Abstract
We introduce Crystal Edge Graph Attention Neural Network (CEGANN) workflow that uses graph attention-based architecture to learn unique feature representations and perform classification of materials across multiple scales (from atomic to mesoscale) and diverse classes ranging from metals, oxides, non-metals to hierarchical materials such as zeolites and semi-ordered mesophases. CEGANN can classify based on a global, structure-level representation such as space group and dimensionality (e.g., bulk, 2D, clusters, etc.). Using representative materials such as polycrystals and zeolites, we demonstrate its transferability in performing local atom-level classification tasks, such as grain boundary identification and other heterointerfaces. CEGANN classifies in (thermal) noisy dynamical environments as demonstrated for representative zeolite nucleation and growth from an amorphous mixture. Finally, we use CEGANN to classify multicomponent systems with thermal noise and compositional diversity. Overall, our approach is material agnostic and allows for multiscale feature classification ranging from atomic-scale crystals to heterointerfaces to microscale grain boundaries.
Similar content being viewed by others
Introduction
Characterization of materials with unique properties1,2,3,4,5 is at the core of data-driven material design and discovery6,7. A relatively small fraction of materials has been characterized either experimentally or with computational methods, compared to their anticipated potential diversity across a vast chemical space. Given the surge in the development of materials databases8,9,10 in recent years, there is an urgent need for automated tools to analyze large amounts of structural data. In this regard, distinguishing the unique characteristics across different classes of materials with varying dimensionality can provide key insights into learnable aspects which are crucial for state-of-the-art machine learning (ML) tools to be successfully implemented in the design and discovery of new materials with unique properties. To achieve such distinction, ML models typically involve the use of fingerprints or descriptors11,12,13,14,15,16 that allow a learning algorithm to map the fingerprint to a user-desired property of interest. A descriptor that maps the crystal features in a vector space should always be (1) invariant to basis symmetries such as rotation, reflection, translation, and permutation of atoms12, (2) unique to the system applied, but sensitive towards variation in properties, and (3) simple and robust. Additionally, these features play a crucial role in a wide range of applications such as quantitative structure-property relationship (QSPR)11,17,18,19,20, development of interatomic potentials13,21,22,23, prediction of atomistic configurations based on targeted properties24,25,26,27,28, surface phenomena29, etc.
A feature representation is constructed primarily in two ways (i) using a predefined mathematical formulation, or (ii) learning the representation by combining fundamental low-level features and correlating them to the relevant task being performed using ML methods. A plethora of mathematical formulation-based descriptors13,14,15,30,31,32,33,34 such as radial distribution functions (RDF), angular distribution function (ADF), common neighbor analysis (CNA)30, adaptive CNA30, centro-symmetry parameter (CSP)30, Voronoi analysis30, Steinhardt order parameter (SP)31, bond angle analysis (BAA)32, and neighbor distance analysis (NDA)30 are widely used for featurization. A majority of these are very simple and of a very local nature, i.e., mostly capable of differentiating ordered and disordered structures. Improving upon these, a set of features can be developed using pairwise feature matrices and their transformations12,18,19,20,33,34. These features may be as simple as pairwise distances e.g., Weyl matrices34, Z-matrices35, or pairwise electrostatic interactions between atoms (Coulomb matrix18 and sine matrix19). A more comprehensive representation of these matrices is permutation histograms15 e.g., MBTR (Many-Body Tensor Representation)12, BOB (Bag of Bonds)20. The advantages of these methods are that the pairwise features are translationally invariant, and these matrices present a unique representation of the system. However, a major setback of these matrix representations is that they are not invariant to changes in atom ordering. A very popular approach involves the use of smooth overlap of atomic positions (SOAP)14 descriptors constructed by expanding the atomic neighbor density ρ on a spherical harmonics basis and then further expanding it on a radial basis to obtain the rotational invariant power spectrum33. SOAP provides a robust representation of the local environment in a smooth and continuous manner which makes it very suitable for mapping potential energy surfaces. Nevertheless, most of the existing approaches for structure characterization that involve global comparison between two structures are either based on a simple aggregation-based method (average kernel) that causes loss in resolution or are computationally expensive such as, for example, the best match kernel method14. Another setback is that the number of descriptors increases quadratically6 with the increase in the number of chemical species, precluding their applicability to multicomponent systems.
In this context, graph neural networks (GNNs)36,37,38 have been widely used in node-level as well as graph-level classification tasks with remarkable success. Furthermore, recent developments in the area of graph attention networks39 make the task more accurate by learning the interaction between node-level features. These networks tend to learn flexible representations by combining very fundamental low-level features (interatomic distances, bond angles, etc.), and yet produce a graph-based input that very accurately maps to the target-specific tasks. This caters to the limitations of static descriptors bound by predefined mathematical formulations.
In crystal systems, GNNs can be made to operate on atom-based graph representation to create node-level embedding through convolutions on neighboring nodes and edges17,40,41,42,43,44,45,46. More layers of convolutions tend to capture higher-level information. A widely used framework for crystal systems is the Crystal Graph Convolutional Neural Network (CGCNN)17,46. Xie and Grossman have shown that CGCNN can directly learn material properties from the connectivity of atoms in a crystal, thus enabling an interpretable representation of crystalline materials47. Graph attention-based architecture43 has also recently been implemented for the structure-to-property mapping in atomistic systems. Traditional CGCNN architecture tends to map structure to the property by using a diverse set of atom-level features (e.g., group number, period number, atomic number, electronic structure, etc.), and crystal graphs with simple edge feature such as pairwise interatomic distances.
Predefined mathematical formulation-based descriptors are useful when there isn’t sufficient data to learn from. However, they largely suffer from transferability issues due to a lack of flexibility. On the other hand, current existing graph-based CGCNN architectures do not incorporate orientational features40,44 that are very relevant for classification tasks in a multitude of atomic environments. Moreover, these features tend to play a more significant role in classification tasks than features belonging to different atomic species. Although there have been recent applications40,44 that include orientational features in their network architecture, they are more complex in nature and mostly focus on property prediction. To elucidate the issues involving transferability and applicability, we present two distinct classification scenarios (Fig. 1). To begin with, we classify the liquid and glassy-amorphous phases of a representative material such as silicon (Si). Both liquid and amorphous Si phases are disordered with no symmetry whatsoever and only differ in density and coordination number. Figure 1a displays the variability in the coordination environment of the liquids and amorphous Si phases used in this study. We use a dataset containing 2000 Si structures with 50% of them being liquid and rest 50% amorphous (see Supplementary Notes 1 for the details on the data generation methods) and train a traditional CGCNN model using a train-to-test split of 80:20. From the t-SNE (t-distributed stochastic neighbor embedding) of feature representation of the test dataset, with SOAP (cutoff 6 Å) (Fig. 1b), there is no distinct separation of the phases in features space indicating the inability of SOAP to distinguish the individual phases (see Supplementary Fig. 1c, d) for different cutoffs). On the other hand, simple bond order based features (Q2, Q4, Q6) + CN (Coordination Number) (cutoff 6 Å) (Fig. 1c) and a trained CGCNN (Fig. 1d) can clearly characterize the two phases with decent separation in feature space. The second task involves the identification of particles belonging to local motifs (hexagonal or cubic) in a stacking disordered (ABABCB), ice. The correlated bond order-based CHILL+48 is used as a benchmark for labeling the data (Fig. 1e). Similar to the earlier case, we employ order parameters, SOAP, and CGCNN for this classification. The training data of CGCNN comprises a pure cubic, hexagonal, and stacking faulted (ABCBCB) ice structure. The results in Fig. 1f–h indicate that while SOAP is able to classify local motifs, CGCNN or the order parameters-based features fail to do so. This is converse to the fact that SOAP could not characterize structures belonging to a liquid or an amorphous class while its two counterparts could. This is an indication of the transferability issue in existing characterization techniques across various problems at different scales. Although traditional GNNs (such as CGCNN) showing exceptional promise in learning flexible feature representation at a graph level (global), their performance in local environments is not as good as their predictability of global attributes (e.g., properties such as energy, bandgap, etc.) and remains mostly unexplored.
a Typical coordination number distribution of liquid and amorphous Si structures. b t-SNE plot of SOAP representation of the crystalline and amorphous Si phases in the test dataset. c t-SNE plot for bond-order parameters (Q2, Q4, Q6) + CN (Coordination Number), feature representation. d t-SNE plot of embeddings of the test dataset obtained by training a classic CGCNN model. e classification of hexagonal and cubic stacked ice in an ABABCB stacked ice with CHILL + 48 algorithm. f classification using SOAP feature vector g by using order parameters (Q2, Q4, Q6) + CN, and h using the trained CGCNN model.
Clearly, there is a need for a method that is not only transferable, but adaptable to variabilities in the material environment while providing accurate characterization at different scales. To the best of our knowledge, most efforts on crystal graph neural networks have been restricted to map structures to properties and a few property-based prediction tasks. There is still an immense untapped potential for GNNs in classification at both the structure (global) and atomic (local) levels. In this work, we introduce a graph attention-based51 workflow that operates on edge graphs, convoluting edges, and bond angle features and passing messages in between (Fig. 2), to learn feature representation of material environments. An advantage of attention-based architectures is that they can learn the importance of feature vectors (i.e., bonds and angles) in the neighborhood of each atom and put emphasis on the ones unique to the task being performed. This helps in increasing performance by ignoring redundant and unnecessary information. We demonstrate the efficacy of our workflow in classification tasks at both the atom-level (local) and structure-level (global) using a wide range of representative examples from materials applications. For global-level classification, we perform two tasks. The first is classifying a diverse range of materials based on their space groups, and the second is classifying them based on their dimensionality (bulk, 2D, cluster, etc.). We base the local atom-level classification on structural motifs (FCC, BCC, HCP, and diamond cubic), and demonstrate its use on a classic problem of grain boundary identification and grain size distribution. To validate the efficacy of our workflow in environments with thermal variations or noise, we deploy our classification workflow to facilitate the study of nucleation and growth of a zeolite, a complex porous crystal, in molecular dynamics simulations of synthesis. Often, practical materials application involves the characterization of phases with structural and compositional variances along with thermal noise. We address these challenges through the identification of ice and liquid along simulations of water crystallization, and the classification of disordered, mesophase, and crystalline orders in simulations of binary mixtures involving transformations between these phases.
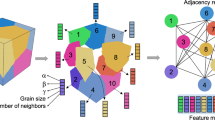
a, b Crystals are converted into atom graphs and edge graphs are obtained from atom graphs. c Shows alternate message passing and hierarchical interaction between edge and angle convolutions. Finally, the atom feature, convolved edge, and angle features are concatenated to produce the final representation. d t-SNE plot of the feature vector of liquid and amorphous structures as predicted by CEGANN workflow on an identical test dataset as Fig. 1. e Shows identification of particles belonging to a hexagonal and cubic motif in an ABABCB stacked ice by a trained CEGANN workflow.
Results
Edge graph representation
Edge graphs are higher-order representations of atomic graphs with edges as nodes and bond angles as connections between a pair of edges (Fig. 2b). We start from a crystal structure, creating its atom-graph (atom as nodes, bonds as edges) based on a fixed number of nearest neighbors. The edge graph is extracted from the atom-graph afterward (Fig. 2a, b). The edge features (eij) are obtained by expanding the pairwise distance on Gaussian basis functions while the bond angle features (θijk) are obtained by expanding the cosines of the bond angles on a Gaussian basis as well.
Hierarchical message passing
One main feature of the proposed architecture is the hierarchical interaction between edge and angle layers (Fig. 2c) (see Methods section). The edge layer always gets updated first. This follows the hierarchy that the bond angles are constructed from a pair of edges and any change at the edge level should get updated first before passing the information onto the corresponding angle. This gives n-1 angle convolution operations for n edge convolutions, where n is an integer.
CEGANN workflow for multiscale classification
The architecture of the CEGANN workflow used to perform multiscale classification of materials is shown in Fig. 2c. The edge-graph feature representation of the structures is passed to the hierarchical message passing block for convolution operations. The output of the convolved feature vectors from the edge and angle convolution layers are then passed to the aggregation block via dense layers (linear transformation), where feature representations of each of the structures are generated for the prediction task. For multicomponent systems, additional chemical information can be included in the input edge feature vector \(f_{e_{ij}}\) (Fig. 2c) as one-hot encoding, depending on the characterization task being performed. CEGANN architecture also has an inherent ability of learning to distinguish atomic species from the interatomic distances of nearest neighbor atoms (see Supplementary Note 2 and Supplementary Fig. 2). The choice of the number of edges and angle convolution layers to be employed depends on the scale at which the classification tasks are being performed. For local-level tasks, it is preferable to have fewer convolutions while its global application requires more. In this work, we select an optimal number of convolutions that results in the best performance of our model for each of the tasks being performed (Table 1). Similar to the choice of the number of convolutions, the number of neighbors considered for the graph constructions also affects the model performance. A grid study can be performed to obtain an optimal set of hyperparameters for the specific task (see Supplementary Note 3 and Supplementary Fig. 3a–c). In the end, the selection becomes entirely dependent on the choice of the problem, the computational cost associated, and the accuracy of the prediction. The number of neighbors and the number of convolutions used for each of the tasks is reported in Table 1. It is to be noted that for all classification tasks performed in this work, we keep the input dimension of edge and angle feature vectors to be 80. We maintain uniformity of samples belonging to each class in both training and testing data while the splitting of any individual class is done randomly at a given ratio.
Classification of liquid and amorphous silicon and stacking-disordered ice
We start by employing our CEGANN workflow for the classification tasks as discussed in Fig. 1. (a) Classification of liquid and amorphous phases (Silicon) Fig. 1a–d. (b) Characterization of local motifs (Hexagonal or Cubic) in stacking-disordered ice (ABABCB) Fig. 1e–h. CEGANN is trained on the same training data as CGCNN (Fig. 1d, h). The test data is also kept identical. From Fig. 2d, the t-SNE plot of the feature vectors of liquid and amorphous Si structures as predicted by CEGANN, it is evident that CEGANN has been able to distinguish amorphous and liquid phases of silicon conspicuously. Figure 2e also depicts the ability of CEGANN to precisely classify local cubic and hexagonal motifs in stacking fault structures where CGCNN performs poorly. CEGANN is shown to overcome the challenge of transferability for both applications ranging from global to local levels while its counterparts, such as the traditional CGCNN, and descriptors such as SOAP fail to do so (Fig. 1).
Characterization of crystal structures based on their space groups
The space group of a crystalline system directly correlates to its structural motif, albeit at a global level. We demonstrate that the CEGANN framework can classify several different material classes based on their space groups. For this classification task, we use the same dataset as in ref. 49. The space group of each crystal is calculated using a Pymatgen50 package. The dataset contains a total of 10,517 crystal structures with seven crystal classes belonging to eight different space groups. For the elemental system, the classes are body-centered tetragonal (bct, 139 and 141), rhombohedral (rh, 166), hexagonal (hex, 194), simple cubic (sc, 221), face-centered cubic (fcc, 225), diamond (dia, 227), and body-centered cubic (bcc, 229), respectively (see Supplementary Fig. 4a).
We start with the dataset having a train-to-test ratio of 90:10 and train CEGANN, CGCNN, and a SOAP_ML workflow on this dataset. It is worth noting that our goal is to map SOAP feature vectors (cutoff 6 Å) directly to the space group. So, instead of passing the SOAP features through consecutive dense layers (linear transformation) with nonlinear activations51, we have only one dense layer that directly maps it to the target space (SOAP_ML workflow) conforming to the specification used in CEGANN after the aggregation block (Fig. 2c). The accuracy on the test dataset is shown in Fig. 3d. The CEGANN workflow achieves an accuracy of ~100% on the test set. The confusion matrix of CEGANN (Fig. 3a) also demonstrates a perfect identification (no off-diagonal entries) (also see Supplementary Fig. 4b–i) of each class belonging to different space groups. The CGCNN, on the other hand, achieves an accuracy of ~83% on the test dataset with major confusion (Fig. 3b) between the hex (194) and fcc (225) structures. This is evident from the fact that fcc and hcp are close-packed with a 74% atomic packing factor, and 12 nearest neighbors for both, which results in an identical graphical representation of the structures unless the orientational order of the particles are considered. The CGCNN not having these attributes in its graphical representation, significantly impacts its performance. The performance of SOAP_ML workflow is poor indicating that SOAP in its current mathematical state, however, does contain all the information but is not flexible enough to be directly mappable to the target space group. The degree of characterization can also be visualized in the t-SNE plot of the feature space representation on the test dataset (Fig. 3e–g). There is a clear distinction in the representation of each class for CEGANN, while CGCNN and SOAP feature vectors display a lack of resolution in the representation of each class in the feature space.
Classification of polymorphs across various structural dimensionalities
Next, we demonstrate the ability of CEGANN to perform classification on material polymorphs across various dimensionalities, from clusters (0D) to sheets (2D) to bulk (3D). Carbon is known to have a diverse range of allotropes across these dimensionalities, making it an excellent candidate for validating the performance of our network for dimensionality classification. We start with a dataset of 511 bulk structures collected from the Samara Carbon Allotrope Database (SACADA)52. Monolayer C polymorphs53, Graphite with varying interlayer distances, and a collection of different Graphite allotrope and 2D polymorphs Carbon sampled using CASTING framework1,54 and LCBOP potential55 making a total of 612, 2D structures. The addition of 704 C nanoclusters56,57 result in a total dataset of 1827 configurations (see Supplementary Fig. 5). We divide our dataset into 80% training and 20% test.
Figure 4a shows the confusion matrix for the dimensionality classification. CEGANN workflow can classify the structures with ~100% accuracy. Figure 4b shows the t-SNE plot of the embeddings of the test set data. A clear distinction between phases can also be observed in the feature space which displays the capability of CEGANN to characterize polymorphs of different dimensions. It is worth mentioning that dimensionality is a defining material parameter, depending on which material can exhibit dramatically different properties58. Identification of materials based on their dimensionality is a crucial aspect of new material design and prediction25. While 3D crystalline objects are well documented among the experimentally known crystals, the same is not true for low dimensional structures such as 2D or 0D. For example, in a few cases, isolated 2D carbon layers tend to form porous bulk-like polymorphs which makes it difficult to categorize and distinguish them from typical layered structures.
Grain boundary identification
Characterization of local motifs in full 3D samples of polycrystalline materials and accurately identifying grains and boundaries is a nontrivial task with a plethora of applications in material science. Although there are many methods used for grain characterization30,32, there is no gold standard for identifying the grain size distribution in polycrystalline materials, as the predictions widely vary with the methodology used. We use CNA (Common Neighbor Analysis)30 as a benchmark to generate labels for the training and test data. CNA has been widely utilized for the characterization of local motifs in ordered and disordered systems30,59,60,61. The original CNA method is based on generating the signatures of the local neighborhood of an atom and matching it to a reference one. The neighborhood of an atom is constructed based of a fixed cutoff (rcut). The overall atomic signature of the atom consists of three features: (1) the number of neighbor atoms the central atom and its bonded neighbor have in common, ncn, (2) the total number of bonds between these common neighbors, nb, and (3) the number of bonds in the longest chain of bonds connecting the common neighbors, nlcb. However, traditional CNA, and even its variations (such as adaptive CNA), not only show variability in results, but their performance also deteriorates under conditions with physical deformtion61.
Here, we consider 4 representative polycrystal classes for the prediction task. These are (i) Face-Centered Cubic (FCC- Al), Body-Centered cubic (BCC-W), Diamond (Si), and Hexagonal Closed Pack (HCP-Mg) with 40 grains. For the prediction of each of the aforementioned classes, we generate 10 polycrystalline training samples (see Supplementary Fig. 6a–d) using the atomsk62 package. The overall characterization is carried out with a two-step approach. First, we label the atoms locally based on their crystalline motifs (e.g, FCC, BCC, etc.) and then, we apply an unsupervised learning DBSCAN63,64 clustering to identify the size of the grains in the polycrystal samples. The grain size distribution and the number of particles belonging to crystalline motifs as predicted by CEGANN and CNA have been compared in Fig. 5. It is to be noted that the ordinary CNA cannot classify the diamond structure. Hence, we use a modified CNA65 for the creation of the labels of the Si (diamond structures). The number of nearest neighbors used for the construction of the graphs for each of the classifications is reported in Table 1. This conforms to the number of neighbors that traditional CNA30 uses for the prediction tasks.
The predictions of CEGANN (Fig. 5a–d) are almost identical to those of CNA, both in terms of the grain size distribution and the number of particles belonging to crystalline motifs of the grains. This clearly demonstrates the ability of the CEGANN in learning the different local motifs and distinguishing them from disordered atoms. The predictions of CEGANN on the local-level classification tasks are largely dependent on the selection of the number of convolutional layers in the model as well as the number of neighbors used for the local neighborhood of the edge graphs. Adding more convolution layers will cause the compression of too much information at a single node. This may result in a loss of resolution, which in turn would deteriorate the CEGANN performance. As we increase the number of convolutional layers for fixed 12 neighbors of graph construction (Fig. 6a), the performance severely declines at four edge-convolutional (+3 angle convolutions) layers. However, it seems that with an increase in the number of neighbors, CEGANN tends to slightly underpredict grain sizes (Fig. 6b). The amount of information being compressed in each node of a graph using subsequent convolutions follows the equation:
where, Ninformation is the information from surrounding neighbors in terms the number of atoms, “NN” is the number of nearest neighbors of an atom in the graph, and “CONV” is the number of convolutions being used. The mean grain size of the Mg (HCP) system is ~1200 with a maximum value of ~2500. In Fig. 6c, beyond the operation point 12, 3 (NN, CONV) the amount of information being compressed is ~8000, which is much larger than the maximum grain size. Hence, there is a severe mix-up between the information on grain boundary and grains. Thus, the model tends to perform poorly at 4 edge-convolutional (+3 angle convolutions) (Fig. 6a). An increase in NN will cause this deterioration very slowly and will result in an underprediction of grain sizes (Fig. 6b). It is also worth mentioning that, unlike CNA, CEGANN is very flexible in learning environments with local noises, such as thermal noise, which is essential for practical applications.
a Effect number of edge-convolutional layer on the prediction of grain size distribution. b Effect of number nearest neighbors used for graph construction on grain boundary prediction. c The amount of information (in terms of the number of atoms) being compressed in a node of the graph for different edge convolutions (& n-1 angle convolutions) and the number of neighbors used for the graph construction.
Dynamical classification of structures with thermal noise
Zeolites are ordered microporous silicates or aluminosilicate66,67 materials widely used as solid catalysts in the chemical industry. Knowledge about the mechanistic pathways of the formation of zeolites is still limited, which is a key to realizing new zeolites for catalysis and separations. The stochastic nature of nucleation processes and the small, nanoscopic size of critical nuclei within the heterogeneous reaction mixture, make the detection of the birth of a new phase challenging in experimental hydrothermal synthesis. Molecular simulations have the right spatial resolution. However, in the synthesis mixture, the zeolite crystallites and the surrounding amorphous matrix have very similar local and medium-range orders68. Figure 7a, b shows that, indeed, the zeolite and the network former silica in the amorphous phase has very similar radial and Qn (number of silica neighbors) distributions. Moreover, unlike the case of simple crystals, such as ice, where the unit cell consists of 1–2 atoms, the unit cell of zeolites typically has ~100 silica nodes. Even though each silicon has a coordination number of 4, the environment of each silicon node in a zeolite is diverse. This makes the identification of the nascent zeolite inside an amorphous matrix a very challenging endeavor.
a, b The radial distribution function between silica nodes (gT-T) and the number of silica neighbors (Qn) is very similar between the amorphous and zeolite phases. c CEGANN predicts the fraction of silica sites that are part of the zeolite, as it nucleates and grows from the synthesis mixture. The snapshots of the simulation box corresponding to points A–D are shown in the lower panel. Silica nodes of the amorphous phase are shown in orange, whereas the crystalline silica detected by CEGANN is shown in green. For clarity, the organic cations and water molecules are not shown.
Traditional approaches, such as the bond-orientational order parameter q6, could be used to detect the nucleation process of zeolites. However, the requirement of the large cutoff distance makes it inefficient to detect very small nuclei69,70,71. Moreover, the bond-order parameter approach is specific to a particular zeolite polymorph. Identification of crystal based on mobility criteria is not zeolite specific, but it assumes that there is a considerable mobility difference between the new crystal phase from the mother phase. This approach does not work if the new phase crystallizes from a glassy state, as is the case in zeolite synthesis68. These necessitate the development of a classification technique that distinguishes the zeolite nucleus from the amorphous phase during the formation of zeolites.
We use the CEGANN framework to probe the evolution of the zeolite nucleus and growth in the simulation mentioned above. To train our network, we use a total of 400 structures consisting of 50% pure crystalline zeolites at different temperatures, noisy zeolite crystals (added Gaussian noise to the atomic positions) as well as 50% amorphous structures at different temperatures (see Supplementary Note 4). We use 12 NN (nearest neighbors) (see Table 1) for the graph construction, although the effects of four and eight nearest neighbors on the construction of the graph are also explored (see Supplementary Fig. 7c). Figure 7c shows the zeolite fraction in the simulation trajectory as a function of time for the case of 12NN. A sharp change in the fraction of zeolite starting at time 16.5 ns suggests the formation of stable nuclei of zeolite Z1 that grow into a full slab at a time >25 ns. The same is evident from the snapshots presented at different instances during the crystallization (panel A–D in Fig. 7). This case study clearly illustrates that the proposed CEGANN workflow is not only capable of performing accurate classification in static local environments but also equally effective in heterogeneous simulation environments with considerable thermal noise.
Multilabel characterization of mesophases in binary mixtures
Mesophases have an ordered intermediate between that of amorphous and crystalline phases. They are traditionally observed in block copolymers and solutions surfactants but can also occur in other systems with frustrated attraction72,73. Mesophases occur on multiple morphologies such as lamellar, gyroid, and hexagonal72,74. The intermediate nature of the ordering in mesophases makes them challenging to identify in simulations. We use the CEGANN workflow to characterize the formation of mesophases, and subsequent crystallization during the cooling of a binary isotropic mixture of representative species A and B72. We also characterize the order of the species in the system as the phase transition is taking place. The dataset consisted of 22 lamellar, 22 crystalline, and 22 isotropic mixture structures (see Supplementary Note 5 and Supplementary Fig. 8). The transitions are validated with the potential energy changes in the system (side panel of Fig. 8a). Figure 8a demonstrates that CEGANN successfully characterizes the amorphous, lamellar and crystalline phases individually, and also accurately detects the transition between phases along a cooling simulation.
a Characterization of isotropic liquid, lamellar mesophase, and layered crystal in a binary synthesis mixture. CEGANN identifies each phase along the thermal trajectories of phase transformation. Respective potential energy changes with time and corresponding predictions of existing phases by CEGANN (b–d) are the confusion matrix of the overall chemical species predicted by CEGANN at different time steps (A, B, C) for mesophase characterization. e Predicting the growth of ice from liquid at 235 K, along with the atoms (“H” & “O”) present in the system (Multilabel). f–h are the confusion matrix of the atomic species predicted at different time steps (A, B, C) as the system is crystalizing.
Multilabel classification of interface evolution during ice growth
The crystallization of water is ubiquitous in natural environments. Development in the last two decades, have enabled simulations of ice nucleation and growth with molecular resolution75,76,77,78. Here, we implement CEGANN for the characterization of the early stages of growth of ice I from liquid water, a polyatomic molecule. The molecular dynamics (MD) simulation was carried out using the TIP4P/200579 water model (see Supplementary Notes 5 and Supplementary Fig. 8). Using our multilabel classification approach, we classify whether a particle in the MD trajectory belongs to either crystalline or liquid phase, and also identify the local order of each water molecules. In Fig. 8e, we show that CEGANN precisely characterizes the crystallization of water (reflected in the decrease of the potential energy of the system).
The above two examples—phase transitions in the binary mixture and crystallization of water—display CEGANN’s ability to characterize complex environments with multiple components or polyatomic species, in the presence of thermal noise.
Discussion
Characterization of materials at different scales and domains of application is a must for any data-driven material science application. In this work, we develop the graph attention-based CEGANN workflow, which is transferrable across scales and adaptable to variabilities in the material environment, while also providing an accurate characterization. We demonstrate the efficacy of our workflow on challenging and relevant classification problems in material science. Unlike similar graph-based architecture (CGCNN) or mathematical formulation-based descriptors (SOAP, order parameters), CEGANN is not only able to classify disordered (liquid and amorphous phases) at a global level but is equally accurate in classifying local motifs in stacking-disordered structures, displaying transferability in the application domain. It is equally effective in performing global-level classification tasks such as space group classification and characterization of structures based on their dimensionality, at the same time, it can characterize local motifs, grain boundaries, and grain size distribution in polycrystalline materials accurately.
We further extend the applicability of CEGANN in systems with significant practical implications. Systems that have compositional variability accompanied by thermal fluctuations. CEGANN can identify the formation of complex crystals with large unit cells, identifying of onset of nucleation and growth of a zeolite from a synthesis solution with strong thermal fluctuations, even when the size of the nucleus is much smaller than the unit cell of the zeolite, and captures the growth process accurately. It can also identify crystalline and amorphous phases in polyatomic systems with thermal noise, as well as distinguish liquid, mesophase, and crystalline order in binary mixtures. These applications showcase the applicability of CEGANN to problems involving variability in the environments. Overall, our approach is agnostic to the problem and allows the classification of features at different scales with equal efficacy.
Methods
Angle convolution
The angle convolutional layer uses bond angle (θijk) cosines expanded on a gaussian basis as the initial input. The idea is that each angle learns and collects the messages from its adjacent edges through the convolutions. We use a simple graph attention-based architecture and convolutional operation is performed according to
where \(e_{ij}^l,\;e_{jk}^l\) are edge features from previous edge convolution layers and αijkl is the attention coefficient calculated using39
where \(W_{ijkl}^f,\;W_{ijkl}^{att}\) and \(b_{ijkl}^f,\;b_{ijkl}^{att}\) are feature and attention weights and biases, respectively. We use softmax activation as a normalizer for calculating the attention coefficient and the final output of the angle convolution is passed through a softplus activation to obtain the final representation. Batch normalization is applied after the aggregation operation.
Edge convolution
We follow a similar attention-type mechanism for the edge-convolutional layer. The convolutional function is represented as
where \(W_{ijk}^f\) and \(b_{ijk}^f\) are the weights and biases for the feature matrix and \(\theta _{ijk}^l\) is the angle features from the previous angle convolutional stage. αijk, the attention coefficient computed using an equation analogous to Eq. 2, with different weights and biases. We apply a nonlinear softplus activation function before and after the aggregation over the neighborhood; the additional nonlinearity helps the features to adapt to the target task. There is also a provision for adding explicit one hot-coded atomic feature xi based on the characterizing task being performed. The incorporation of the chemical information is done before each edge convolution. For l + 1th edge convolution layer, with \(e_{ij}^l\) as input form the lth layer, the atomic features of atom i and j (xi, xj) are included as a concatenation of the features (Fig. 2c).
Feature aggregation and concatenation
The aggregation block (Fig. 2c) consists of three stages. First, the edge and angle features are aggregated as
The final feature representation is given as concatenation \(Z_i = e_i^{l + 1} \oplus \theta _i^{l + 1}\). To provide more resolution to the aggregated feature, we take a linear transformation before the aggregation stage. The pooling of the features follows the concatenation operation. It should be noted that the pooling (average-pooling) on the features is applied only if a global-level classification task is being performed. For local classification tasks, no pooling is applied to the features. Batch normalization is applied after the aggregation operation. We also apply dropouts’ (0.5 rates) before subsequent transformation after the convolutional layer. This helps in reducing overfitting. We use cross-entropy loss as the loss metric17.
Training the model
The network is trained on 1 GPU-accelerated to compute node on the NERSC computing cluster with 20-core Intel Xeon Gold 6148 (‘Skylake’) @ 2.40 GHz and 1 NVIDIA Tesla V100 (‘Volta’) GPU. The feature vector for the Angle convolution and edge convolutions are kept being 80. The hidden feature for the dense layer following the edge and angle convolution layers is 256. Upon aggregation, the overall dimension of the feature vector is 512.
Data availability
The dataset used for space group classification is available via https://www.nomad-coe.eu/49. The Carbon bulk structures used in this work are available via SACADA database https://www.sacada.info/. All the other datasets used for different classification task are available at https://github.com/sbanik2/CEGANN.
Code availability
The CEGANN code along with the trained models are available via https://github.com/sbanik2/CEGANN.
References
Zhang, H.-T. et al. Reconfigurable perovskite nickelate electronics for artificial intelligence. Science 375, 533–539 (2022).
Dwivedi, N. et al. Unusual high hardness and load-dependent mechanical characteristics of hydrogenated carbon–nitrogen hybrid films. ACS Appl. Mater. Interfaces 14, 20220–20229 (2022).
Mansouri Tehrani, A. et al. Machine learning directed search for ultraincompressible, superhard materials. J. Am. Chem. Soc. 140, 9844–9853 (2018).
Schaibley, J. R. et al. Valleytronics in 2D materials. Nat. Rev. Mater. 1, 1–15 (2016).
Gogotsi, Y. & Anasori, B. The rise of MXenes. ACS Nano 13, 8491–8494 (2019).
Schmidt, J., Marques, M. R. G., Botti, S. & Marques, M. A. L. Recent advances and applications of machine learning in solid-state materials science. npj Comput. Mater. 5, 1–36 (2019).
Liu, Y., Zhao, T., Ju, W. & Shi, S. Materials discovery and design using machine learning. J. Materiomics 3, 159–177 (2017).
Curtarolo, S. et al. AFLOWLIB. ORG: a distributed materials properties repository from high-throughput ab initio calculations. Comput. Mater. Sci. 58, 227–235 (2012).
Jain, A. et al. Commentary: the materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Saal, J. E., Kirklin, S., Aykol, M., Meredig, B. & Wolverton, C. Materials design and discovery with high-throughput density functional theory: the open quantum materials database (OQMD). JOM 65, 1501–1509 (2013).
Isayev, O. et al. Universal fragment descriptors for predicting properties of inorganic crystals. Nat. Commun. 8, 1–12 (2017).
Huo, H. & Rupp, M. Unified representation of molecules and crystals for machine learning. Mach. Learn. Sci. Technol. 3, 045017 (2022).
Behler, J. Atom-centered symmetry functions for constructing high-dimensional neural network potentials. J. Chem. Phys. 134, 074106 (2011).
De, S., Bartók, A. P., Csányi, G. & Ceriotti, M. Comparing molecules and solids across structural and alchemical space. Phys. Chem. Chem. Phys. 18, 13754–13769 (2016).
Musil, F. et al. Physics-inspired structural representations for molecules and materials. Chem. Rev. 121, 9759–9815 (2021).
Ward, L., Agrawal, A., Choudhary, A. & Wolverton, C. A general-purpose machine learning framework for predicting properties of inorganic materials. npj Comput. Mater. 2, 1–7 (2016).
Xie, T. & Grossman, J. C. Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys. Rev. Lett. 120, 145301 (2018).
Rupp, M., Tkatchenko, A., Müller, K.-R. & Von Lilienfeld, O. A. Fast and accurate modeling of molecular atomization energies with machine learning. Phys. Rev. Lett. 108, 058301 (2012).
Faber, F., Lindmaa, A., von Lilienfeld, O. A. & Armiento, R. Crystal structure representations for machine learning models of formation energies. Int. J. Quant. Chem. 115, 1094–1101 (2015).
Hansen, K. et al. Machine learning predictions of molecular properties: accurate many-body potentials and nonlocality in chemical space. J. Phys. Chem. Lett. 6, 2326–2331 (2015).
Bartók, A. P., Payne, M. C., Kondor, R. & Csányi, G. Gaussian approximation potentials: the accuracy of quantum mechanics, without the electrons. Phys. Rev. Lett. 104, 136403 (2010).
Seko, A., Takahashi, A. & Tanaka, I. Sparse representation for a potential energy surface. Phys. Rev. B 90, 024101 (2014).
Jiang, B. & Guo, H. Permutation invariant polynomial neural network approach to fitting potential energy surfaces. J. Chem. Phys. 139, 054112 (2013).
Revard, B. C., Tipton, W. W. & Hennig, R. G. Genetic algorithm for structure and phase prediction. GitHub repository. https://doi.org/10.5281/zenodo.2554076 (2018).
Oganov, A. R., Pickard, C. J., Zhu, Q. & Needs, R. J. Structure prediction drives materials discovery. Nat. Rev. Mater. 4, 331–348 (2019).
Banik, S. et al. Learning with delayed rewards—a case study on inverse defect design in 2D materials. ACS Appl. Mater. Interfaces 13, 36455–36464 (2021).
Loeffler, T. D., Banik, S., Patra, T. K., Sternberg, M. & Sankaranarayanan, S. K. R. S. Reinforcement learning in discrete action space applied to inverse defect design. J. Phys. Commun. 5, 031001 (2021).
Ryan, K., Lengyel, J. & Shatruk, M. Crystal structure prediction via deep learning. J. Am. Chem. Soc. 140, 10158–10168 (2018).
Jäger, M. O. J., Morooka, E. V., Federici Canova, F., Himanen, L. & Foster, A. S. Machine learning hydrogen adsorption on nanoclusters through structural descriptors. npj Comput. Mater. 4, 1–8 (2018).
Stukowski, A. Structure identification methods for atomistic simulations of crystalline materials. Model. Simul. Mater. Sci. Eng. 20, 045021 (2012).
Steinhardt, P. J., Nelson, D. R. & Ronchetti, M. Bond-orientational order in liquids and glasses. Phys. Rev. B 28, 784 (1983).
Ackland, G. J. & Jones, A. P. Applications of local crystal structure measures in experiment and simulation. Phys. Rev. B 73, 054104 (2006).
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B 87, 184115 (2013).
Weyl, H. The Classical Groups: Their Invariants and Representations (Princeton Univ. Press, 1946).
Jensen, F. Introduction to Computational Chemistry (Wiley, 2017).
Scarselli, F., Gori, M., Tsoi, A. C., Hagenbuchner, M. & Monfardini, G. The graph neural network model. IEEE Trans. Neural Netw. 20, 61–80 (2008).
Xu, K., Hu, W., Leskovec, J. & Jegelka, S. How powerful are graph neural networks? Preprint at https://doi.org/10.48550/arXiv.1810.00826 (2018). Also, published as proceedings of The International Conference on Learning Representations (2019).
Zhou, J. et al. Graph neural networks: a review of methods and applications. AI Open 1, 57–81 (2020).
Velickovic, P. et al. Graph attention networks. Stat 1050, 20 (2017).
Klicpera, J., Groß, J. & Günnemann S. Directional message passing for molecular graphs. Preprint at https://doi.org/10.48550/arXiv.2003.03123 (2020). Also, published as proceedings of The International Conference on Learning Representations (2020).
Duvenaud, D. K. et al. Convolutional networks on graphs for learning molecular fingerprints. Adv. Neural Inform. Process. Syst. 28, 1–9 (2015).
Fung, V., Zhang, J., Juarez, E. & Sumpter, B. G. Benchmarking graph neural networks for materials chemistry. npj Comput. Mater. 7, 1–8 (2021).
Schmidt, J., Pettersson, L., Verdozzi, C., Botti, S. & Marques, M. A. L. Crystal graph attention networks for the prediction of stable materials. Sci. Adv. 7, eabi7948 (2021).
Choudhary, K. & DeCost, B. Atomistic line graph neural network for improved materials property predictions. npj Comput. Mater. 7, 1–8 (2021).
Chen, C., Ye, W., Zuo, Y., Zheng, C. & Ong, S. P. Graph networks as a universal machine learning framework for molecules and crystals. Chem. Mater. 31, 3564–3572 (2019).
Park, C. W. & Wolverton, C. Developing an improved crystal graph convolutional neural network framework for accelerated materials discovery. Phys. Rev. Mater. 4, 063801 (2020).
Xie, T. & Grossman, J. C. Hierarchical visualization of materials space with graph convolutional neural networks. J. Chem. Phys. 149, 174111 (2018).
Nguyen, A. H. & Molinero, V. Identification of clathrate hydrates, hexagonal ice, cubic ice, and liquid water in simulations: the CHILL+ algorithm. J. Phys. Chem. B 119, 9369–9376 (2015).
Ziletti, A., Kumar, D., Scheffler, M. & Ghiringhelli, L. M. Insightful classification of crystal structures using deep learning. Nat. Commun. 9, 1–10 (2018).
Ong, S. P. et al. Python materials genomics (pymatgen): a robust, open-source python library for materials analysis. Comput. Mater. Sci. 68, 314–319 (2013).
Leitherer, A., Ziletti, A. & Ghiringhelli, L. M. Robust recognition and exploratory analysis of crystal structures via Bayesian deep learning. Nat. Commun. 12, 1–13 (2021).
Hoffmann, R., Kabanov, A. A., Golov, A. A. & Proserpio, D. M. Homo citans and carbon allotropes: for an ethics of citation. Angew. Chem. Int. Ed. 55, 10962–10976 (2016).
Sharma, B. R., Manjanath, A. & Singh, A. K. pentahexoctite: a new two-dimensional allotrope of carbon. Sci. Rep. 4, 1–6 (2014).
Banik, S. et al. A continuous action space tree search for INverse desiGn (CASTING) framework for materials discovery. Preprint at https://doi.org/10.48550/arXiv.2212.12106 (2022).
Los, J. H. & Fasolino, A. Intrinsic long-range bond-order potential for carbon: Performance in Monte Carlo simulations of graphitization. Phys. Rev. B 68, 024107 (2003).
Manna, S. et al. A database of low-energy atomically precise nanoclusters. Preprint at https://doi.org/10.26434/chemrxiv-2021-0fq3q (2021).
Manna, S. et al. Learning in continuous action space for developing high dimensional potential energy models. Nat. Commun. 13, 1–10 (2022).
Novoselov, K. S. et al. Two-dimensional atomic crystals. Proc. Natl Acad. Sci. USA 102, 10451–10453 (2005).
Faken, D. & Jónsson, H. Systematic analysis of local atomic structure combined with 3D computer graphics. Comput. Mater. Sci. 2, 279–286 (1994).
Tsuzuki, H., Branicio, P. S. & Rino, J. P. Structural characterization of deformed crystals by analysis of common atomic neighborhood. Comput. Phys. Commun. 177, 518–523 (2007).
Polak, W. Z. Efficiency in identification of internal structure in simulated monoatomic clusters: comparison between common neighbor analysis and coordination polyhedron method. Comput. Mater. Sci. 201, 110882 (2022).
Hirel, P. Atomsk: a tool for manipulating and converting atomic data files. Comput. Phys. Commun. 197, 212–219 (2015).
Schubert, E., Sander, J., Ester, M., Kriegel, H. P. & Xu, X. DBSCAN revisited, revisited: why and how you should (still) use DBSCAN. ACM Trans. Database Syst. 42, 1–21 (2017).
Chan, H., Cherukara, M., Loeffler, T. D., Narayanan, B. & Sankaranarayanan, S. K. R. S. Machine learning enabled autonomous microstructural characterization in 3D samples. npj Comput. Mater. 6, 1–9 (2020).
Stukowski, A. Visualization and analysis of atomistic simulation data with OVITO–the open visualization tool. Model. Simul. Mater. Sci. Eng. 18, 015012 (2009).
Chen, Y. et al. Pressure-induced phase transformation in β-eucryptite: an X-ray diffraction and density functional theory study. Scr. Mater. 122, 64–67 (2016).
Chen, Y., Manna, S., Ciobanu, C. V. & Reimanis, I. E. Thermal regimes of Li‐ion conductivity in β‐eucryptite. J. Am. Ceramic Soc. 101, 347–355 (2018).
Bertolazzo, A. A., Dhabal, D. & Molinero, V. Polymorph selection in zeolite synthesis occurs after nucleation. J. Phys. Chem. Lett. 13, 977–981 (2022).
Bertolazzo, A. A., Dhabal, D., Lopes, L. J. S., Walker, S. K. & Molinero, V. Unstable and metastable mesophases can assist in the nucleation of porous crystals. J. Phys. Chem. C 126, 3776–3786 (2022).
Kumar, A. & Molinero, V. Two-step to one-step nucleation of a zeolite through a metastable gyroid mesophase. J. Phys. Chem. Lett. 9, 5692–5697 (2018).
Kumar, A., Nguyen, A. H., Okumu, R., Shepherd, T. D. & Molinero, V. Could mesophases play a role in the nucleation and polymorph selection of zeolites? J. Am. Chem. Soc. 140, 16071–16086 (2018).
Kumar, A. & Molinero, V. Self-assembly of mesophases from nanoparticles. J. Phys. Chem. Lett. 8, 5053–5058 (2017).
Hustad, P. D., Marchand, G. R., Garcia-Meitin, E. I., Roberts, P. L. & Weinhold, J. D. Photonic polyethylene from self-assembled mesophases of polydisperse olefin block copolymers. Macromolecules 42, 3788–3794 (2009).
Kumar, A. & Molinero, V. Why is gyroid more difficult to nucleate from disordered liquids than lamellar and hexagonal mesophases? J. Phys. Chem. B 122, 4758–4770 (2018).
Moore, E. B. & Molinero, V. Ice crystallization in water’s “no-man’s land”. J. Chem. Phys. 132, 244504 (2010).
Moore, E. B., De La Llave, E., Welke, K., Scherlis, D. A. & Molinero, V. Freezing, melting and structure of ice in a hydrophilic nanopore. Phys. Chem. Chem. Phys. 12, 4124–4134 (2010).
García Fernández, R., Abascal, J. L. F. & Vega, C. The melting point of ice I h for common water models calculated from direct coexistence of the solid-liquid interface. J. Chem. Phys. 124, 144506 (2006).
Carignano, M. A., Shepson, P. B. & Szleifer, I. Molecular dynamics simulations of ice growth from supercooled water. Mol. Phys. 103, 2957–2967 (2005).
Abascal, J. L. F. & Vega, C. A general purpose model for the condensed phases of water: TIP4P/2005. The J. Chem. Phys. 123, 234505 (2005).
Acknowledgements
The authors acknowledge support from the US Department of Energy through BES award DE-SC0021201. This material is based on work supported by the DOE, Office of Science, BES Data, Artificial Intelligence and Machine Learning at DOE Scientific User Facilities programme (MLExchange). Use of the Center for Nanoscale Materials, an Office of Science user facility, was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. This research also used resources from the Argonne Leadership Computing Facility at Argonne National Laboratory, which is supported by the Office of Science of the U.S. Department of Energy under contract DE-AC02-06CH11357. This research used resources of the National Energy Research Scientific Computing Center; a DOE Office of Science User Facility supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231. We gratefully acknowledge the computing resources provided via high-performance computing clusters operated by the Laboratory Computing Resource Center (LCRC) at Argonne National Laboratory.
Author information
Authors and Affiliations
Contributions
S.B. and S.K.R.S.S. conceived the project. S.B. developed the CEGANN workflow for multiscale classification. H.C. provided feedback on the workflow. S.B. evaluated the performance of workflow on different classification tasks and analyzed the results. S.B. wrote the manuscript with guidance from S.K.R.S.S. and V.M. D.D. and V.M. generated the dataset and contributed to the writing, and analysis of the dynamical classification, Mesophase, and Ice growth characterization task presented in the manuscript. S.M., H.C., and D.D. provided feedback on the manuscript. M.C. assisted with computing resources. All authors participated in discussing the results and provided comments and suggestions on the various sections of the manuscript. S.K.R.S.S. supervised and directed the overall project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Banik, S., Dhabal, D., Chan, H. et al. CEGANN: Crystal Edge Graph Attention Neural Network for multiscale classification of materials environment. npj Comput Mater 9, 23 (2023). https://doi.org/10.1038/s41524-023-00975-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-023-00975-z
This article is cited by
-
A Continuous Action Space Tree search for INverse desiGn (CASTING) framework for materials discovery
npj Computational Materials (2023)