Featured
-
-
Article
| Open Access Multiple communication mechanisms between sensor kinases are crucial for virulence in Pseudomonas aeruginosa
Multiple communication mechanisms between sensor kinases are crucial for virulence in Pseudomonas aeruginosa
Bacteria respond to stresses using two-component systems consisting of sensor kinases (SKs) and response regulators. Here, Francis et al. reveal three specific interaction mechanisms between a pair of SKs that are important for regulation of virulence in the pathogen Pseudomonas aeruginosa.
- Vanessa I. Francis
- , Elaine M. Waters
- & Steven L. Porter
-
Article
| Open Access Precise temporal regulation of alternative splicing during neural development
Precise temporal regulation of alternative splicing during neural development
The precise timing of neurodevelopmental splicing switches and the underlying regulatory mechanisms remain poorly understood. This study identifies two major waves of developmental switches under the control of distinct combinations of RNA-binding proteins in central and peripheral nervous systems.
- Sebastien M. Weyn-Vanhentenryck
- , Huijuan Feng
- & Chaolin Zhang
-
Article
| Open Access Multi-omics analysis reveals neoantigen-independent immune cell infiltration in copy-number driven cancers
Multi-omics analysis reveals neoantigen-independent immune cell infiltration in copy-number driven cancers
Neoantigen load has been associated with tumour immune infiltration. Here, the authors show that while this is true for tumours with recurrent mutations, cancers with recurrent CNAs show neoantigen-independent infiltration driven by cytokine production downstream of the DNA damage sensor ATM.
- Daniel J. McGrail
- , Lorenzo Federico
- & Nidhi Sahni
-
Article
| Open Access Reconstruction of complex single-cell trajectories using CellRouter
Reconstruction of complex single-cell trajectories using CellRouter
Single cell analysis provides insight into cell states and transitions, but to interpret the data, improved algorithms are needed. Here, the authors present CellRouter as a method to analyse single-cell trajectories from RNA-sequencing data, and provide insight into erythroid, myeloid and lymphoid differentiation.
- Edroaldo Lummertz da Rocha
- , R. Grant Rowe
- & George Q. Daley
-
Article
| Open Access Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Innate immunity combines intra- and intercellular signalling to develop responses that limit pathogen spread. Here the authors analyse feedback and feedforward loops connecting IRF3, NF-κB and STAT pathways, and suggest they allow coordinating cell fate decisions in cellular populations in response to the virus-mimicking agent poly(I:C).
- Maciej Czerkies
- , Zbigniew Korwek
- & Tomasz Lipniacki
-
Article
| Open Access Control of primary metabolism by a virulence regulatory network promotes robustness in a plant pathogen
Control of primary metabolism by a virulence regulatory network promotes robustness in a plant pathogen
How pathogens maintain phenotypic robustness during infection is poorly understood. Here the authors couple the virulence regulatory network (VRN) of the pathogen R. solanacearum to a model of its metabolic network, and find that the VRN activates functionally redundant primary metabolism genes to promote phenotypic robustness during infection.
- Rémi Peyraud
- , Ludovic Cottret
- & Stéphane Genin
-
Article
| Open Access Monitoring single-cell gene regulation under dynamically controllable conditions with integrated microfluidics and software
Monitoring single-cell gene regulation under dynamically controllable conditions with integrated microfluidics and software
How gene regulatory pathways control cell fate decisions in single cells is not fully understood. Here the authors present an integrated dual-input microfluidic chip and a linked analysis software, enabling tracking of gene regulatory responses of single bacterial cells to changing conditions.
- Matthias Kaiser
- , Florian Jug
- & Erik van Nimwegen
-
Article
| Open Access Genomic regression analysis of coordinated expression
Genomic regression analysis of coordinated expression
Somatic copy number alterations (SCNA) can confound gene co-expression analysis in cancers. Here the authors develop a method to remove the effects of SCNA in co-expression analysis, improving the analysis of network rewiring in cancer, and provide a database with adjusted data from TCGA.
- Ling Cai
- , Qiwei Li
- & Guanghua Xiao
-
Article
| Open Access Network dynamics-based cancer panel stratification for systemic prediction of anticancer drug response
Network dynamics-based cancer panel stratification for systemic prediction of anticancer drug response
Genomic alterations underlie the variability of drug responses between cancers, but our mechanistic understanding is limited. Here the authors use the p53 network to study how rewiring of signalling networks by genomic alterations impact their dynamic response to pharmacological perturbation.
- Minsoo Choi
- , Jue Shi
- & Kwang-Hyun Cho
-
Article
| Open Access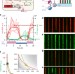 Balancing a genetic toggle switch by real-time feedback control and periodic forcing
Balancing a genetic toggle switch by real-time feedback control and periodic forcing
Cybergenetics aims to monitor and regulate cellular processes in real-time using computer monitoring and feedback of biological readouts. Here the authors use a feedback loop and periodic forcing to maintain cells with a bistable synthetic circuit near its unstable state.
- Jean-Baptiste Lugagne
- , Sebastián Sosa Carrillo
- & Pascal Hersen
-
Article
| Open Access Quorum sensing integrates environmental cues, cell density and cell history to control bacterial competence
Quorum sensing integrates environmental cues, cell density and cell history to control bacterial competence
Peptide CSP regulates natural competence in pneumococci and has been proposed as a quorum-sensing signal or a probe for sensing environmental cues. Here, the authors show that CSP levels can indeed act as an indicator of cell density and also incorporate information on environmental factors or cell history.
- Stefany Moreno-Gámez
- , Robin A. Sorg
- & Jan-Willem Veening
-
Article
| Open Access Evolution of new regulatory functions on biophysically realistic fitness landscapes
Evolution of new regulatory functions on biophysically realistic fitness landscapes
Gene networks evolve by transcription factor (TF) duplication and divergence of their binding site specificities, but little is known about the global constraints at play. Here, the authors study the coevolution of TFs and binding sites using a biophysical-evolutionary approach, and show that the emerging complex fitness landscapes strongly influence regulatory evolution with a role for crosstalk.
- Tamar Friedlander
- , Roshan Prizak
- & Gašper Tkačik
-
Article
| Open Access PLATE-Seq for genome-wide regulatory network analysis of high-throughput screens
PLATE-Seq for genome-wide regulatory network analysis of high-throughput screens
Despite the importance of pharmacological and functional genomic screens the readouts are of low complexity. Here the authors introduce PLATE-Seq, a low-cost genome-wide mRNA profiling method to complement high-throughput screening.
- Erin C. Bush
- , Forest Ray
- & Peter A. Sims
-
Article
| Open Access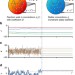 Exploratory adaptation in large random networks
Exploratory adaptation in large random networks
Recent works suggest that cellular networks may respond to novel challenges on the time-scale of cellular lifetimes through large-scale perturbation of gene expression and convergence to a new state. Here, the authors demonstrate the theoretical feasibility of exploratory adaptation in cellular networks by showing that convergence to new states depends on known features of these networks.
- Hallel I. Schreier
- , Yoav Soen
- & Naama Brenner
-
Article
| Open Access Cell fate decisions emerge as phages cooperate or compete inside their host
Cell fate decisions emerge as phages cooperate or compete inside their host
The bacteriophage lambda and its hostEscherichia coli provide a model system to study cell-fate decisions. Here, Trinh et al. develop a four-colour fluorescence system at the single-cell/single-virus/single-viral-DNA level and find phages cooperate during lysogenization and compete during lysis.
- Jimmy T. Trinh
- , Tamás Székely
- & Lanying Zeng
-
Article
| Open Access The impact of microRNAs on transcriptional heterogeneity and gene co-expression across single embryonic stem cells
The impact of microRNAs on transcriptional heterogeneity and gene co-expression across single embryonic stem cells
MicroRNAs can posttranscriptionally repress multiple targets in a cell population. Here the authors use single-cell sequencing to investigate the effects of an individual miRNA on transcriptional heterogeneity and gene co-expression
- Gennaro Gambardella
- , Annamaria Carissimo
- & Robert Blelloch
-
Article
| Open Access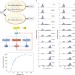 Noise reduction facilitated by dosage compensation in gene networks
Noise reduction facilitated by dosage compensation in gene networks
Cells must function despite the noisiness of their processes by tolerating or reducing such variability. Here, the authors combine experiment and modelling to show that a network motif that mediates network-dosage compensation also reduces noise in network output, suggesting that noise is tuneable.
- Weilin Peng
- , Ruijie Song
- & Murat Acar
-
Article
| Open Access Intrinsic limits to gene regulation by global crosstalk
Intrinsic limits to gene regulation by global crosstalk
Limited specificity of transcription factor-DNA interactions leads to crosstalk in gene regulation. Here the authors consider global crosstalk in regulatory networks of growing size and complexity, and show that it imposes constraints on gene regulation and on the evolution of regulatory networks.
- Tamar Friedlander
- , Roshan Prizak
- & Gašper Tkačik
-
Article
| Open Access The Parkinson’s disease-associated genes ATP13A2 and SYT11 regulate autophagy via a common pathway
The Parkinson’s disease-associated genes ATP13A2 and SYT11 regulate autophagy via a common pathway
Mutations in ATP13A2 are associated with lysosomal dysfunction and early onset Parkinson’s disease. Here Bento et al. show that ATP13A2 depletion negatively regulates SYT11, at both transcriptional and post-translational levels, which in turn impairs function of the autophagy-lysosome pathway.
- Carla F. Bento
- , Avraham Ashkenazi
- & David C. Rubinsztein
-
Article
| Open Access Extreme multifunctional proteins identified from a human protein interaction network
Extreme multifunctional proteins identified from a human protein interaction network
Proteins are sometimes implicated in separate and seemingly unrelated processes, so called moonlighting functions. Here the authors use bioinformatics tools to identify extreme multifunctional proteins and define a signature of extreme multifunctionality.
- Charles E. Chapple
- , Benoit Robisson
- & Christine Brun
-
Article |
 Orphan receptor IL-17RD regulates Toll-like receptor signalling via SEFIR/TIR interactions
Orphan receptor IL-17RD regulates Toll-like receptor signalling via SEFIR/TIR interactions
Toll-like receptors detect conserved microbial features to initiate host defence and are tightly regulated. Here the authors show that the orphan receptor interleukin-17 receptor D negatively regulates signalling downstream of Toll-like receptors to prevent excessive inflammation.
- Mark Mellett
- , Paola Atzei
- & Paul N. Moynagh
-
Article
| Open Access The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment
The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment
Autism genes converge in midfetal cortical co-expression networks, and chromatin regulators such as CHD8 are increasingly associated with autism spectrum disorder (ASD). Here the authors map CHD8 targets in developing brain, and find that CHD8 directly regulates other ASD risk genes during human neurodevelopment.
- Justin Cotney
- , Rebecca A. Muhle
- & James P. Noonan
-
Article |
 Deciphering Fur transcriptional regulatory network highlights its complex role beyond iron metabolism in Escherichia coli
Deciphering Fur transcriptional regulatory network highlights its complex role beyond iron metabolism in Escherichia coli
The ferric uptake regulator, Fur, is involved in the transcriptional regulation of iron metabolism. Here the authors show that Fur exhibits genome-wide regulatory effects in Escherichia colithat control many fundamental cellular processes linked to iron metabolism.
- Sang Woo Seo
- , Donghyuk Kim
- & Bernhard O. Palsson
-
Article |
 miRNAs confer phenotypic robustness to gene networks by suppressing biological noise
miRNAs confer phenotypic robustness to gene networks by suppressing biological noise
MicroRNAs are thought to confer robustness to biological processes, but clear experimental evidence is still needed. Here, Siciliano et al. construct a toggle-switch in mammalian cells to show that microRNAs buffer fluctuations in protein levels, thereby providing phenotypic robustness to gene regulatory networks.
- Velia Siciliano
- , Immacolata Garzilli
- & Diego di Bernardo
-
Article |
 Characterizing the interplay between multiple levels of organization within bacterial sigma factor regulatory networks
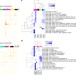
Characterizing the interplay between multiple levels of organization within bacterial sigma factor regulatory networks
Sigma factors are proteins controlling gene expression that allow bacteria to adapt to changing environmental conditions. Qiu and colleagues investigate sigma factor regulatory networks in Geobacter sulfurreducens, providing insights into how sigma factors regulate bacterial growth and energy metabolism.
- Yu Qiu
- , Harish Nagarajan
- & Karsten Zengler
 Oxidized phospholipids regulate amino acid metabolism through MTHFD2 to facilitate nucleotide release in endothelial cells
Oxidized phospholipids regulate amino acid metabolism through MTHFD2 to facilitate nucleotide release in endothelial cells