Abstract
Single-cell transcriptomics can profile thousands of cells in a single experiment and identify novel cell types, states and dynamics in a wide variety of tissues and organisms. Standard experimental protocols and analysis workflows have been developed to create single-cell transcriptomic maps from tissues. This tutorial focuses on how to interpret these data to identify cell types, states and other biologically relevant patterns with the objective of creating an annotated map of cells. We recommend a three-step workflow including automatic cell annotation (wherever possible), manual cell annotation and verification. Frequently encountered challenges are discussed, as well as strategies to address them. Guiding principles and specific recommendations for software tools and resources that can be used for each step are covered, and an R notebook is included to help run the recommended workflow. Basic familiarity with computer software is assumed, and basic knowledge of programming (e.g., in the R language) is recommended.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
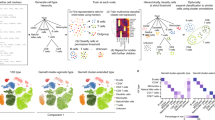
The data used to generate this tutorial are openly available at the following sources. The sequence data used to generate Figs. 1 and 4 are available from MacParland et al.100 through the NCBI GEO accession GSE115469. The analyzed data from which the map was directly created can also be accessed interactively as the R package HumanLiver from https://github.com/BaderLab/HumanLiver. The sequence data used to generate Figs. 3 and 7 are available from 10x Genomics and can be downloaded from https://support.10xgenomics.com/single-cell-gene-expression/datasets. The sequence data used to generate Fig. 6 are available through the NCBI GEO accession GSE129788, as reported by Ximerakis et al.101. The analyzed data can be accessed interactively at http://shiny.baderlab.org/AgingMouseBrain/. The human bulk RNA-seq data used to generate the reference data set in the accompanying R code (https://github.com/BaderLab/CellAnnotationTutorial and https://codeocean.com/capsule/d67541eb-43f8-4cae-a258-5ef0069e5301/) are available from the Database of Immune Cell Expression and can be downloaded in R through the package ‘celldex’43 by the command DatabaseImmuneCellExpressionData(). The query data set used in the accompanying R code is available from 10x Genomics and can be downloaded from https://cf.10xgenomics.com/samples/cell-exp/1.1.0/pbmc3k/pbmc3k_filtered_gene_bc_matrices.tar.gz. The collection of PBMC marker genes used in the accompanying R code is available from Diaz-Mejia et al.37 with read data from NCBI Sequence Read Archive accession number SRX1723926. The supplementary data from the Diaz-Mejia et al. paper can be accessed at https://zenodo.org/record/3369934/#.X3CGN5NKjGI. The Gene Matrix Transposed (GMT) file used for pathway analysis in the accompanying R code can be downloaded from http://download.baderlab.org/EM_Genesets/current_release/Human/symbol/Pathways/Human_MSigdb_March_01_2021_symbol.gmt.
Code availability
An R script that implements the main workflow described in this proposal is available at https://github.com/BaderLab/CellAnnotationTutorial, and a version-controlled copy is available through Code Ocean at https://codeocean.com/capsule/d67541eb-43f8-4cae-a258-5ef0069e5301/.
References
Sasagawa, Y., Hayashi, T. & Nikaido, I. Strategies for converting RNA to amplifiable cDNA for single-cell RNA sequencing methods. Adv. Exp. Med. Biol. 1129, 1–17 (2019).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Han, X. et al. Construction of a human cell landscape at single-cell level. Nature 581, 303–309 (2020).
Tabula Muris Consortium. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Grün, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
Xia, B. & Yanai, I. A periodic table of cell types. Development 146, dev169854 (2019).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943.e22 (2019).
Ziegenhain, C. et al. Comparative analysis of single-cell RNA sequencing methods. Mol. Cell 65, 631–643.e4 (2017).
Lafzi, A., Moutinho, C., Picelli, S. & Heyn, H. Tutorial: guidelines for the experimental design of single-cell RNA sequencing studies. Nat. Protoc. 13, 2742–2757 (2018).
Hwang, B., Lee, J. H. & Bang, D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp. Mol. Med. 50, 1–14 (2018).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Henry, G. H., Mathews, J. A. & Malladi, V. S. BICF Cellranger count analysis workflow (version publish_1.2.0). Zenodo. https://zenodo.org/record/3373749#.YGzmGhRucdU (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol 19, 15 (2018).
Duò, A., Robinson, M. D. & Soneson, C. A systematic performance evaluation of clustering methods for single-cell RNA-seq data. F1000Res. 7, 1141 (2018).
Kiselev, V. Y., Andrews, T. S. & Hemberg, M. Challenges in unsupervised clustering of single-cell RNA-seq data. Nat. Rev. Genet. 20, 273–282 (2019).
Menon, V. Clustering single cells: a review of approaches on high-and low-depth single-cell RNA-seq data. Brief. Funct. Genomics 17, 240–245 (2018).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. (86), 2579–2605 (2008).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Gene Set Enrichment Analysis. Archived: SCSig collection: Signatures of Single Cell Identities; https://www.gseamsigdb.org/gsea/msigdb/supplementary_genesets.jsp#SCSig
Franzén, O., Gan, L.-M. & Björkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA sequencing data. Database (Oxford) 2019, baz046 (2019).
Zhang, X. et al. CellMarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 47, D721–D728 (2019).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Papatheodorou, I. et al. Expression Atlas update: from tissues to single cells. Nucleic Acids Res. 48, D77–D83 (2020).
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
HuBMAP Consortium. The human body at cellular resolution: the NIH Human Biomolecular Atlas Program. Nature 574, 187–192 (2019).
Yuzwa, S. A. et al. Developmental emergence of adult neural stem cells as revealed by single-cell transcriptional profiling. Cell Rep. 21, 3970–3986 (2017).
Kurial, S. N. T. & Willenbring, H. Transcriptomic traces of adult human liver progenitor cells. Hepatology 71, 1504–1507 (2020).
Stanley, G., Gokce, O., Malenka, R. C., Südhof, T. C. & Quake, S. R. Continuous and discrete neuron types of the adult murine striatum. Neuron 105, 688–699.e8 (2020).
Satpathy, A. Curated, multi-omic, ML-driven single-cell atlas for characterizing the human immune system across disease states. J. Immunol. 204, 11–159.11 (2020).
Abdelaal, T. et al. A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Biol. 20, 194 (2019).
Zhang, Z. et al. SCINA: a semi-supervised subtyping algorithm of single cells and bulk samples. Genes (Basel) 10, 531 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Diaz-Mejia, J. J. et al. Evaluation of methods to assign cell type labels to cell clusters from single-cell RNA-sequencing data. [version 3; peer review: 2 approved, 1 approved with reservations]. F1000Res. 8, ISCB Comm J-296 (2019).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Han, X. et al. Mapping the mouse cell atlas by Microwell-seq. Cell 172, 1091–1107.e17 (2018).
Regev, A. et al. The Human Cell Atlas White Paper. https://doi.org/10.17863/CAM.40032 (2017).
Kiselev, V. Y., Yiu, A. & Hemberg, M. scmap: projection of single-cell RNA-seq data across data sets. Nat. Methods 15, 359–362 (2018).
Tan, Y. & Cahan, P. SingleCellNet: a computational tool to classify single cell RNA-seq data across platforms and across species. Cell Syst. 9, 207–213.e2 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Bais, A. S. & Kostka, D. scds: computational annotation of doublets in single-cell RNA sequencing data. Bioinformatics 36, 1150–1158 (2020).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Niwa, H. The principles that govern transcription factor network functions in stem cells. Development 145, dev157420 (2018).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Clark, J. Z. et al. Representation and relative abundance of cell-type selective markers in whole-kidney RNA-Seq data. Kidney Int. 95, 787–796 (2019).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Dal Molin, A., Baruzzo, G. & Di Camillo, B. Single-cell RNA-sequencing: assessment of differential expression analysis methods. Front. Genet. 8, 62 (2017).
Adossa, N. A., Schauser, L., Gregersen, V. G. & Elo, L. L. Feature extraction approach in single-cell gene expression profiling for cell-type marker identification. Preprint at bioRxiv https://doi.org/10.1101/686659 (2019).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Diehl, A. D. et al. The Cell Ontology 2016: enhanced content, modularization, and ontology interoperability. J. Biomed. Semantics 7, 44 (2016).
Meehan, T. F. et al. Logical development of the cell ontology. BMC Bioinformatics 12, 6 (2011).
Aevermann, B. D. et al. Cell type discovery using single-cell transcriptomics: implications for ontological representation. Hum. Mol. Genet. 27, R40–R47 (2018).
Hsiao, C. J. et al. Characterizing and inferring quantitative cell cycle phase in single-cell RNA-seq data analysis. Genome Res. 30, 611–621 (2020).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308.e36 (2018).
Adler, M., Korem Kohanim, Y., Tendler, A., Mayo, A. & Alon, U. Continuum of gene-expression profiles provides spatial division of labor within a differentiated cell type. Cell Syst. 8, 43–52.e5 (2019).
Liu, S. & Trapnell, C. Single-cell transcriptome sequencing: recent advances and remaining challenges. F1000Res. 5, F1000 Faculty Rev-182 (2016).
Schumacher, L. J. Neural crest migration with continuous cell states. J. Theor. Biol. 481, 84–90 (2019).
Chung, N. C. Statistical significance of cluster membership for unsupervised evaluation of cell identities. Bioinformatics 36, 3107–3114 (2020).
Rosati, E. et al. Overview of methodologies for T-cell receptor repertoire analysis. BMC Biotechnol. 17, 61 (2017).
Setliff, I. et al. High-throughput mapping of B cell receptor sequences to antigen specificity. Cell 179, 1636–1646.e15 (2019).
Park, D. et al. Differences in the molecular signatures of mucosal-associated invariant T cells and conventional T cells. Sci. Rep. 9, 7094 (2019).
Moter, A. & Göbel, U. B. Fluorescence in situ hybridization (FISH) for direct visualization of microorganisms. J. Microbiol. Methods 41, 85–112 (2000).
Ren, X. et al. Reconstruction of cell spatial organization from single-cell RNA sequencing data based on ligand-receptor mediated self-assembly. Cell Res. 30, 763–778 (2020).
Porter, J. R., Telford, W. G. & Batchelor, E. Single-cell gene expression profiling using FACS and qPCR with internal standards. J. Vis. Exp. (120), 55219 (2017).
Wu, A. R. et al. Quantitative assessment of single-cell RNA-sequencing methods. Nat. Methods 11, 41–46 (2014).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Liu, F. et al. Systematic comparative analysis of single-nucleotide variant detection methods from single-cell RNA sequencing data. Genome Biol. 20, 242 (2019).
Fan, J. et al. Linking transcriptional and genetic tumor heterogeneity through allele analysis of single-cell RNA-seq data. Genome Res. 28, 1217–1227 (2018).
Serin Harmanci, A., Harmanci, A. O. & Zhou, X. CaSpER identifies and visualizes CNV events by integrative analysis of single-cell or bulk RNA-sequencing data. Nat. Commun. 11, 89 (2020).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Tickle, T., Gc Ti, Brown, M. & Haas, B. InferCNV of the Trinity CTAT Project. https://github.com/broadinstitute/inferCNV (Klarman Cell Observatory, Broad Institute of MIT and Harvard, 2019).
AlJanahi, A. A., Danielsen, M. & Dunbar, C. E. An introduction to the analysis of single-cell RNA-sequencing data. Mol. Ther. Methods Clin. Dev. 10, 189–196 (2018).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Zhao, Q. et al. A mitochondrial specific stress response in mammalian cells. EMBO J. 21, 4411–4419 (2002).
Guantes, R. et al. Global variability in gene expression and alternative splicing is modulated by mitochondrial content. Genome Res. 25, 633–644 (2015).
Jiang, L., Chen, H., Pinello, L. & Yuan, G.-C. GiniClust: detecting rare cell types from single-cell gene expression data with Gini index. Genome Biol. 17, 144 (2016).
Innes, B. T. & Bader, G. D. scClustViz – Single-cell RNAseq cluster assessment and visualization. F1000Res. 7, ISCB Comm J-1522 (2018).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. Gigascience 7, giy083 (2018).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
Fleming, S. J., Marioni, J. C. & Babadi, M. CellBender remove-background: a deep generative model for unsupervised removal of background noise from scRNA-seq datasets. Preprint at bioRxiv https://doi.org/10.1101/791699 (2019).
Mohanraj, S. et al. Crescent: cancer single cell expression toolkit. Nucleic Acids Res. 48, W372–W379 (2020).
David, F. P. A., Litovchenko, M., Deplancke, B. & Gardeux, V. ASAP 2020 update: an open, scalable and interactive web-based portal for (single-cell) omics analyses. Nucleic Acids Res. 48, W403–W414 (2020).
Franzén, O. & Björkegren, J. L. M. alona: a web server for single-cell RNA-seq analysis. Bioinformatics 36, 3910–3912 (2020).
Hillje, R., Pelicci, P. G. & Luzi, L. Cerebro: interactive visualization of scRNA-seq data. Bioinformatics 36, 2311–2313 (2020).
Zhang, A. W. et al. Probabilistic cell-type assignment of single-cell RNA-seq for tumor microenvironment profiling. Nat. Methods 16, 1007–1015 (2019).
Miao, Z. et al. Putative cell type discovery from single-cell gene expression data. Nat. Methods 17, 621–628 (2020).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Angermueller, C. et al. Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat. Methods 13, 229–232 (2016).
Baron, M. & Yanai, I. New skin for the old RNA-Seq ceremony: the age of single-cell multi-omics. Genome Biol. 18, 159 (2017).
Guilhamon, P. et al. Chromatin blueprint of glioblastoma stem cells reveals common drug candidates for distinct subtypes. Preprint at bioRxiv https://doi.org/10.1101/370726 (2018).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Ximerakis, M. et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 (2019).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. (85), 2825–2830 (2011).
Van de Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J. P. GSEA-P: a desktop application for Gene Set Enrichment Analysis. Bioinformatics 23, 3251–3253 (2007).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887.e17 (2019).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Wold, S., Esbensen, K. & Geladi, P. Principal component analysis. Chemometr. Intell. Lab. Syst. 2, 37–52 (1987).
Kobak, D. & Berens, P. The art of using t-SNE for single-cell transcriptomics. Nat. Commun. 10, 5416 (2019).
Halladin-Dąbrowska, A., Kania, A. & Kopeć, D. The t-SNE algorithm as a tool to improve the quality of reference data used in accurate mapping of heterogeneous non-forest vegetation. Remote Sens. (Basel) 12, 39 (2019).
Kobak, D. & Linderman, G. C. Initialization is critical for preserving global data structure in both t-SNE and UMAP. Nat. Biotechnol. 39, 156–157 (2019).
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform Manifold Approximation and Projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Ringnér, M. What is principal component analysis? Nat. Biotechnol. 26, 303–304 (2008).
Hicks, S. C., Townes, F. W., Teng, M. & Irizarry, R. A. Missing data and technical variability in single-cell RNA-sequencing experiments. Biostatistics 19, 562–578 (2018).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Clamp, M. et al. Ensembl 2002: accommodating comparative genomics. Nucleic Acids Res. 31, 38–42 (2003).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Geirsdottir, L. et al. Cross-species single-cell analysis reveals divergence of the primate microglia program. Cell 179, 1609–1622.e16 (2019); erratum: 181, 746 (2020).
Ding, H., Blair, A., Yang, Y. & Stuart, J. M. Biological process activity transformation of single cell gene expression for cross-species alignment. Nat. Commun. 10, 4899 (2019).
Acknowledgements
This project has been made possible in part by grant CZF2019-002429 from the Chan Zuckerberg Initiative DAF, an advised fund of the Silicon Valley Community Foundation. We acknowledge Shirley Hui, Daniel Stueckmann and Ronald Xie for their help in reviewing the tutorial.
Author information
Authors and Affiliations
Contributions
Z.A.C., T.S.A., J.A., D.P. and S.A.M. initially wrote different sections of the document and collaboratively joined them together. Z.A.C., T.S.A., J.A., D.P. and B.T.I. designed the figures. Z.A.C. and G.D.B. organized the process and undertook major edits. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Åsa Björklund and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Heat map and UMAP visualizations.
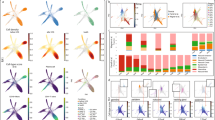
An extension of Fig. 6 incorporating the visualization of marker genes for identified cell types as a heat map (a) and a Mllt11 expression overlaid on a UMAP plot (b). Mllt11 is expressed at various levels across clusters, suggesting a cell-type gradient across NRP, ImmN, NendC and mNEUR. Both plots are generated from scRNA-seq data from young and old mouse brains101.
Supplementary information
Supplementary Information
Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Clarke, Z.A., Andrews, T.S., Atif, J. et al. Tutorial: guidelines for annotating single-cell transcriptomic maps using automated and manual methods. Nat Protoc 16, 2749–2764 (2021). https://doi.org/10.1038/s41596-021-00534-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00534-0
This article is cited by
-
Shaoxia: a web-based interactive analysis platform for single cell RNA sequencing data
BMC Genomics (2024)
-
AnnoSpat annotates cell types and quantifies cellular arrangements from spatial proteomics
Nature Communications (2024)
-
Characterizing the impacts of dataset imbalance on single-cell data integration
Nature Biotechnology (2024)
-
UPP1 promotes lung adenocarcinoma progression through the induction of an immunosuppressive microenvironment
Nature Communications (2024)
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.