploid Ac. gramineus and the tetraploid Ac. calamus, reconstruct an ancestral monocot karyotype and gene toolkit, and discuss the origin and evolution of the two species and other monocots.
- Liang Ma
- , Ke-Wei Liu
- & Zhong-Jian Liu
-
Article
| Open Access The genome of Acorus deciphers insights into early monocot evolution
The genome of Acorus deciphers insights into early monocot evolution
Monocots are one of the most diverse and dominant clades of flowering plants. Here, the authors assemble the genome of Acorus gramineus, confirm its phylogenetic position as sister to the rest of monocots and reveal the absence of tau (τ) whole-genome duplication observed in the majority of monocot clades.
- Xing Guo
- , Fang Wang
- & Huan Liu
-
Article
| Open Access Phylodynamic of SARS-CoV-2 during the second wave of COVID-19 in Peru
Phylodynamic of SARS-CoV-2 during the second wave of COVID-19 in Peru
The second SARS-CoV-2 wave in Peru had a high case fatality rate with Lambda and Gamma causing most cases. Using phylodynamics, the authors here show that Lambda most likely originated in Peru from where it spread to other South American countries and that the center of Peru played a key role in transmission to other regions.
- Santiago Justo Arevalo
- , Carmen Sofia Uribe Calampa
- & Joao Renato Rebello Pinho
-
Article
| Open Access The monoaminergic system is a bilaterian innovation
The monoaminergic system is a bilaterian innovation
Monoamines act as neuromodulators in the nervous system, but their evolutionary origins are unclear. Here, the authors examine the evolution of genes involved in monoamine production, and processing suggesting that the monoaminergic system evolved in the bilaterian stem-group.
- Matthew Goulty
- , Gaelle Botton-Amiot
- & Roberto Feuda
-
Article
| Open Access Independent rediploidization masks shared whole genome duplication in the sturgeon-paddlefish ancestor
Independent rediploidization masks shared whole genome duplication in the sturgeon-paddlefish ancestor
Whole genome duplication can generate new genes and support survival through mass extinctions. Here, the authors show that paddlefish and sturgeon shared a genome duplication event 200 million years ago that was previously unrecognised due to the mixed signals from independent rediploidisation.
- Anthony K. Redmond
- , Dearbhaile Casey
- & Aoife McLysaght
-
Article
| Open Access Resurgence of Omicron BA.2 in SARS-CoV-2 infection-naive Hong Kong
Resurgence of Omicron BA.2 in SARS-CoV-2 infection-naive Hong Kong
Hong Kong experienced a large wave of COVID-19 in early 2022 driven by Omicron BA.2. Here, the authors describe the epidemiological dynamics of this wave and show discordant inferences based on genomic and epidemiological data that underscore the need to improve near real-time epidemic growth estimates.
- Ruopeng Xie
- , Kimberly M. Edwards
- & Vijaykrishna Dhanasekaran
-
Article
| Open Access The genomic epidemiology of Escherichia albertii infecting humans and birds in Great Britain
The genomic epidemiology of Escherichia albertii infecting humans and birds in Great Britain
Escherichia albertii is an emerging gastrointestinal pathogen that causes disease in humans and animals, notably birds. In this genomic epidemiology study, the authors investigate characteristics of isolates sampled from humans and birds in Great Britain and find that they tend to cluster separately.
- Rebecca J. Bengtsson
- , Kate S. Baker
- & Becki Lawson
-
Article
| Open Access Molecular exploration of fossil eggshell uncovers hidden lineage of giant extinct bird
Molecular exploration of fossil eggshell uncovers hidden lineage of giant extinct bird
The evolution and systematics of Madagascar’s extinct elephant birds remains unclear. Here, the authors recover genetic, stable isotope, morphological, and geographic data from fossil eggshell to describe variation among clades, identifying cryptic diversity and potential drivers of speciation.
- Alicia Grealy
- , Gifford H. Miller
- & Michael Bunce
-
Article
| Open Access Bacterial origins of thymidylate metabolism in Asgard archaea and Eukarya
Bacterial origins of thymidylate metabolism in Asgard archaea and Eukarya
Asgard archaea include the closest known archaeal relatives of eukaryotes. Here, the authors provide evidence that eukaryotic and Asgard thymidylate synthases (required for DNA synthesis) may have a bacterial origin, and additional lateral transfer of bacterial genes may have shaped the metabolism of Asgard archaea.
- Jonathan Filée
- , Hubert F. Becker
- & Hannu Myllykallio
-
Article
| Open Access A latitudinal gradient of deep-sea invasions for marine fishes
A latitudinal gradient of deep-sea invasions for marine fishes
This study finds that high-latitude fish clades with the fastest speciation rates also exhibit elevated rates of depth evolution, creating a prevailing latitudinal gradient of deep-sea invasions concentrated in poleward regions. These results advance our understanding of how niche lability and climate shape global patterns of species distributions.
- Sarah T. Friedman
- & Martha M. Muñoz
-
Matters Arising
| Open Access Reply to: Available data do not rule out Ctenophora as the sister group to all other Metazoa
Reply to: Available data do not rule out Ctenophora as the sister group to all other Metazoa
- Anthony K. Redmond
- & Aoife McLysaght
-
Matters Arising
| Open Access Available data do not rule out Ctenophora as the sister group to all other Metazoa
Available data do not rule out Ctenophora as the sister group to all other Metazoa
- Nathan V. Whelan
- & Kenneth M. Halanych
-
Article
| Open Access Macroevolutionary diversity of traits and genomes in the model yeast genus Saccharomyces
Macroevolutionary diversity of traits and genomes in the model yeast genus Saccharomyces
Here, the authors describe the geographies, hosts, substrates, and phylogenetic relationships for 1,794 Saccharomyces strains. They provide insight into the genetic and phenotypic diversity in the genus, not seen through prior work focused on the model species Saccharomyces cerevisiae.
- David Peris
- , Emily J. Ubbelohde
- & Chris Todd Hittinger
-
Article
| Open Access Phylogeography and transmission of Mycobacterium tuberculosis spanning prisons and surrounding communities in Paraguay
Phylogeography and transmission of Mycobacterium tuberculosis spanning prisons and surrounding communities in Paraguay
To role that carceral institutions play in Mycobacterium tuberculosis transmission remains somewhat unknown. Authors perform a prospective genomic surveillance study, to assess transmission dynamics in prisons and surrounding communities in Paraguay.
- Gladys Estigarribia Sanabria
- , Guillermo Sequera
- & Katharine S. Walter
-
Article
| Open Access The macroevolutionary impact of recent and imminent mammal extinctions on Madagascar
The macroevolutionary impact of recent and imminent mammal extinctions on Madagascar
Madagascar is a threatened biodiversity hotspot. Here, using a newly assembled dataset and island biogeography models, the authors estimate how many millions of years of evolutionary history have been lost since human colonisation and may be further lost in the future for Malagasy mammals.
- Nathan M. Michielsen
- , Steven M. Goodman
- & Luis Valente
-
Article
| Open Access Ruminant inner ear shape records 35 million years of neutral evolution
Ruminant inner ear shape records 35 million years of neutral evolution
External ecological interactions and intrinsic biological parameters affect evolutionary pathways and animal diversity. Here, the authors use ruminant inner ear morphology to investigate patterns of diversity through 33 million years, finding clade-dependent climate and paleogeographic trends.
- Bastien Mennecart
- , Laura Dziomber
- & Loïc Costeur
-
Article
| Open Access Evolutionary origins of the prolonged extant squamate radiation
Evolutionary origins of the prolonged extant squamate radiation
Here, the authors present two well preserved fossil lizard skulls from the Late Jurassic of North America. These fossils, placed at the base of the clade Pan-Scincoidea, suggest that squamates had a wide geographic distribution and preserve characteristics that show the complex early evolutionary history of squamate anatomy.
- Chase D. Brownstein
- , Dalton L. Meyer
- & Jacques A. Gauthier
-
Article
| Open Access The evolution of reproductive modes and life cycles in amphibians
The evolution of reproductive modes and life cycles in amphibians
Here, the authors use reproductive mode data with matching phylogenetic data to explore the evolution of reproductive mode, transitions between reproductive modes, and diversification rates in amphibians.
- H. Christoph Liedtke
- , John J. Wiens
- & Ivan Gomez-Mestre
-
Article
| Open Access Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny
Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny
Multiple sequence alignments are widely used to predict protein structure, function, and phylogeny, but are uncertain with more diverged sequences. Muscle5 generates ensembles of alternative high-accurate alignments, enabling novel confidence estimates in alignments, trees, and other inferences.
- Robert C. Edgar
-
Article
| Open Access Ordovician opabiniid-like animals and the role of the proboscis in euarthropod head evolution
Ordovician opabiniid-like animals and the role of the proboscis in euarthropod head evolution
Here, the authors describe two opabiniid-like euarthropods with anterior proboscises from the Middle Ordovician Castle Bank Biota, Wales, UK. Phylogenetic analysis suggests that these specimens may be sister to radiodonts and deuteropods.
- Stephen Pates
- , Joseph P. Botting
- & Joanna M. Wolfe
-
Article
| Open Access Fertilization mode differentially impacts the evolution of vertebrate sperm components
Fertilization mode differentially impacts the evolution of vertebrate sperm components
The location where fertilization takes place can be highly variable across species, and especially between internal and external fertilizers. Kahrl et al. find that fertilization environment plays a significant role in the evolution and diversification of sperm morphology across vertebrate species.
- Ariel F. Kahrl
- , Rhonda R. Snook
- & John L. Fitzpatrick
-
Article
| Open Access Past and present giant viruses diversity explored through permafrost metagenomics
Past and present giant viruses diversity explored through permafrost metagenomics
Although giant viruses are abundant in aquatic environments, less is known about giant viruses in soil. Here, the authors use permafrost metagenomics to reveal giant virus diversity and heterogeneity, as well as gene transfers between viruses from different families.
- Sofia Rigou
- , Sébastien Santini
- & Matthieu Legendre
-
Article
| Open Access Predicting the evolution of the Lassa virus endemic area and population at risk over the next decades
Predicting the evolution of the Lassa virus endemic area and population at risk over the next decades
It is currently unknown how climate and land use changes could affect the endemic area of Lassa virus, a zoonotic pathogen responsible for Lassa fever. Here, the authors show that by 2070, new regions in Africa will likely become ecologically suitable for Lassa virus, drastically increasing the population living in conditions favourable for virus circulation.
- Raphaëlle Klitting
- , Liana E. Kafetzopoulou
- & Simon Dellicour
-
Article
| Open Access Using multiple sampling strategies to estimate SARS-CoV-2 epidemiological parameters from genomic sequencing data
Using multiple sampling strategies to estimate SARS-CoV-2 epidemiological parameters from genomic sequencing data
SARS-CoV-2 genome sequencing data can be used to infer epidemiological parameters, but the impact of the strategy used to select samples on these estimates is rarely considered. Here, the authors produce estimates using different sampling strategies and compare results to those based on case reporting data.
- Rhys P. D. Inward
- , Kris V. Parag
- & Nuno R. Faria
-
Article
| Open Access Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Here, the authors develop software that uses photographs of birds to extract information on plumage UV reflectance. They use these data to show that UV reflectance is phylogenetically conserved and associated with the light environment.
- Yichen He
- , Zoë K. Varley
- & Christopher R. Cooney
-
Article
| Open Access Regional connectivity drove bidirectional transmission of SARS-CoV-2 in the Middle East during travel restrictions
Regional connectivity drove bidirectional transmission of SARS-CoV-2 in the Middle East during travel restrictions
The dynamics of SARS-CoV-2 transmission in the Middle East have been relatively under-studied. Here, the authors integrate genomic and travel data and show that introductions to the region were initially driven by intercontinental air travel, after which regional land travel became a more important driver.
- Edyth Parker
- , Catelyn Anderson
- & Issa Abu-Dayyeh
-
Article
| Open Access Dynamics of competing SARS-CoV-2 variants during the Omicron epidemic in England
Dynamics of competing SARS-CoV-2 variants during the Omicron epidemic in England
This study presents data from the REACT-1 SARS-CoV-2 community sampling study in England from November 2021 to March 2022. They show that the Omicron variant peaked in January with a prevalence of ~7% and that the BA.2 sublineage had a 1.5x higher reproduction number compared to other Omicron sublineages.
- Oliver Eales
- , Leonardo de Oliveira Martins
- & Marc Chadeau-Hyam
-
Article
| Open Access A Bayesian approach to infer recombination patterns in coronaviruses
A Bayesian approach to infer recombination patterns in coronaviruses
Genetic recombination can confound standard phylogenetic approaches. Here, the authors present a method to reconstruct virus recombination networks, and show the importance of recombination in shaping the ongoing evolution of SARS-like, MERS and 3 human seasonal coronaviruses.
- Nicola F. Müller
- , Kathryn E. Kistler
- & Trevor Bedford
-
Article
| Open Access Recovery of Lutacidiplasmatales archaeal order genomes suggests convergent evolution in Thermoplasmatota
Recovery of Lutacidiplasmatales archaeal order genomes suggests convergent evolution in Thermoplasmatota
Genome recovery of the order Lutacidiplasmatales reveals that key genes for environmental specialisation were acquired multiple times in the Thermoplasmatota phylum, suggesting a crucial role of convergent evolution in archaeal habitat adaptation.
- Paul O. Sheridan
- , Yiyu Meng
- & Cécile Gubry-Rangin
-
Article
| Open Access Detection of SARS-CoV-2 intra-host recombination during superinfection with Alpha and Epsilon variants in New York City
Detection of SARS-CoV-2 intra-host recombination during superinfection with Alpha and Epsilon variants in New York City
Here, the authors characterize a case of SARS-CoV-2 superinfection with Alpha and Epsilon variants, in which, via full genome sequencing analyses, they identify recombinant haplotypes in the spike, nucleocapsid, and ORF 8 coding regions, suggesting recombination could play a role in SARS-CoV-2 genetic diversity.
- Joel O. Wertheim
- , Jade C. Wang
- & Scott Hughes
-
Article
| Open Access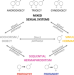 Switches, stability and reversals in the evolutionary history of sexual systems in fish
Switches, stability and reversals in the evolutionary history of sexual systems in fish
Fish have a diversity of sexual systems. Pla et al. analyse the transitions in these systems across fish, supporting that simultaneous hermaphroditism cannot evolve directly from separate sexes but requires sequential hermaphroditism as an intermediate step.
- Susanna Pla
- , Chiara Benvenuto
- & Francesc Piferrer
-
Article
| Open Access Tempo and drivers of plant diversification in the European mountain system
Tempo and drivers of plant diversification in the European mountain system
Here, the authors use full-plastome phylogenomics and multiclade comparative models to reconstruct the tempo and drivers of six European Alpine angiosperm lineages before and during the Pleistocene. They find that geographic divergence and bedrock shifts drive speciation events, while diversification rates remained steady.
- Jan Smyčka
- , Cristina Roquet
- & Sébastien Lavergne
-
Article
| Open Access Genomic diversity across the Rickettsia and ‘Candidatus Megaira’ genera and proposal of genus status for the Torix group
Genomic diversity across the Rickettsia and ‘Candidatus Megaira’ genera and proposal of genus status for the Torix group
The bacterial genus Rickettsia includes vector-borne pathogens and arthropod symbionts that are close relatives of symbionts of microeukaryotes classified under the genus ‘Candidatus Megaira’. Here, Davison et al. clarify the evolutionary relationships between these organisms by assembling 28 genomes of understudied species, and propose that a distinct clade known as Torix Rickettsia should be considered a separate genus.
- Helen R. Davison
- , Jack Pilgrim
- & Stefanos Siozios
-
Article
| Open Access Genetic diversity of SARS-CoV-2 infections in Ghana from 2020-2021
Genetic diversity of SARS-CoV-2 infections in Ghana from 2020-2021
In this genomic epidemiology study from Ghana, the authors sequence ~1,000 SARS-CoV-2 whole genomes from March 2020 to September 2021. They describe changes in the predominant circulating lineages over time and infer how variants of concern were likely introduced into the country.
- Collins M. Morang’a
- , Joyce M. Ngoi
- & Gordon A. Awandare
-
Article
| Open Access Phylotranscriptomic insights into a Mesoproterozoic–Neoproterozoic origin and early radiation of green seaweeds (Ulvophyceae)
Phylotranscriptomic insights into a Mesoproterozoic–Neoproterozoic origin and early radiation of green seaweeds (Ulvophyceae)
“Ulvophyceae is a remarkably morphologically and ecologically diverse clade of green algae. Here, the authors reconstruct the Ulvophyceae phylogeny, showing that these algae originated earlier than expected and may have influenced biogeochemical cycles at the Mesoproterozoic-Neoproterozoic transition.”
- Zheng Hou
- , Xiaoya Ma
- & Bojian Zhong
-
Article
| Open Access Phylogenomic analyses highlight innovation and introgression in the continental radiations of Fagaceae across the Northern Hemisphere
Phylogenomic analyses highlight innovation and introgression in the continental radiations of Fagaceae across the Northern Hemisphere
Fagaceae are diverse family including trees of ecological and economic importance. This phylogenomic analysis of nuclear and plastid genomes reconstructs evolutionary history and finds evidence of multiple adaptive introgression events in this important plant family.
- Biao-Feng Zhou
- , Shuai Yuan
- & Baosheng Wang
-
Article
| Open Access Recurring adaptive introgression of a supergene variant that determines social organization
Recurring adaptive introgression of a supergene variant that determines social organization
Solenopsis fire ants have a polymorphic social system in which some colonies have multiple queens. Here, Stolle, Pracana et al. show that the supergene that produces the multiple-queen phenotype has spread repeatedly between Solenopsis species by introgression.
- Eckart Stolle
- , Rodrigo Pracana
- & Yannick Wurm
-
Article
| Open Access Fossil coleoid cephalopod from the Mississippian Bear Gulch Lagerstätte sheds light on early vampyropod evolution
Fossil coleoid cephalopod from the Mississippian Bear Gulch Lagerstätte sheds light on early vampyropod evolution
The authors describe a new cephalopod from the Carboniferous (Mississippian) Bear Gulch Lagerstätte of Montana, USA. This specimen extends the fossil record of vampyropods back by ~82 million years and changes our understanding of their evolution.
- Christopher D. Whalen
- & Neil H. Landman
-
Article
| Open Access Antibody escape and global spread of SARS-CoV-2 lineage A.27
Antibody escape and global spread of SARS-CoV-2 lineage A.27
The A.27 SARS-CoV-2 lineage spread globally in 2021 but did not become dominant. Here, the authors show that A.27 shares some mutations in the spike gene that are present in variants of concern, but lacks the D614G mutation, indicating independent evolution of immune escape properties.
- Tamara Kaleta
- , Lisa Kern
- & Jonas Fuchs
-
Article
| Open Access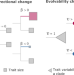 General statistical model shows that macroevolutionary patterns and processes are consistent with Darwinian gradualism
General statistical model shows that macroevolutionary patterns and processes are consistent with Darwinian gradualism
‘Macroevolution posed difficulties for Darwin and later theorists because species frequently change abruptly, or experience long periods of stasis, both counter to the theory of incremental change or gradualism. Here, the authors propose a macroevolutionary statistical model that accommodates this uneven evolutionary landscape, and shows how even abrupt macroevolutionary changes are compatible with gradualist microevolutionary processes.’
- Mark Pagel
- , Ciara O’Donovan
- & Andrew Meade
-
Article
| Open Access Genomic epidemiology of SARS-CoV-2 under an elimination strategy in Hong Kong
Genomic epidemiology of SARS-CoV-2 under an elimination strategy in Hong Kong
Hong Kong has used an elimination strategy to control SARS-CoV-2 with stringent measures including traveller quarantine. Here, the authors show that the majority of community-acquired cases until January 2021 resulted from three importations, and that increased transmission followed prolonged periods of restrictions, likely due to adherence fatigue.
- Haogao Gu
- , Ruopeng Xie
- & Leo L. M. Poon
-
Article
| Open Access Buxus and Tetracentron genomes help resolve eudicot genome history
Buxus and Tetracentron genomes help resolve eudicot genome history
Gamma triplication arises via two whole-genome duplications early in eudicot history, but the relative timing of these is unclear. Here, the authors report the genomes of Buxales and Trochodendrales and reject the hypothesis of gamma arising via inter-lineage hybridization between ancestral eudicot lineages.
- Andre S. Chanderbali
- , Lingling Jin
- & Pamela S. Soltis
-
Article
| Open Access Neuroanatomy in a middle Cambrian mollisoniid and the ancestral nervous system organization of chelicerates
Neuroanatomy in a middle Cambrian mollisoniid and the ancestral nervous system organization of chelicerates
Here, the authors report a preserved central nervous system in the soft-bodied stem-group chelicerate Mollisonia symmetrica from the mid-Cambrian Burgess Shale. The neuroanatomy described here is proposed to represent the ancestral state for the stem group Chelicerata.
- Javier Ortega-Hernández
- , Rudy Lerosey-Aubril
- & James C. Weaver
-
Article
| Open Access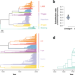 Genomic signatures of pre-resistance in Mycobacterium tuberculosis
Genomic signatures of pre-resistance in Mycobacterium tuberculosis
Signals of antimicrobial resistance in pathogen genomes may be detectable before the organism evolves an antimicrobial resistance phenotype. Here, the authors investigate this hypothesis using Mycobacterium tuberculosis data from Peru and identify candidate “pre-resistance” markers.
- Arturo Torres Ortiz
- , Jorge Coronel
- & Louis Grandjean
-
Article
| Open Access The Chloranthus sessilifolius genome provides insight into early diversification of angiosperms
The Chloranthus sessilifolius genome provides insight into early diversification of angiosperms
Chloranthales remain the last lineage of core angiosperms that lacks a nuclear genome assembly. Here, the authors report the genome assembly of Chloranthus sesilifolius and show that both hybridization and incomplete lineage sorting may have contributed to the phylogenetic incongruities in the literature.
- Jianxiang Ma
- , Pengchuan Sun
- & Yongzhi Yang
-
Article
| Open Access Chloranthus genome provides insights into the early diversification of angiosperms
Chloranthus genome provides insights into the early diversification of angiosperms
Chloranthales remain the last lineage of core angiosperms that lacks a nuclear genome assembly. Here, the authors report the genome assembly of Chloranthus spicatus and show its contribution to deepen our understanding on diversification, phylogeny, and genome evolution in angiosperms.
- Xing Guo
- , Dongming Fang
- & Huan Liu
-
Article
| Open Access Co-evolution based machine-learning for predicting functional interactions between human genes
Co-evolution based machine-learning for predicting functional interactions between human genes
With the rise in number of eukaryotic species being fully sequenced, large scale phylogenetic profiling can give insights on gene function, Here, the authors describe a machine-learning approach that integrates co-evolution across eukaryotic clades to predict gene function and functional interactions among human genes.
- Doron Stupp
- , Elad Sharon
- & Yuval Tabach
-
Article
| Open Access Population structure, biogeography and transmissibility of Mycobacterium tuberculosis
Population structure, biogeography and transmissibility of Mycobacterium tuberculosis
Mycobacterium tuberculosis is a clonal pathogen that has co-evolved with humans for millennia. Here, Freschi et al. reevaluate the population structure of M. tuberculosis, providing an in-depth analysis of the ancient Indo-Oceanic Lineage 1 and the modern Central Asian Lineage 3, and expanding our understanding of Lineages 2 and 4.
- Luca Freschi
- , Roger Vargas Jr.
- & Maha Reda Farhat
-
Article
| Open Access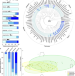 Multiple evolutionary origins and losses of tooth complexity in squamates
Multiple evolutionary origins and losses of tooth complexity in squamates
Tooth morphology has provided many insights into the tempo and mode of dietary evolution in mammals. A study of fossil and extant squamates shows that this group also repeatedly evolved increasingly complex teeth with more flexibility than mammals, and that higher tooth complexity and herbivory likely led to higher speciation rates.
- Fabien Lafuma
- , Ian J. Corfe
- & Nicolas Di-Poï

 Integrating full and partial genome sequences to decipher the global spread of canine rabies virus
Integrating full and partial genome sequences to decipher the global spread of canine rabies virus