Abstract
AlphaFold2 revolutionized structural biology with the ability to predict protein structures with exceptionally high accuracy. Its implementation, however, lacks the code and data required to train new models. These are necessary to (1) tackle new tasks, like protein–ligand complex structure prediction, (2) investigate the process by which the model learns and (3) assess the model’s capacity to generalize to unseen regions of fold space. Here we report OpenFold, a fast, memory efficient and trainable implementation of AlphaFold2. We train OpenFold from scratch, matching the accuracy of AlphaFold2. Having established parity, we find that OpenFold is remarkably robust at generalizing even when the size and diversity of its training set is deliberately limited, including near-complete elisions of classes of secondary structure elements. By analyzing intermediate structures produced during training, we also gain insights into the hierarchical manner in which OpenFold learns to fold. In sum, our studies demonstrate the power and utility of OpenFold, which we believe will prove to be a crucial resource for the protein modeling community.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
OpenProteinSet and OpenFold model parameters are hosted on the Registry of Open Data on AWS and can be accessed at https://registry.opendata.aws/openfold/. Both are available under the permissive CC BY 4.0 license. Throughout the study, we use validation sets derived from the PDB via CAMEO. We also use CASP evaluation sets. Source data are provided with this paper.
Code availability
OpenFold can be accessed at https://github.com/aqlaboratory/openfold. It is available under the permissive Apache 2 Licence.
References
Anfinsen, C. B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973).
Dill, K. A., Ozkan, S. B., Shell, M. S. & Weikl, T. R. The protein folding problem. Annu. Rev. Biophys. 37, 289–316 (2008).
Jones, D. T., Singh, T., Kosciolek, T. & Tetchner, S. MetaPSICOV: combining coevolution methods for accurate prediction of contacts and long range hydrogen bonding in proteins. Bioinformatics 31, 999–1006 (2015).
Golkov, V. et al. Protein contact prediction from amino acid co-evolution using convolutional networks for graph-valued images. In Advances in Neural Information Processing Systems (eds Lee, D. et al.) (Curran Associates, 2016).
Wang, S., Sun, S., Li, Z., Zhang, R. & Xu, J. Accurate de novo prediction of protein contact map by ultra-deep learning model. PLoS Comput. Biol. 13, e1005324 (2017).
Liu, Y., Palmedo, P., Ye, Q., Berger, B. & Peng, J. Enhancing evolutionary couplings with deep convolutional neural networks. Cell Syst. 6, 65–74 (2018).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
Xu, J., McPartlon, M. & Li, J. Improved protein structure prediction by deep learning irrespective of co-evolution information. Nat. Mach. Intell. 3, 601–609 (2021).
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 577, 583–589 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Baek, M. Adding a big enough number for ‘residue_index’ feature is enough to model hetero-complex using AlphaFold (green&cyan: crystal structure / magenta: predicted model w/ residue_index modification). Twitter twitter.com/minkbaek/status/1417538291709071362?lang=en (2021).
Tsaban, T. et al. Harnessing protein folding neural networks for peptide–protein docking. Nat. Commun. 13, 176 (2022).
Roney, J. P. & Ovchinnikov, S. State-of-the-art estimation of protein model accuracy using AlphaFold. Phys. Rev. Lett. 129, 238101 (2022).
Baltzis, A. et al. Highly significant improvement of protein sequence alignments with AlphaFold2. Bioinformatics 38, 5007–5011 (2022).
Bryant, P., Pozzati, G. & Elofsson, A. Improved prediction of protein–protein interactions using AlphaFold2. Nat. Commun. 13, 1265 (2022).
Wayment-Steele, H. K., Ovchinnikov, S., Colwell, L. & Kern, D. Prediction of multiple conformational states by combining sequence clustering with AlphaFold2. Nature 625, 832–839 (2024).
Tunyasuvunakool, K. et al. Highly accurate protein structure prediction for the human proteome. Nature 596, 590–596 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2021).
Callaway, E. ‘The entire protein universe’: AI predicts shape of nearly every known protein. Nature 608, 15–16 (2022).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2021).
Ahdritz, G. et al. OpenProteinSet: training data for structural biology at scale. In Advances in Neural Information Processing Systems (eds Oh, A. et al.) 4597-4609 (Curran Associates, 2023).
Paszke, A. et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Advances in Neural Information Processing Systems (eds Wallach, H. et al.) 8026–8037 (Curran Associates, 2019).
Bradbury, J. et al. JAX: composable transformations of Python+NumPy programs. GitHub github.com/google/jax (2018).
Rasley, J., Rajbhandari, S., Ruwase, O. & He, Y. DeepSpeed: system optimizations enable training deep learning models with over 100 billion parameters. In Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, KDD ’20 3505–3506 (Association for Computing Machinery, 2020).
Charlier, B., Feydy, J., Glaunès, J., Collin, F.-D. & Durif, G. Kernel operations on the GPU, with autodiff, without memory overflows. J. Mach. Learn. Res. 22, 1–6 (2021).
Falcon, W. & the PyTorch Lightning team. PyTorch Lightning (PyTorch Lightning, 2019).
Dao, T., Fu, D. Y., Ermon, S., Rudra, A. & Ré, C. FlashAttention: fast and memory-efficient exact attention with IO-awareness. In Advances in Neural Information Processing Systems (eds Koyejo, S. et al.) 16344–16359 (Curran Associates, 2022).
Mirdita, M. et al. Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res. 45, D170–D176 (2017).
wwPDB Consortium. Protein Data Bank: the single global archive for 3D macromolecular structure data. Nucleic Acids Res. 47, D520–D528 (2018).
Haas, J. ürgen et al. Continuous automated model evaluation (CAMEO) complementing the critical assessment of structure prediction in CASP12. Proteins 86, 387–398 (2018).
Mariani, V., Biasini, M., Barbato, A. & Schwede, T. lDDT: a local superposition-free score for comparing protein structures and models using distance difference tests. Bioinformatics 29, 2722–2728 (2013).
Orengo, C. A. et al. CATH—a hierarchic classification of protein domain structures. Structure 5, 1093–1108 (1997).
Sillitoe, I. et al. CATH: increased structural coverage of functional space. Nucleic Acids Res. 49, D266–D273 (2021).
Andreeva, A., Kulesha, E., Gough, J. & Murzin, A. G. The SCOP database in 2020: expanded classification of representative family and superfamily domains of known protein structures. Nucleic Acids Res. 48, D376–D382 (2020).
Saitoh, Y. et al. Structural basis for high selectivity of a rice silicon channel Lsi1. Nat. Commun. 12, 6236 (2021).
Mota, DaniellyC. A. M. et al. Structural and thermodynamic analyses of human TMED1 (p241) Golgi dynamics. Biochimie 192, 72–82 (2022).
Vaswani, A. et al. Attention is all you need. In Advances in Neural Information Processing Systems (eds Guyon, I. et al.) (Curran Associates, 2017).
Rabe, M. N. & Staats, C. Self-attention does not need O(n2) memory. Preprint at https://doi.org/10.48550/arXiv.2112.05682 (2021).
Cheng, S. et al. FastFold: Optimizing AlphaFold Training and Inference on GPU Clusters. In Proceedings of the 29th ACM SIGPLAN Annual Symposium on Principles and Practice of Parallel Programming 417–430 (Association for Computing Machinery, 2024).
Li, Z. et al. Uni-Fold: an open-source platform for developing protein folding models beyond AlphaFold. Preprint at bioRxiv https://doi.org/10.1101/2022.08.04.502811 (2022).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Science 22, 2577–2637 (1983).
Zemla, A. LGA: a method for finding 3D similarities in protein structures. Nucleic Acids Res. 31, 3370–3374 (2003).
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS ONE 6, e28766 (2011).
Sułkowska, J. I., Morcos, F., Weigt, M., Hwa, T. & Onuchic, José Genomics-aided structure prediction. Proc. Natl Acad. Sci. USA 109, 10340–10345 (2012).
Kaplan, J. et al. Scaling laws for neural language models. Preprint at https://doi.org/10.48550/arXiv.2001.08361 (2020).
Hoffmann, J. et al. An empirical analysis of compute-optimal large language model training. In Advances in Neural Information Processing Systems (eds Oh, A. H. et al.) 30016–30030 (NeurIPS, 2022).
Tay, Y. et al. Scaling laws vs model architectures: how does inductive bias influence scaling? In Findings of the Association for Computational Linguistics: EMNLP 2023 (eds Bouamor, H. et al.) 12342–12364 (Association for Computational Linguistics, 2023).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Alley, E. C., Khimulya, G., Biswas, S., AlQuraishi, M. & Church, G. M. Unified rational protein engineering with sequence-based deep representation learning. Nat. Methods 16, 1315–1322 (2019).
Chowdhury, R. et al. Single-sequence protein structure prediction using a language model and deep learning. Nat. Biotechnol. 40, 1617–1623 (2022).
Wu, R. et al. High-resolution de novo structure prediction from primary sequence. Preprint at bioRxiv https://doi.org/10.1101/2022.07.21.500999 (2022).
Singh, J., Paliwal, K., Litfin, T., Singh, J. & Zhou, Y. Predicting RNA distance-based contact maps by integrated deep learning on physics-inferred secondary structure and evolutionary-derived mutational coupling. Bioinformatics 38, 3900–3910 (2022).
Baek, M., McHugh, R., Anishchenko, I., Baker, D. & DiMaio, F. Accurate prediction of protein–nucleic acid complexes using RoseTTAFoldNA. Nat. Methods 21, 117–121 (2024).
Pearce, R., Omenn, G. S. & Zhang, Y. De novo RNA tertiary structure prediction at atomic resolution using geometric potentials from deep learning. Preprint at bioRxiv https://doi.org/10.1101/2022.05.15.491755 (2022).
McPartlon, M., Lai, B. & Xu, J. A deep SE(3)-equivariant model for learning inverse protein folding. Preprint at bioRxiv https://doi.org/10.1101/2022.04.15.488492 (2022).
McPartlon, M. & Xu, J. An end-to-end deep learning method for protein side-chain packing and inverse folding. In Proceedings of the National Academy of Sciences e2216438120 (PNAS, 2023).
Knox, H. L., Sinner, E. K., Townsend, C. A., Boal, A. K. & Booker, S. J. Structure of a B12-dependent radical SAM enzyme in carbapenem biosynthesis. Nature 602, 343–348 (2022).
Zhang, Y. & Skolnick, J. Scoring function for automated assessment of protein structure template quality. Proteins 57, 702–710 (2004).
Rajbhandari, S., Rasley, J., Ruwase, O. & He, Y. Zero: memory optimizations toward training trillion parameter models. In Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis (IEEE Press, 2020).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. In 3rd International Conference on Learning Representations (eds Bengio, Y. & LeCun, Y.) (ICLR, 2015).
Wang, G. et al. HelixFold: an efficient implementation of AlphaFold2 using PaddlePaddle. Preprint at https://doi.org/10.48550/arXiv.2207.05477 (2022).
Yuan, J. et al. OneFlow: redesign the distributed deep learning framework from scratch. Preprint at https://doi.org/10.48550/arXiv.2110.15032 (2021).
Ovchinnikov, S. Weekend project! nerd-face So now that OpenFold weights are available. I was curious how different they are from AlphaFold weights and if they can be used for AfDesign evaluation. More specifically, if you design a protein with AlphaFold, can OpenFold predict it (and vice-versa)? (1/5). Twitter twitter.com/sokrypton/status/1551242121528520704?lang=en (2022).
Wei, X. et al. The α-helical cap domain of a novel esterase from gut Alistipes shahii shaping the substrate-binding pocket. J. Agric. Food Chem. 69, 6064–6072 (2021).
Carroll, B. L. et al. Caught in motion: human NTHL1 undergoes interdomain rearrangement necessary for catalysis. Nucleic Acids Res. 49, 13165–13178 (2021).
Acknowledgements
We thank the Flatiron Institute, OpenBioML, Stability AI, the Texas Advanced Computing Center and NVIDIA for providing compute for experiments in this paper. Individually, we thank M. Mirdita, M. Steinegger and S. Ovchinnikov for valuable support and expertise. This research used resources of the National Energy Research Scientific Computing Center, which is supported by the Office of Science of the US Department of Energy under contract no. DE-AC02-05CH11231. We acknowledge the Texas Advanced Computing Center at the University of Texas at Austin for providing HPC resources that have contributed to the research results reported within this paper. G.A. is supported by a Simons Investigator Fellowship, NSF grant DMS-2134157, DARPA grant W911NF2010021, DOE grant DE-SC0022199 and a graduate fellowship from the Kempner Institute at Harvard University. N.B. is supported by DARPA Panacea program grant HR0011-19-2-0022 and NCI grant U54-CA225088. C.F. and S.K. are supported by NIH grant R35GM150546. B.Z. and Z.Z. are supported by grants NSF OAC-2112606 and OAC-2106661. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
G.A. wrote and optimized the OpenFold codebase, generated data, trained the model, performed experiments and maintained the GitHub repository. C.F. wrote and tested code for the OpenFold implementation of AlphaFold-Multimer. S.K. and W.G. wrote data preprocessing code. G.A., N.B. and M.A. conceived of and managed the project, designed experiments, analyzed results and wrote the manuscript. G.A., B.Z., Z.Z., N.Z. and A.N. ran ablations. All authors read and approved the manuscript. The Flatiron Institute (via I.F., A.M.W., S.R. and R.B.) provided compute for ablations, all data generation and our main training experiments. NVIDIA (A.N., B. Wang, M.M.S.-D., S.Z., A.O., M.E.G. and P.R.L.) performed training stability experiments, fixed critical bugs in the codebase, added new model features and provided compute for ablations. Stability AI (via N.Z., S.B. and E.M.) provided compute for ablations. The DeepSpeed team at Microsoft (S.C., M.Z., C.L., S.L.S. and Y.H.) wrote custom optimized attention kernels. Q.X. and T.J.O.’D. debugged code and provided feedback.
Corresponding authors
Ethics declarations
Competing interests
M.A. is a member of the scientific advisory boards of Cyrus Biotechnology, Deep Forest Sciences, Nabla Bio, Oracle Therapeutics and FL2021-002, a Foresite Labs company. P.K.S. is a cofounder and member of the BOD of Glencoe Software, member of the BOD for Applied BioMath and a member of the SAB for RareCyte, NanoString, Reverb Therapeutics and Montai Health; he holds equity in Glencoe, Applied BioMath and RareCyte. L.N. is an employee of Cyrus Biotechnology. The other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Arunima Singh, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
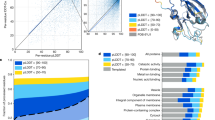
Extended Data Fig. 1 OpenFold matches the accuracy of AlphaFold2 on CASP15 targets.
Scatter plot of GDT-TS values of AlphaFold and OpenFold ‘Model 1’ predictions against all currently available ‘all groups’ CASP15 domains (n = 90). OpenFold’s mean accuracy (95% confidence interval = 68.6-78.8) is on par with AlphaFold’s (95% confidence interval = 69.7-79.2) and OpenFold does at least as well as the latter on exactly 50% of targets. Confidence intervals of each mean are estimated from 10,000 bootstrap samples.
Extended Data Fig. 3 Fine-tuning does not materially improve prediction accuracy on long proteins.
Mean lDDT-Cα over validation proteins with at least 500 residues as a function of fine-tuning step.
Extended Data Fig. 4 The ‘Mostly alpha’ CATH class contains some beta sheets, and vice versa.
Counts for alpha helices and beta sheets in the mostly alpha and mostly beta CATH class-stratified training sets from Fig. 2, based on 1,000 random samples. Counts are binned by size, defined as the number of residues for alpha helices and number of strands for beta sheets.
Extended Data Fig. 5 Reduced dataset diversity disproportionately affects global structure.
Mean GDT-TS and lDDT-Cα of non-overlapping protein fragments from CAMEO validation set as a function of the percentage of CATH clusters in elided training sets. Data for both topology and architecture elisions are included. The fragmenting procedure is the same as that described in Fig. 5a.
Extended Data Fig. 6 Early predictions crudely approximate lower-dimensional PCA projections.
(A) Mean dRMSD, as a function of training step, between low- dimensional PCA projections of predicted structures and the final 3D prediction at step 5,000 (denoted by *). Averages are computed over the CAMEO validation set. Insets show idealized behavior corresponding to unstaggered, simultaneous growth in all dimensions and perfectly staggered growth. Empirical training behavior more closely resembles the staggered scenario. (B) Low-dimensional projections as in (A) compared to projections of the final predicted structures at step 5,000. (C) Mean displacement, as a function of training step, of C? atoms along the directions of their final structure’s PCA eigenvectors. Results are shown for two individual proteins (PDB accession codes 7DQ9_A ref. 66 and 7RDT_A ref. 67). Shaded regions correspond loosely to ‘1D,’ ‘2D,’ and ‘3D’ phases of dimensionality.
Extended Data Fig. 7 Radius of gyration as an order parameter for learning protein phase structure.
Radii of gyration for proteins in the CAMEO validation set (or- ange) as a function of sequence length over training time, plotted on a log-log scale against experimental structures (blue). Legends show equations of best fit curves, computed using non-linear least squares. The training steps chosen correspond loosely to four phases of dimensional growth. See Supplementary Information section B.3 for extended discussion.
Extended Data Fig. 8 Contact prediction for beta sheets at different ranges.
Binned contact F1 scores (8 Å threshold) for beta sheets of various widths as a function of training step at different residue-residue separation ranges (SMLR ≥ 6 residues apart; LR ≥ 24 residues apart, as in8). Sheet widths are weighted averages of sheet thread counts within each bin, as in Fig. 5b.
Supplementary information
Supplementary Information
Supplementary Discussion
Supplementary Video 1
Folding animation for PDB protein 7B3A, chain A. Predictions are from successive early checkpoints of an OpenFold model (training step is shown at the bottom left).
Supplementary Video 2
Folding animation for PDB protein 7DMF, chain A. Predictions are from successive early checkpoints of an OpenFold model (training step is shown at the bottom left).
Supplementary Video 3
Folding animation for PDB protein 7DQ9, chain A. Predictions are from successive early checkpoints of an OpenFold model (training step is shown at the bottom left).
Supplementary Video 4
Folding animation for PDB protein 7LBU, chain A. Predictions are from successive early checkpoints of an OpenFold model (training step is shown at the bottom left).
Supplementary Video 5
Folding animation for PDB protein 7RDT, chain A. Predictions are from successive early checkpoints of an OpenFold model (training step is shown at the bottom left).
Source data
Source Data Fig. 1
Numerical source data.
Source Data Fig. 2
Numerical source data.
Source Data Fig. 3
Numerical source data.
Source Data Fig. 4
Numerical source data.
Source Data Fig. 5
Numerical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ahdritz, G., Bouatta, N., Floristean, C. et al. OpenFold: retraining AlphaFold2 yields new insights into its learning mechanisms and capacity for generalization. Nat Methods (2024). https://doi.org/10.1038/s41592-024-02272-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41592-024-02272-z