Featured
-
-
Article
| Open Access Blood monocyte transcriptome and epigenome analyses reveal loci associated with human atherosclerosis
Blood monocyte transcriptome and epigenome analyses reveal loci associated with human atherosclerosis
The molecular mechanisms mediating the impact of environmental factors in atherosclerosis are unclear. Here, the authors examine CD14+ blood monocyte’s transcriptome and epigenome signatures to find differential methylation and expression of ARID5B to be associated with human atherosclerosis.
- Yongmei Liu
- , Lindsay M. Reynolds
- & James H. Stein
-
Article
| Open Access HDAC3 is a molecular brake of the metabolic switch supporting white adipose tissue browning
HDAC3 is a molecular brake of the metabolic switch supporting white adipose tissue browning
Histone deacetylases, such as HDAC3, have been shown to alter cellular metabolism in various tissues. Here the authors show that HDAC3 regulates WAT metabolism by activating a futile cycle of fatty acid synthesis and oxidation, which supports WAT browning.
- Alessandra Ferrari
- , Raffaella Longo
- & Maurizio Crestani
-
Article
| Open Access A transcriptomics data-driven gene space accurately predicts liver cytopathology and drug-induced liver injury
A transcriptomics data-driven gene space accurately predicts liver cytopathology and drug-induced liver injury
Predicting the hepatotoxic effects of new drugs is still a challenge. Using toxicogenomics data, the authors here define a predictive toxicogenomic space (PTGS), the component gene space capturing dose-dependent cytotoxicity, and demonstrate that it can be used to accurately predict drug-induced liver pathology, including human drug-induced liver injury fromin vitrodata.
- Pekka Kohonen
- , Juuso A. Parkkinen
- & Roland C. Grafström
-
Article
| Open Access Transcriptomic and macroevolutionary evidence for phenotypic uncoupling between frog life history phases
Transcriptomic and macroevolutionary evidence for phenotypic uncoupling between frog life history phases
In animals with complex life cycles, selection on one life phase may constrain adaptation in another phase. Here the authors find that, during the adaptive radiation of mantellid frogs, the evolution of tadpole and adult morphologies has been uncoupled through phase-specific gene expression.
- Katharina C. Wollenberg Valero
- , Joan Garcia-Porta
- & Miguel Vences
-
Article
| Open Access Image-guided genomics of phenotypically heterogeneous populations reveals vascular signalling during symbiotic collective cancer invasion
Image-guided genomics of phenotypically heterogeneous populations reveals vascular signalling during symbiotic collective cancer invasion
The mechanisms linking phenotypic heterogeneity to collective cancer invasion are unclear. Here the authors develop an image-guided genomic technique to select and amplify leader and follower cells fromin vitroinvading cell packs and find a cooperative symbiotic relationship between these two cell populations.
- J. Konen
- , E. Summerbell
- & A. I. Marcus
-
Article
| Open Access Quantification of differential gene expression by multiplexed targeted resequencing of cDNA
Quantification of differential gene expression by multiplexed targeted resequencing of cDNA
Transcriptome sequencing is a powerful tool for functional analysis of different types of RNA molecules in a wide range of applications. Here the authors use targeted resequencing of cDNA with single-molecule molecular inversion probes as a cost-effective, high-throughput tool for mRNA quantification.
- Peer Arts
- , Jori van der Raadt
- & Cornelis A. Albers
-
Article
| Open Access Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Global patterns of gene transcription can be represented with reduced dimensionality. Here, the authors devise a method called Tradict that learns and uses 100 marker genes to predict transcriptome-wide pathway expression levels and patterns that reflect cell activity and state.
- Surojit Biswas
- , Konstantin Kerner
- & Philip A. Wigge
-
Article
| Open Access Neurons and neuronal activity control gene expression in astrocytes to regulate their development and metabolism
Neurons and neuronal activity control gene expression in astrocytes to regulate their development and metabolism
How neurons and neuronal activity regulate astrocyte functions is poorly understood. Haselet al. identify two large groups of astrocytic genes that are regulated by neuronal contact and synaptic activity respectively, with distinct roles in astrocytic function; interestingly, many of these genes are dysregulated in neurodegeneration.
- Philip Hasel
- , Owen Dando
- & Giles E. Hardingham
-
Article
| Open Access FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons
FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons
The RNA binding protein FUS functions in several RNA biosynthetic processes and has been linked to the pathogenesis of amyotrophic lateral sclerosis (ALS). Here the authors show that FUS controls back-splicing reactions leading to circular RNA (circRNA) production in stem cell-derived motor neurons and that ALS-associated FUS mutations affect the biogenesis of circRNAs.
- Lorenzo Errichelli
- , Stefano Dini Modigliani
- & Irene Bozzoni
-
Article
| Open Access Epigenetically-driven anatomical diversity of synovial fibroblasts guides joint-specific fibroblast functions
Epigenetically-driven anatomical diversity of synovial fibroblasts guides joint-specific fibroblast functions
Arthritis affects different joints variably despite systemic inflammatory cues. Here the authors show anatomical differences in the transcriptome, epigenome and function of synovial fibroblasts that might affect susceptibility to site-specific joint diseases.
- Mojca Frank-Bertoncelj
- , Michelle Trenkmann
- & Caroline Ospelt
-
Article
| Open Access Genomic innovations linked to infection strategies across emerging pathogenic chytrid fungi
Genomic innovations linked to infection strategies across emerging pathogenic chytrid fungi
Batrachochytrium dendrobatidis and B. salamandrivoransare both important pathogens of amphibians, but they differ in their host ranges, infection strategies, and host immune responses. Here, Farrer and colleagues compare their genomes and transcriptomes to identify the genetic basis of these differences.
- Rhys A. Farrer
- , An Martel
- & Christina A. Cuomo
-
Article
| Open Access CLK-dependent exon recognition and conjoined gene formation revealed with a novel small molecule inhibitor
CLK-dependent exon recognition and conjoined gene formation revealed with a novel small molecule inhibitor
The phosphorylation of serine/arginine-rich proteins by CDC-like kinase is a central regulatory mechanism for RNA splicing reactions. Here, the authors synthesize a novel small molecule CLK inhibitor and map CLK-responsive alternative splicing events and discover an effect on conjoined gene transcription.
- Tyler Funnell
- , Shinya Tasaki
- & Samuel Aparicio
-
Article
| Open Access Massively parallel digital transcriptional profiling of single cells
Massively parallel digital transcriptional profiling of single cells
Single-cell gene expression analysis is challenging. This work describes a new droplet-based single cell RNA-seq platform capable of processing tens of thousands of cells across 8 independent samples in minutes, and demonstrates cellular subtypes and host–donor chimerism in transplant patients.
- Grace X. Y. Zheng
- , Jessica M. Terry
- & Jason H. Bielas
-
Article
| Open Access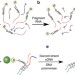 Global repositioning of transcription start sites in a plant-fermenting bacterium
Global repositioning of transcription start sites in a plant-fermenting bacterium
Bacteria may respond to a change in environment by using alternative transcriptional start sites. Here, the authors use a novel genome-wide capture and reverse transcription method to find substrate-specific start sites for hundreds of genes at single base resolution inClostridium phytofermentans.
- Magali Boutard
- , Laurence Ettwiller
- & Andrew C. Tolonen
-
Article
| Open Access Tetrapod limb and sarcopterygian fin regeneration share a core genetic programme
Tetrapod limb and sarcopterygian fin regeneration share a core genetic programme
Salamanders are unique among extant tetrapods for their ability to completely regenerate their limbs. Here, Nogueira and colleagues show that lungfishes, the sister clade of tetrapods, regenerate their fins using analogous gene regulatory changes and morphological steps.
- Acacio F. Nogueira
- , Carinne M. Costa
- & Igor Schneider
-
Article
| Open Access Molecular characterization of Thy1 expressing fear-inhibiting neurons within the basolateral amygdala
Molecular characterization of Thy1 expressing fear-inhibiting neurons within the basolateral amygdala
Activation of Thy-1 expressing neurons in the basolateral amygdala (BLA) has been shown to result in inhibition of fear. Here the authors present a comprehensive workflow to identify genes that are upregulated in this population and can be pharmacologically targeted to enable fear extinction.
- Kenneth M. McCullough
- , Dennis Choi
- & Kerry J. Ressler
-
Article
| Open Access Breast cancer genome and transcriptome integration implicates specific mutational signatures with immune cell infiltration
Breast cancer genome and transcriptome integration implicates specific mutational signatures with immune cell infiltration
Recent studies using in depth DNA sequencing techniques led to the identification of cancer driver genes but mainly focused on the effect on their expression. Here, the authors analyse 266 cases of breast cancer and report gene expression signatures associated with the number and character of signature mutations.
- Marcel Smid
- , F. Germán Rodríguez-González
- & John W. M. Martens
-
Article
| Open Access Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Long non-coding RNAs are increasingly recognised to be important factors in regulating cellular processes and comprise a large faction of the transcriptome, however most are uncharacterised. Here the authors present RACE-Seq, a tool to improve and extend the annotation of low-expression transcripts.
- Julien Lagarde
- , Barbara Uszczynska-Ratajczak
- & Jennifer Harrow
-
Article
| Open Access Laser capture microscopy coupled with Smart-seq2 for precise spatial transcriptomic profiling
Laser capture microscopy coupled with Smart-seq2 for precise spatial transcriptomic profiling
Laser capture microscopy (LCM) coupled with global transcriptome profiling requires relatively large numbers of cells. Here, the authors show that LCM coupled with full-length mRNA-sequencing (LCM-seq) can sequence single cells, and that LCM-seq can provide biological insight on highly similar neuronal populations.
- Susanne Nichterwitz
- , Geng Chen
- & Eva Hedlund
-
Article
| Open Access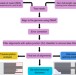 A survey of the sorghum transcriptome using single-molecule long reads
A survey of the sorghum transcriptome using single-molecule long reads
Alternative splicing and alternative polyadenylation (APA) contribute to mRNA diversity but are difficult to assess using short read RNA-seq data. Here, the authors use single molecule long-read isoform sequencing and develop a computational pipeline to identify full-length splice isoforms and APA sites in sorghum.
- Salah E. Abdel-Ghany
- , Michael Hamilton
- & Anireddy S. N. Reddy
-
Article
| Open Access Unveiling the complexity of the maize transcriptome by single-molecule long-read sequencing
Unveiling the complexity of the maize transcriptome by single-molecule long-read sequencing
Zea mays is an important crop species and genetic model but uncertainties remain regarding the structure of the transcriptome. Here Wang et al. use single-molecule sequencing and size-fractionated libraries to identify novel transcripts and isoforms illustrating the complexity of maize mRNA.
- Bo Wang
- , Elizabeth Tseng
- & Doreen Ware
-
Article
| Open Access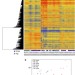 Joint-specific DNA methylation and transcriptome signatures in rheumatoid arthritis identify distinct pathogenic processes
Joint-specific DNA methylation and transcriptome signatures in rheumatoid arthritis identify distinct pathogenic processes
Rheumatoid arthritis is an inflammatory disease that selectively affects different joints. Here the authors show that gene expression and DNA methylation patterns of fibroblast-like synoviocytes differ between hip and knee joints in patients with RA, thus providing epigenetic and transcriptomic evidence for this anatomic selectivity of inflammation.
- Rizi Ai
- , Deepa Hammaker
- & Wei Wang
-
Article
| Open Access Survival trade-offs in plant roots during colonization by closely related beneficial and pathogenic fungi
Survival trade-offs in plant roots during colonization by closely related beneficial and pathogenic fungi
Colletotrichum tofieldiae is a beneficial root endophyte, whereas the closely related C. incanumis pathogenic. Here the authors compare the genomes and transcriptomes during host plant interaction and demonstrate that the host plant can respond differently to the beneficial endophyte according to phosphate status.
- Stéphane Hacquard
- , Barbara Kracher
- & Richard J. O’Connell
-
Article
| Open Access Highly variable cancer subpopulations that exhibit enhanced transcriptome variability and metastatic fitness
Highly variable cancer subpopulations that exhibit enhanced transcriptome variability and metastatic fitness
Phenotypic and genetic intra-tumor heterogeneity have an important role in cancer progression and therapeutic resistance. Here, the authors show that phenotypically variable tumor subpopulations exhibit higher metastatic potential and display enhanced intra-clonal transcriptomic variability, likely promoted by deregulated spliceosome activity.
- Alexander Nguyen
- , Mitsukuni Yoshida
- & Sohail F. Tavazoie
-
Article
| Open Access Genome-scale study reveals reduced metabolic adaptability in patients with non-alcoholic fatty liver disease
Genome-scale study reveals reduced metabolic adaptability in patients with non-alcoholic fatty liver disease
Non-alcoholic fatty liver disease (NAFLD) is a risk factor for other types of liver diseases. Here, the authors integrate transcriptomic and metabolomic data from patients with NAFLD with a genome-scale metabolic model to paint a comprehensive picture of liver function in NAFLD.
- Tuulia Hyötyläinen
- , Livnat Jerby
- & Matej Orešič
-
Article
| Open Access Abasic pivot substitution harnesses target specificity of RNA interference
Abasic pivot substitution harnesses target specificity of RNA interference
RNA interference inadvertently represses off-target transcripts. Here, Lee et al.report that substituting nucleotide in position 6 of the seed region of the small interfering RNAs with abasic spacers can significantly decrease miRNA-like off-target repression while preserving on-target activity.
- Hye-Sook Lee
- , Heeyoung Seok
- & Sung Wook Chi
-
Article
| Open Access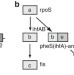 The propagation of perturbations in rewired bacterial gene networks
The propagation of perturbations in rewired bacterial gene networks
Expression of transcription factors to alter gene regulation can cause substantial changes to expression across a genome. Here the authors ‘rewire’ E. coliand analyse the global transcriptome alterations to identify novel network interactions.
- Rebecca Baumstark
- , Sonja Hänzelmann
- & Mark Isalan
-
Article
| Open Access Endothelial Gata5 transcription factor regulates blood pressure
Endothelial Gata5 transcription factor regulates blood pressure
Unravelling the molecular basis of hypertension remains a major challenge. Here, the authors identify the transcription factor GATA5 as a novel regulator of blood pressure and potential genetic determinant of human hypertension and describe a unique mouse model for research of salt-sensitive hypertension.
- Smail Messaoudi
- , Ying He
- & Mona Nemer
-
Article
| Open Access Genomic and transcriptomic evidence for scavenging of diverse organic compounds by widespread deep-sea archaea
Genomic and transcriptomic evidence for scavenging of diverse organic compounds by widespread deep-sea archaea
The contribution of marine archaea to the ocean's carbon cycle is unclear. Here, Li et al. analyse the genomes and transcriptomes from five deep-sea archaeal groups to reveal their metabolic characteristics, suggesting a crucial role in modulating the carbon cycle in deep oceans.
- Meng Li
- , Brett J. Baker
- & Gregory J. Dick
-
Article
| Open Access Transcriptomes of parents identify parenting strategies and sexual conflict in a subsocial beetle
Transcriptomes of parents identify parenting strategies and sexual conflict in a subsocial beetle
The burying beetle shows flexible parenting behaviour. Here, the authors show that offspring fare equally well regardless of the sex or number of parents present and find similar gene expression profiles in uniparental and biparental females and in uniparental males, which suggests no specialization in parenting.
- Darren J. Parker
- , Christopher B. Cunningham
- & Allen J. Moore
-
Article
| Open Access RC3H1 post-transcriptionally regulates A20 mRNA and modulates the activity of the IKK/NF-κB pathway
RC3H1 post-transcriptionally regulates A20 mRNA and modulates the activity of the IKK/NF-κB pathway
The RNA-binding protein RC3H1/ROQUIN1 promotes the degradation of mRNA by binding to a consensus CDE present in the 3′UTR. Here the authors expand the set of consensus sequences through which RCH31 binds and regulates mRNA encoding members of the DNA damage response and IKK/NF-κB pathway.
- Yasuhiro Murakawa
- , Michael Hinz
- & Markus Landthaler
-
Article
| Open Access TCF12 is mutated in anaplastic oligodendroglioma
TCF12 is mutated in anaplastic oligodendroglioma
Anaplastic oligodendrogliomas are rare and incurable primary brain tumours with few treatment options. Here Labrecheet al. perform whole-exome sequencing and identify recurring mutations in transcription factor TCF12, which are associated with aggressive tumours.
- Karim Labreche
- , Iva Simeonova
- & Michel Wager
-
Article
| Open Access The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome
The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome
The evolutionarily conserved MTREC complex promotes degradation of meiotic mRNAs and regulatory ncRNAs. Here the authors show that MTREC also targets cryptic unstable transcripts and unspliced pre-mRNAs for degradation by the nuclear exosome, while the TRAMP complex has only a minor role in this process.
- Yang Zhou
- , Jianguo Zhu
- & Tamás Fischer
-
Article
| Open Access Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state
Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state
The key regulators that allow transition from proliferative to invasive phenotype in melanoma cells have not been identified yet. The authors perform chromatin and transcriptome profiling followed by comprehensive bioinformatics analysis identifying new candidate regulators for two distinct cell states of melanoma.
- Annelien Verfaillie
- , Hana Imrichova
- & Stein Aerts
-
Article |
 ID4 controls mammary stem cells and marks breast cancers with a stem cell-like phenotype
ID4 controls mammary stem cells and marks breast cancers with a stem cell-like phenotype
Basal-like breast cancer is a heterogeneous disease with poor prognosis; however, its cellular origins and aetiology are poorly understood. Here the authors provide evidence that ID4 is a key controller of mammary stem/progenitor cell self-renewal, acting upstream of Notch signalling to repress luminal fate commitment.
- Simon Junankar
- , Laura A. Baker
- & Alexander Swarbrick
-
Article |
 Transformation of the intestinal epithelium by the MSI2 RNA-binding protein
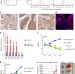
Transformation of the intestinal epithelium by the MSI2 RNA-binding protein
In mammals there are two Musashi proteins, MSI1 and MSI2, orthologues of the Drosophila protein, with roles in asymmetric stem cell division and cell fate determination. Here the authors report new functions for MSI2 in colorectal cancer using in vitro loss of function and in vivoectopic overexpression.
- Shan Wang
- , Ning Li
- & Christopher J. Lengner
-
Article
| Open Access Macrotene chromosomes provide insights to a new mechanism of high-order gene amplification in eukaryotes
Macrotene chromosomes provide insights to a new mechanism of high-order gene amplification in eukaryotes
Copy number variation is an important source of genetic variation in natural populations and may have a role in human disease. Here, the authors identify high-order amplification structures that form large extended chromosomes and suggest that these may occur due to accidental template switching in stress conditions.
- Agnès Thierry
- , Varun Khanna
- & Bernard Dujon
-
Article |
 Sequencing of first-strand cDNA library reveals full-length transcriptomes
Sequencing of first-strand cDNA library reveals full-length transcriptomes
Strand-specific RNA-seq (ssRNA-seq) data often lack information on 5′ and 3′ ends of transcripts. Here the authors present a novel method for ssRNA-seq that enables the simultaneous profiling of gene expression, TSSs and polyadenylation sites at near-base resolution with a single library.
- Saurabh Agarwal
- , Todd S. Macfarlan
- & Shigeki Iwase
-
Article |
 The statistical geometry of transcriptome divergence in cell-type evolution and cancer
The statistical geometry of transcriptome divergence in cell-type evolution and cancer
Body plan complexity is associated with the number of different cell types, yet the processes that create this diversity are unclear. Here the authors use transcriptomics to test the hypothesis that unlike cancer cells, novel normal cell types arise through sub-specialization of an ancestral cell type.
- Cong Liang
- , Alistair R.R. Forrest
- & Günter P. Wagner
-
Article
| Open Access Enhanced transcriptome maps from multiple mouse tissues reveal evolutionary constraint in gene expression
Enhanced transcriptome maps from multiple mouse tissues reveal evolutionary constraint in gene expression
The analysis of mammalian transcriptomes could provide new insights into human biology. Here the authors carry out RNA sequencing in a large collection of mouse tissues and compare these data to human transcriptome profiles, identifying a set of constrained genes that carry out basic cellular functions with remarkably constant expression levels across tissues and species.
- Dmitri D. Pervouchine
- , Sarah Djebali
- & Thomas R. Gingeras
-
Article |
 Transcriptome meta-analysis of lung cancer reveals recurrent aberrations in NRG1 and Hippo pathway genes
Transcriptome meta-analysis of lung cancer reveals recurrent aberrations in NRG1 and Hippo pathway genes
Targeted cancer therapy requires knowledge of driver aberrations. Here the authors perform large-scale transcriptome analysis, and show that gene fusions in NRG1, NF1and Hippo pathway genes are recurrent mostly among lung cancers lacking known driver mutations.
- Saravana M. Dhanasekaran
- , O Alejandro Balbin
- & Arul M. Chinnaiyan
-
Article
| Open Access Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism
Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism
Autism spectrum disorder (ASD) is a common, highly heritable neurodevelopmental condition characterized by marked genetic heterogeneity. In this study, the authors use RNA sequencing analyses to characterize differences in the transcriptome between autistic and typically developing brains.
- Simone Gupta
- , Shannon E. Ellis
- & Dan E. Arking
-
Article |
 microTSS: accurate microRNA transcription start site identification reveals a significant number of divergent pri-miRNAs
microTSS: accurate microRNA transcription start site identification reveals a significant number of divergent pri-miRNAs
microRNAs are short non-coding RNAs that post-transcriptionally regulate gene expression for which the identification of promoter and primary transcripts (pri-miRNAs) has been difficult. Here the authors describe microTSS, an algorithm that supports the precise identification of intergenic pri-miRNA transcription start sites.
- Georgios Georgakilas
- , Ioannis S. Vlachos
- & Artemis G. Hatzigeorgiou
-
Article |
 Somatic transcriptome priming gates lineage-specific differentiation potential of human-induced pluripotent stem cell states
Somatic transcriptome priming gates lineage-specific differentiation potential of human-induced pluripotent stem cell states
Molecular and functional differences between induced pluripotent stem cells (iPSCs) derived from distinct cell types have been described. Here the authors show, by comparing human iPSCs derived from fibroblasts or cord blood, that the competence in activating developmental genes upon differentiation is influenced by the donor cell of origin.
- Jong-Hee Lee
- , Jung Bok Lee
- & Mickie Bhatia
-
Article |
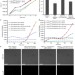 Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles
Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles
Bacteria can grow as free living planktonic cells or as part of surface-associated biofilms. Here the authors show, for the opportunistic pathogen Pseudomonas aeruginosa, that cells recently dispersed from biofilms are physiologically different from, and more virulent than, planktonic and biofilm cells.
- Song Lin Chua
- , Yang Liu
- & Liang Yang
-
Article |
 Genome-wide adaptive complexes to underground stresses in blind mole rats Spalax
Genome-wide adaptive complexes to underground stresses in blind mole rats Spalax
The blind mole rat (BMR), Spalax galili, is perfectly adapted to life underground. Here, the authors sequence the BMR genome and transcriptome and highlight genomic features that may have played a role in adaptation to extreme underground stressors, such as darkness hypercapnia and hypoxia.
- Xiaodong Fang
- , Eviatar Nevo
- & Jun Wang
-
Article
| Open Access A rat RNA-Seq transcriptomic BodyMap across 11 organs and 4 developmental stages
A rat RNA-Seq transcriptomic BodyMap across 11 organs and 4 developmental stages
Gene expression is highly variable between tissues, and changes during development and with age. Here, the authors provide a comprehensive RNA-Seq analysis of the rat transcriptome, spanning eleven organs, four developmental stages and both sexes.
- Ying Yu
- , James C. Fuscoe
- & Charles Wang
-
Article
| Open Access Inferring tumour purity and stromal and immune cell admixture from expression data
Inferring tumour purity and stromal and immune cell admixture from expression data
Tumour biopsies contain contaminating normal cells and these can influence the analysis of tumour samples. In this study, Yoshihara et al.develop an algorithm based on gene expression profiles from The Cancer Genome Atlas to estimate the number of contaminating normal cells in tumour samples.
- Kosuke Yoshihara
- , Maria Shahmoradgoli
- & Roel G.W. Verhaak
-
Article
| Open Access The landscape of viral expression and host gene fusion and adaptation in human cancer
The landscape of viral expression and host gene fusion and adaptation in human cancer
Viruses contribute to the pathogenesis of certain cancers. Using massively parallel sequencing data from The Cancer Genome Atlas to analyse viral expression in 19 tumour types, Tang et al. both confirm and reject previously described viral associations and present new information on viral integration and host interaction.
- Ka-Wei Tang
- , Babak Alaei-Mahabadi
- & Erik Larsson

 Genetic correlations reveal the shared genetic architecture of transcription in human peripheral blood
Genetic correlations reveal the shared genetic architecture of transcription in human peripheral blood