Article
|
Open Access
Featured
-
-
Article
| Open Access 2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
Here the authors captured the structure of human telomerase H/ACA ribonucleoprotein (RNP) by cryo-EM. The structure rationalizes telomere-disorder disease mutations and reveals insights into the mechanism of pseudouridylation by eukaryotic H/ACA RNPs.
- George E. Ghanim
- , Zala Sekne
- & Thi Hoang Duong Nguyen
-
Article
| Open Access The cellular and KSHV A-to-I RNA editome in primary effusion lymphoma and its role in the viral lifecycle
The cellular and KSHV A-to-I RNA editome in primary effusion lymphoma and its role in the viral lifecycle
The Karijolich laboratory describes an atlas of A-to-I RNA editing during the KSHV lifecycle in primary effusion lymphoma. These analyses identified conserved editing events within a viral-encoded microRNA, revealing a critical role for the microRNA and its modification in virus infection.
- Suba Rajendren
- , Xiang Ye
- & John Karijolich
-
Article
| Open Access Structural basis for sequence-independent substrate selection by eukaryotic wobble base tRNA deaminase ADAT2/3
Structural basis for sequence-independent substrate selection by eukaryotic wobble base tRNA deaminase ADAT2/3
The deamination of all adenosines in the wobble base position is essential to empower the individual tRNA to decode several codons. Here, the authors present the cryo-EM structure of the tRNA-bound ADAT2/3 deaminase, revealing how the geometry-specific enzyme acquiesces sequence-divergent tRNAs to its active site.
- Luciano G. Dolce
- , Aubree A. Zimmer
- & Eva Kowalinski
-
Article
| Open Access Sequential action of a tRNA base editor in conversion of cytidine to pseudouridine
Sequential action of a tRNA base editor in conversion of cytidine to pseudouridine
C-to-Ψ conversion is a previously uncharacterized form of base editing. Here, the authors describe how the TrcP enzyme catalyzes this process in a stepwise fashion and how this editing process is controlled by a network of modifications and nutrient availability to optimize translation efficiency.
- Satoshi Kimura
- , Veerasak Srisuknimit
- & Matthew K. Waldor
-
Article
| Open Access Adar-mediated A-to-I editing is required for embryonic patterning and innate immune response regulation in zebrafish
Adar-mediated A-to-I editing is required for embryonic patterning and innate immune response regulation in zebrafish
Additional roles for A-to-I editing of RNA continue to be uncovered. Niescierowicz et al. report prevalent A-to-I editing in the zebrafish transcriptome, and the distinct maternal and zygotic functions of the editing enzyme Adar in embryonic patterning and in the regulation of innate immune response, respectively.
- Katarzyna Niescierowicz
- , Leszek Pryszcz
- & Cecilia Winata
-
Article
| Open Access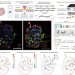 Spatial epitranscriptomics reveals A-to-I editome specific to cancer stem cell microniches
Spatial epitranscriptomics reveals A-to-I editome specific to cancer stem cell microniches
The spatial context of epitranscriptomic features in the tumour microenvironment remains poorly understood. Here, a method for transcriptomic and epitranscriptomic analysis of immunofluorescence-stained tissue, Select-seq, is applied to stem cell-like microniches in triple negative breast cancer.
- Amos C. Lee
- , Yongju Lee
- & Sunghoon Kwon
-
Article
| Open Access Crystal structures and insights into precursor tRNA 5’-end processing by prokaryotic minimal protein-only RNase P
Crystal structures and insights into precursor tRNA 5’-end processing by prokaryotic minimal protein-only RNase P
HARP are member of protein-only RNase P, which catalyzes pre-tRNA 5’-end processing and maturation. Here, the authors present crystal structure and provide mechanistic insights into pre-tRNA binding and cleavage by HARP proteins.
- Yangyang Li
- , Shichen Su
- & Jianhua Gan
-
Article
| Open Access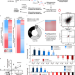 ADARs act as potent regulators of circular transcriptome in cancer
ADARs act as potent regulators of circular transcriptome in cancer
RNA editing and circRNAs are involved in tumorigenesis. Here the authors report that ADARs regulate the circular transcriptome in a bidirectional manner through and beyond their editing function in multiple cancer cells.
- Haoqing Shen
- , Omer An
- & Leilei Chen
-
Article
| Open Access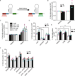 ADAR-mediated RNA editing of DNA:RNA hybrids is required for DNA double strand break repair
ADAR-mediated RNA editing of DNA:RNA hybrids is required for DNA double strand break repair
Different roles of specific RNA-related factors in DNA repair have now been reported. Here the authors reveal a role for RNA-editing by ADAR in DNA end resection following double strand break formation and a change in pattern of ADAR2-mediated A-to-I editing.
- Sonia Jimeno
- , Rosario Prados-Carvajal
- & Pablo Huertas
-
Article
| Open Access ADAR1 RNA editing enzyme regulates R-loop formation and genome stability at telomeres in cancer cells
ADAR1 RNA editing enzyme regulates R-loop formation and genome stability at telomeres in cancer cells
One type of RNA editing involves ADAR-mediated conversion of adenosine to inosine. Here the authors show that ADAR1 nuclear isoform p110 regulates R loop formation and genome stability at telomeres in cancer cells.
- Yusuke Shiromoto
- , Masayuki Sakurai
- & Kazuko Nishikura
-
Article
| Open Access Increased RNA editing in maternal immune activation model of neurodevelopmental disease
Increased RNA editing in maternal immune activation model of neurodevelopmental disease
Interferon can be induced by the viral mimic PolyI:C, which can in turn activate RNA editing enzymes. Here the authors identify transient changes in A-to-I RNA editing following maternal immune activation in mice.
- Hadas Tsivion-Visbord
- , Eli Kopel
- & Erez Y. Levanon
-
Article
| Open Access Adar RNA editing-dependent and -independent effects are required for brain and innate immune functions in Drosophila
Adar RNA editing-dependent and -independent effects are required for brain and innate immune functions in Drosophila
Human RNA editing enzymes ADAR1 and ADAR2 are required for innate immune functions and neurological functions, respectively. Here, the authors show that Drosophila Adar has both innate immune and brain functions, despite being the homolog of mammalian ADAR2.
- Patricia Deng
- , Anzer Khan
- & Liam P. Keegan
-
Article
| Open Access Cis- and trans-regulations of pre-mRNA splicing by RNA editing enzymes influence cancer development
Cis- and trans-regulations of pre-mRNA splicing by RNA editing enzymes influence cancer development
RNA editing and RNA splicing are involved in tumorigenesis. Here the authors report crosstalk between RNA editing and splicing by identifying ADAR1/2-dependent splicing events in esophageal squamous carcinoma cells.
- Sze Jing Tang
- , Haoqing Shen
- & Leilei Chen
-
Article
| Open Access ADAR1 mediated regulation of neural crest derived melanocytes and Schwann cell development
ADAR1 mediated regulation of neural crest derived melanocytes and Schwann cell development
ADAR1 is an RNA editing protein known to regulate immune responses to dsRNA that has been connected to neural crest cell function. Here, the authors show RNA editing by ADAR1 is important for the normal development of neural crest derived melanocytes and Schwann cells.
- Nadjet Gacem
- , Anthula Kavo
- & Nadege Bondurand
-
Article
| Open Access RNA editing is abundant and correlates with task performance in a social bumblebee
RNA editing is abundant and correlates with task performance in a social bumblebee
Bumblebee workers are genetically highly similar but they show different behaviors such as brood care and foraging. Here the authors report a high level of ADAR-mediated RNA editing in the bumblebee Bombus terrestris and its weak correlation to task performance.
- Hagit T. Porath
- , Esther Hazan
- & Guy Bloch
-
Article
| Open Access Pentatricopeptide repeat poly(A) binding protein KPAF4 stabilizes mitochondrial mRNAs in Trypanosoma brucei
Pentatricopeptide repeat poly(A) binding protein KPAF4 stabilizes mitochondrial mRNAs in Trypanosoma brucei
Polyadenylation stabilizes edited mitochondrial mRNAs in Trypanosoma brucei, but the involved poly(A) binding protein is unknown. Here, Mesitov et al. show that a pentatricopeptide repeat factor KPAF4 binds to A-tail and prevents exonucleolytic degradation as well as translation of incompletely edited mRNAs.
- Mikhail V. Mesitov
- , Tian Yu
- & Inna Aphasizheva
-
Article
| Open Access Regulation of gene expression and RNA editing in Drosophila adapting to divergent microclimates
Regulation of gene expression and RNA editing in Drosophila adapting to divergent microclimates
Environmental adaptation is generally studied at the genomic level, but it may also be driven by transcriptional processes. Here, the authors investigate variation in gene expression and RNA editing across diverging populations of Drosophila melanogaster from two microclimates.
- Arielle L. Yablonovitch
- , Jeremy Fu
- & Jin Billy Li
-
Article
| Open Access RNA editing generates cellular subsets with diverse sequence within populations
RNA editing generates cellular subsets with diverse sequence within populations
RNA editing rate detected from bulk RNA-seq data can vary widely. Here, by constructing a hierarchical Bayesian model, the authors report substantial variance in editing signatures detected by RNA-seq data from both single cells and a cognate bulk sample.
- Dewi Harjanto
- , Theodore Papamarkou
- & Anastasia Papavasiliou
-
Article
| Open Access ADAR-mediated RNA editing suppresses sleep by acting as a brake on glutamatergic synaptic plasticity
ADAR-mediated RNA editing suppresses sleep by acting as a brake on glutamatergic synaptic plasticity
Sleep is postulated to offset buildup in net synaptic strength that occurs during waking experience. Here, the authors identify a role for the RNA editing gene Adar in regulating glutamatergic synaptic plasticity and show that disruption in Adarexpression impairs normal waking in flies.
- J. E. Robinson
- , J. Paluch
- & W. J. Joiner
-
Article
| Open Access Activation-induced deoxycytidine deaminase (AID) co-transcriptional scanning at single-molecule resolution
Activation-induced deoxycytidine deaminase (AID) co-transcriptional scanning at single-molecule resolution
Activation-induced deoxycytidine deaminase (AID) induces somatic hypermutation and class-switch recombination during transcription of immunoglobulin genes. Here the authors use single-molecule FRET to show that AID translocates together with RNA polymerase and scans within stalled transcription bubbles.
- Gayan Senavirathne
- , Jeffrey G. Bertram
- & David Rueda
-
Article
| Open Access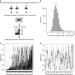 Genetic mapping uncovers cis-regulatory landscape of RNA editing
Genetic mapping uncovers cis-regulatory landscape of RNA editing
Adenosine-to-inosine (A-to-I) RNA editing plays an important role in neurological functions. Here, by a quantitative trait loci (QTL) mapping approach in 131 Drosophila melanogasterstrains, the authors identify 545 QTLs associated with differences in RNA editing.
- Gokul Ramaswami
- , Patricia Deng
- & Jin Billy Li
-
Article |
 Efficient AID targeting of switch regions is not sufficient for optimal class switch recombination
Efficient AID targeting of switch regions is not sufficient for optimal class switch recombination
Activation-induced cytidine deaminase can induce both somatic hypermutation (SHM) and class switch recombination in immunoglobulin loci. Here the authors show that spliced and repetitive switch regions are exquisite SHM targets, but are not sufficient by themselves for class switch recombination.
- Amélie Bonaud
- , Fabien Lechouane
- & Christophe Sirac
-
Article
| Open Access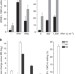 APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages
APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages
Aberrant RNA editing is linked to a range of neuropsychiatric and chronic diseases. Here Sharma et al. show that APOBEC3A can function as an RNA editing protein in response to physiological stimuli, significantly expanding our understanding of RNA editing and the role this may play in diseases.
- Shraddha Sharma
- , Santosh K. Patnaik
- & Bora E. Baysal
-
Article |
 Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways
Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways
ADAR1 is an adenosine deaminase that converts adenosine to inosine (A-to-I) mostly on Alu repeats in human RNA. Here by analysing transcriptome-wide ADAR1–RNA interactions, the authors show that ADAR1 also binds non-Alusequences to regulate alternative 3′ UTR usage and miRNA biogenesis in the nucleus.
- Jae Hoon Bahn
- , Jaegyoon Ahn
- & Xinshu Xiao
-
Article
| Open Access Caste-specific RNA editomes in the leaf-cutting ant Acromyrmex echinatior
Caste-specific RNA editomes in the leaf-cutting ant Acromyrmex echinatior
Post-translational mRNA editing has the potential to enhance the diversity of gene products and alter the functional properties of proteins. Here, Li et al. provide evidence that RNA editing is involved in generating caste-specific contrasting phenotypes in the leaf-cutting ant Acromyrmex echinatior.
- Qiye Li
- , Zongji Wang
- & Guojie Zhang
-
Article
| Open Access A genome-wide map of hyper-edited RNA reveals numerous new sites
A genome-wide map of hyper-edited RNA reveals numerous new sites
Common methods to detect adenosine-to-inosine RNA editing sites rely on mapping short RNA reads to the genome while allowing only a limited number of mismatches. Here, Porath et al. present a novel RNA-seq based approach to identify hyper-edited reads that significantly expands the RNA editome.
- Hagit T. Porath
- , Shai Carmi
- & Erez Y. Levanon
-
Article |
 RNA editing regulates transposon-mediated heterochromatic gene silencing
RNA editing regulates transposon-mediated heterochromatic gene silencing
The Hoppel transposable element mediates heterochromatin formation in Drosophila. Here Savva et al. report that the RNA-editing enzyme, ADAR, edits a long double-stranded RNA generated by the Hoppeltransposon, thereby regulating heterochromatin formation and gene expression.
- Yiannis A. Savva
- , James E. C. Jepson
- & Robert A. Reenan
-
Article |
 Tertiary structural elements determine the extent and specificity of messenger RNA editing
Tertiary structural elements determine the extent and specificity of messenger RNA editing
A central, imperfect duplex RNA secondary structure is generally required for site-specific adenosine-to-inosine RNA editing by ADAR enzymes. Rieder et al. show in Drosophila that conserved and complex long-range RNA tertiary structures form in vivoand can also regulate specific RNA-editing events by ADAR enzymes.
- Leila E. Rieder
- , Cynthia J. Staber
- & Robert A. Reenan
-
Article |
 Auto-regulatory RNA editing fine-tunes mRNA re-coding and complex behaviour in Drosophila
Auto-regulatory RNA editing fine-tunes mRNA re-coding and complex behaviour in Drosophila
Adars are adenosine deaminases that act on RNAs, including those encoding proteins involved in neuronal transmission and also Adar RNA. Here, Savvaet al. engineered knock-in Drosophila mutants with altered Adar autoediting and found that this changed the spectrum of adenosine deamination and Drosophilabehaviour.
- Yiannis A. Savva
- , James E.C Jepson
- & Robert A. Reenan
-
Article
| Open Access Editing of human KV1.1 channel mRNAs disrupts binding of the N-terminus tip at the intracellular cavity
Editing of human KV1.1 channel mRNAs disrupts binding of the N-terminus tip at the intracellular cavity
RNA editing is important in regulating neuronal excitability, and a specific editing event has been shown to alter the permeation pathway of voltage-gate potassium channels. Gonzalezet al.find that the tip of the channel's inactivation gate makes a direct hydrophobic interaction with the edited position.
- Carlos Gonzalez
- , Angelica Lopez-Rodriguez
- & Miguel Holmgren
-
Article
| Open Access Predicting sites of ADAR editing in double-stranded RNA
Predicting sites of ADAR editing in double-stranded RNA
ADAR enzymes edit double-stranded RNA, converting adenosines to inosines, and are essential for neuronal function. Eggingtonet al. quantify edit sites in RNA using a Sanger sequencing protocol and use the resulting data to develop algorithms to predict RNA edit sites.
- Julie M. Eggington
- , Tom Greene
- & Brenda L. Bass

 Unveiling the A-to-I mRNA editing machinery and its regulation and evolution in fungi
Unveiling the A-to-I mRNA editing machinery and its regulation and evolution in fungi