Abstract
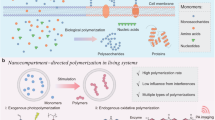
The synthesis of synthetic intracellular polymers offers groundbreaking possibilities in cellular biology and medical research, allowing for novel experiments in drug delivery, bioimaging and targeted cancer therapies. These macromolecules, composed of biocompatible monomers, are pivotal in manipulating cellular functions and pathways due to their bioavailability, cytocompatibility and distinct chemical properties. This protocol details two innovative methods for intracellular polymerization. The first one uses 2-hydroxy-4′-(2-hydroxyethoxy)-2-methylpropiophenone (Irgacure 2959) as a photoinitiator for free radical polymerization under UV light (365 nm, 5 mW/cm2). The second method employs photoinduced electron transfer-reversible addition–fragmentation chain-transfer polymerization with visible light (470 nm, 100 mW/cm2). We further elaborate on isolating these intracellular polymers by streptavidin/biotin interaction or immobilized metal ion affinity chromatography for polymers tagged with biotin or histidine. The entire process, from polymerization to isolation, takes ~48 h. Moreover, the intracellular polymers thus generated demonstrate significant potential in enhancing actin polymerization, in bioimaging applications and as a novel avenue in cancer treatment strategies. The protocol extends to animal models, providing a comprehensive approach from cellular to systemic applications. Users are advised to have a basic understanding of organic synthesis and cell biology techniques.
Key points
-
Light-driven free radical and photoinduced electron transfer-reversible addition–fragmentation chain-transfer polymerization synthesis of structurally defined cellular polymers from biocompatible monomers in the presence of light-sensitive catalysts can modulate cell function and is a potential anticancer therapy.
-
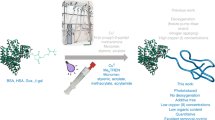
The protocol includes guidelines for the design of biotin and/or histidine-tagged polymers and streptavidin- and/or immobilized metal affinity chromatography-based strategies for their isolation from cell lysate for downstream analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Liu, J. & Liu, B. Living cell-mediated in situ polymerization for biomedical applications. Prog. Polym. Sci. 129, 101545 (2022).
Wu, D. et al. Polymerization in living organisms. Chem. Soc. Rev. 52, 2911–2945 (2023).
Laskar, P., Varghese, O. P. & Shastri, V. P. Advances in intracellular and on-surface polymerization in living cells: implications for nanobiomedicines. Adv. Biomed. Res. 3, 2200174 (2023).
Dergham, M., Lin, S. & Geng, J. Supramolecular self‐assembly in living cells. Angew. Chem. Int. Ed. 61, e202114267 (2022).
Streu, C. & Meggers, E. Ruthenium-induced allylcarbamate cleavage in living cells. Angew. Chem. Int. Ed. 45, 5645 (2006).
Yusop, R. M., Unciti-Broceta, A., Johansson, E. M., Sánchez-Martín, R. M. & Bradley, M. Palladium-mediated intracellular chemistry. Nat. Chem. 3, 239–243 (2011).
Tomás-Gamasa, M., Martínez-Calvo, M., Couceiro, J. R. & Mascareñas, J. L. Transition metal catalysis in the mitochondria of living cells. Nat. Commun. 7, 12538 (2016).
Devaraj, N. K., Thurber, G. M., Keliher, E. J., Marinelli, B. & Weissleder, R. Reactive polymer enables efficient in vivo bioorthogonal chemistry. Proc. Natl Acad. Sci. USA 109, 4762–4767 (2012).
Wang, J., Wang, X., Fan, X. & Chen, P. R. Unleashing the power of bond cleavage chemistry in living systems. ACS Cent. Sci. 7, 929–943 (2021).
Jasinski, F., Zetterlund, P. B., Braun, A. M. & Chemtob, A. Photopolymerization in dispersed systems. Prog. Polym. Sci. 84, 47–88 (2018).
Noè, C., Hakkarainen, M. & Sangermano, M. Cationic UV curing of epoxidized biobased resins. Polymers 13, 89 (2021).
Yuan, Y., Li, C., Zhang, R., Liu, R. & Liu, J. Low volume shrinkage photopolymerization system using hydrogen-bond-based monomers. Prog. Org. Coat. 137, 105308 (2019).
Khudyakov, I. V., Legg, J. C., Purvis, M. B. & Overton, B. J. Kinetics of photopolymerization of acrylates with functionality of 1–6. Ind. Eng. Chem. Res. 38, 3353–3359 (1999).
Xiao, P. et al. Visible light sensitive photoinitiating systems: recent progress in cationic and radical photopolymerization reactions under soft conditions. Prog. Polym. Sci. 41, 32–66 (2015).
Bonardi, A. et al. High performance near-infrared (NIR) photoinitiating systems operating under low light intensity and in the presence of oxygen. Macromolecules 51, 1314–1324 (2018).
Zhao, H. et al. Spatiotemporal control of polymer brush formation through photoinduced radical polymerization regulated by DMD light modulation. Lab Chip 19, 2651–2662 (2019).
Xi, W. et al. Spatial and temporal control of thiol-Michael addition via photocaged superbase in photopatterning and two-stage polymer networks formation. Macromolecules 47, 6159–6165 (2014).
Lang, M., Hirner, S., Wiesbrock, F. & Fuchs, P. A review on modeling cure kinetics and mechanisms of photopolymerization. Polymers 14, 2074 (2022).
Geng, J. et al. Radical polymerization inside living cells. Nat. Chem. 11, 578–586 (2019).
Zhang, Y. et al. Controlled intracellular polymerization for cancer treatment. JACS Au. 2, 579–589 (2022).
Rinnerthaler, M., Bischof, J., Streubel, M. K., Trost, A. & Richter, K. Oxidative stress in aging human skin. Biomolecules 5, 545–589 (2015).
Xu, J., Shanmugam, S., Duong, H. T. & Boyer, C. Organo-photocatalysts for photoinduced electron transfer-reversible addition–fragmentation chain transfer (PET–RAFT) polymerization. Polym. Chem. 6, 5615–5624 (2015).
Xu, Y., Chen, C., Hellwarth, P. B. & Bao, X. Biomaterials for stem cell engineering and biomanufacturing. Bioact. Mater. 4, 366–379 (2019).
Murakami, T. et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron 88, 678–690 (2015).
Shen, Q., Huang, Y., Bai, H., Lv, F. & Wang, S. Polymer materials synthesized through cell-mediated polymerization strategies for regulation of biological functions. Acc. Mater. Res. 4, 57–70 (2022).
Chen, Y. et al. Nanocompartment-confined polymerization in living systems. Nat. Commun. 14, 5229 (2023).
Hai, Z. et al. γ-Glutamyltranspeptidase-triggered intracellular gadolinium nanoparticle formation enhances the T2-weighted MR contrast of tumor. Nano Lett. 19, 2428–2433 (2019).
Shen, Q. et al. Intracellular radical polymerization of paclitaxel-bearing acrylamide for self-inflicted apoptosis of cancer cells. ACS Mater. Lett. 3, 1307–1314 (2021).
Li, L.-L. et al. Intracellular construction of topology-controlled polypeptide nanostructures with diverse biological functions. Nat. Commun. 8, 1276 (2017).
Liu, C. et al. Intracellular hyperbranched polymerization for circumventing cancer drug resistance. ACS Nano 17, 11905–11913 (2023).
Marino, A. et al. Nanostructured Brownian surfaces prepared through two-photon polymerization: investigation of stem cell response. ACS Nano 8, 11869–11882 (2014).
Han, S. B., Kim, J. K., Lee, G. & Kim, D. H. Mechanical properties of materials for stem cell differentiation. Adv. Biosyst. 4, 2000247 (2020).
Ghosh, P. & De, P. Modulation of amyloid protein fibrillation by synthetic polymers: Recent advances in the context of neurodegenerative diseases. ACS Appl. Bio Mater. 3, 6598–6625 (2020).
Sundelacruz, S. & Kaplan, D. L. Stem cell-and scaffold-based tissue engineering approaches to osteochondral regenerative medicine. Semin. Cell Dev. Biol. 20, 646–655 (2009).
Liang, G., Ren, H. & Rao, J. A biocompatible condensation reaction for controlled assembly of nanostructures in living cells. Nat. Chem. 2, 54–60 (2010).
Rong, L. H., Caldona, E. B. & Advincula, R. C. PET–RAFT polymerization under flow chemistry and surface‐initiated reactions. Polym. Int. 72, 145–157 (2023).
Parkatzidis, K., Wang, H. S., Truong, N. P. & Anastasaki, A. Recent developments and future challenges in controlled radical polymerization. Chem 6, 1575–1588 (2020).
Xie, W., Zhao, L., Wei, Y. & Yuan, J. Advances in enzyme-catalysis-mediated RAFT polymerization. Cell Rep. Phys. Sci. 2, 100487 (2021).
Wu, J., Lin, J. & Huang, P. Harnessing abiotic organic chemistry in living systems for biomedical applications. Chem. Soc. Rev. 52, 3973–3990 (2023).
Joshi, T. & Nebhani, L. Light-regulated growth of polymer chains from the surface of RAFT agent primed mesoporous silica nanoparticles. Surf. Interfaces 29, 101764 (2022).
Dufrêne, Y. F. Atomic force microscopy and chemical force microscopy of microbial cells. Nat. Protoc. 3, 1132–1138 (2008).
Stribbling, S. M. & Ryan, A. J. The cell-line-derived subcutaneous tumor model in preclinical cancer research. Nat. Protoc. 17, 2108–2128 (2022).
Heise, C. C., Williams, A., Olesch, J. & Kirn, D. H. Efficacy of a replication-competent adenovirus (ONYX-015) following intratumoral injection: intratumoral spread and distribution effects. Cancer Gene Ther. 6, 499–504 (1999).
Watanabe, S. et al. Transplantation of intestinal organoids into a mouse model of colitis. Nat. Protoc. 17, 649–671 (2022).
Martí, M. et al. Characterization of pluripotent stem cells. Nat. Protoc. 8, 223–253 (2013).
Kaiser, E., Colescott, R. L., Bossinger, C. D. & CookMartí, P. I. Color test for detection of free terminal amino groups in the solid-phase synthesis of peptides. Anal. Biochem. 34, 595–598 (1970).
Acknowledgements
The authors thank the National Natural Science Foundation of China (22071263), the Guangdong Province Zhujiang Talents Program (2019QN01Y127) and the Shenzhen Fundamental Research Program (JCYJ20200109110215774). We thank M. Galluzzi for the AFM measurement.
Author information
Authors and Affiliations
Contributions
M.A. and Q.G. contributed equally to the work. All authors contributed extensively to the work presented and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Greg Qiao, Dayong Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Geng, J. et al. Nat. Chem. 11, 578–586 (2019): https://doi.org/10.1038/s41557-019-0240-y
Zhang, Y. et al. JACS Au 2, 579–589 (2022): https://doi.org/10.1021/jacsau.1c00373
Extended data
Extended Data Fig. 1 Concentration calibrations of DMA, HPMA, CA-CTA, and Eosin Y and 1H NMR of isolated polymer.
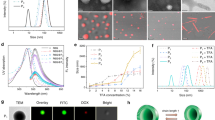
a, UV-vis spectral representation of DMA at concentrations of 1, 5, 10, 20, and 50 mM. b, UV-vis spectra showcasing HPMA at concentrations of 2, 5, 10, 20, and 50 mM. c, UV-vis spectra of CA-CTA at concentrations of 0.02, 0.1, 0.2, 0.5, and 1.0 mM. d, UV-vis spectral plots of Eosin Y at concentrations of 0.002, 0.01, 0.02, 0.05, and 0.1 mM. e, The graphical depiction of cellular uptake data. This data was quantified utilizing UV-vis spectroscopy. f, g, 1H NMR spectra (recorded in D2O) of isolated polymers. f, Displays the 1H NMR spectrum for His-PDMA. The upper spectrum represents the polymer when polymerized in PBS, whereas the lower spectrum is for the polymerized variant in cells. g, Showcases the 1H NMR spectrum for His-PHPMA. As with (f), the upper graph pertains to the polymer processed in PBS, and the lower represents the cell-polymerized version. For both (f) and (g), the polymer isolated from PBS underwent a dialysis process against water with a molecular weight cut-off (MWCO) of 1,000. On the other hand, the polymers derived from cell lysates were isolated using magnetic beads and then released at a temperature of 60 °C. Panels a–e reproduced with permission from ref. 20, American Chemical Society.
Extended Data Fig. 2 Characterizations of His-polymers isolated from HeLa cells.
PET-RAFT polymerization was conducted within cells utilizing a hexa-histidine-tagged CTA known as His-CTA (for detailed polymerization cocktail compositions, refer to Table 1). Polymers synthesized intracellularly were procured through dialysis (with a molecular weight cut-off, MWCO, of 1,000 Da) and IMAC isolation. The following polymers were characterized using 1H NMR (in d6-DMSO): a, His-PHPMA-1. b, His-PHPMA-2. c, His-PDMA-3. d, His-PDMA-1. e, His-PDMA-2. f, His-PDMA-3. g, GPC traces for isolated polymers: His-PHPMA-1, His-PHPMA-2, and His-PHPMA-3. h, GPC traces for isolated polymers: His-PDMA-1, His-PDMA-2, and His-PDMA-3. Figure adapted with permission from ref. 20, American Chemical Society.
Extended Data Fig. 3 Exploration of intracellular retention time of monomers, CTAs, and Eosin Y.
Cells were exposed to PC1, supplemented with YFMA (1.0 µM) and RF-CTA (100 µM), for a duration of 4 h. Then, cells were washed and subjected to either a 10-minute illumination (+hv) or left without illumination (-hv). Representative confocal microscopy images are presented here to show the cellular state immediately after (0 h) and 24 hours post-illumination. The columns, from left to right, depict the cellular uptake and distribution of Eosin Y, monomers, CTAs, and a merged view, respectively. The top two rows showcase cells without illumination (- hν) and the bottom two rows present cells with illumination (+ hν). Scale bar = 50 µm. Figure reproduced with permission from ref. 20, American Chemical Society.
Extended Data Fig. 4 Intracellular polymerization induces cell death.
a, Viability assessment of HeLa, 4T1, and MDA-MB-231 cells post-intracellular polymerization using the CCK-8 assay. Cells underwent treatment with PC1 for a duration of 4 hours, were subsequently washed, and then illuminated for 10 minutes. This was followed by an incubation period of 24 hours. The results are expressed as the mean ± SD, based on 6 independent samples per group. For statistical analysis, one-way ANOVA with Dunnett post-test was utilized, comparing to the untreated cells. Significance levels are indicated as follows: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, with indicating non-significance. b, Evaluation of apoptosis and necroptosis levels induced by intracellular polymerization, determined using flow cytometry. The cells undergoing apoptosis are Annexin-V-positive and PI-negative, while those undergoing necroptosis are indicated in the graph. Numerical analysis regarding the number of cells evaluated and the absolute numbers or percentages of the relevant cell populations within post-sort fractions is provided in the figure. c, Quantitative representation of apoptosis and necroptosis levels from flow cytometry data presented in panel b. Data are presented as percentages of cells undergoing apoptosis and necroptosis. Significance levels are indicated as: *P < 0.05, **P < 0.01, ****P < 0.0001, ns (not significant). d, e, Analysis of crucial biological markers using techniques such as immunoblotting and immunoprecipitation assays. The displayed bands correspond to markers like PARP, C-PARP, p53, BCL-2, BAX, AIF, and γ-H2AX. The molecular weight (in kDa) of each protein is indicated on the right. Figure adapted with permission from ref. 20, American Chemical Society.
Extended Data Fig. 5 Intracellular polymerization inhibited proliferation.
a, Colony growth assay depicting the impact of intracellular polymerization on the proliferative ability of HeLa cells. Left panel shows cells treated with PBS in the absence of light, while the right panel displays cells under similar conditions but with light exposure. The scale bar is indicative of 1 cm. b, Representation of cell cycle phases for HeLa cells subjected to treatments with or without intracellular polymerization. The four groups depicted are cells treated with PBS in the absence and presence of light, and cells treated with PC1 in the absence and presence of light. The percentages of cells in the G0/G1, S, and G2/M phases of the cell cycle are presented. Data were based on n = 6 samples. c, Immunoblotting assays highlight the expression levels of key cell cycle regulators, specifically Cyclin B1 and Cyclin E1, in HeLa cells under different treatment conditions. GAPDH serves as the loading control, and the molecular weight (in kDa) of each protein is denoted on the right. Statistical evaluations were executed using one-way ANOVA accompanied by a Dunnett post-test in comparison with the untreated control groups. Levels of significance are denoted as: *P < 0.05, ***P < 0.001, ****P < 0.0001, ns (not significant). Figure reproduced with permission from ref. 20, American Chemical Society.
Extended Data Fig. 6 analysis of cell mobility and physiological changes induced by intracellular polymerization.
a, Representative wound-healing assay images capture cell mobilities across different treatment conditions and time points. Each image denotes the percentage of the area not covered by cells in the wound area at the indicated time post-treatment. b, Quantitative representation of the wound-healing assay in A, presenting the normalized gap percentage over the incubation periods of 0, 24, 48, and 72 hours. Data are from n = 3 samples per condition. c, Immunoblotting assays showing the expression levels of cell-motility-associated proteins E-cadherin, Snail, and Vimentin. GAPDH is used as the loading control, with the molecular weight (in kDa) for each protein mentioned on the right. d, Illustration of the viscosity p’obe’s molecular structure, which is activated at an excitation/emission wavelength (λex/em) of 640/660 nm. The increase in viscosity corresponds to a higher fluorescence intensity. e, Flow cytometric analysis detailing cellular viscosity across various treatment groups. The measurements employ a Cy5-based viscosity probe, with the fluorescence intensities depicted on the x-axis. f, Atomic force microscopy results indicating the stiffness of cells subjected to different treatments. In all figures, data points represent individual measurements with mean values shown. Statistical analysis was performed using one-way ANOVA with a Dunnett post-test compared to untreated control groups, ****P < 0.0001, ns (not significant). Panels a–e reproduced with permission from ref. 20, American Chemical Society.
Extended Data Fig. 7 In vivo evaluation of intracellular polymerization for cancer treatment.
a–c, Tumor size progression in BALB/c nude mice bearing different tumor models: HeLa (a), MDA-MB-231 (b), and 4T1 (c) over a specified number of days. Mice were subjected to different treatment regimens, and data are representative of n = 5 mice per group. Error bars represent for standard deviations. d, Histological analyses of tumor sections stained with hematoxylin/eosin (H&E), TUNEL, and Ki67. Each staining method illuminates different features within the tumor tissue, including cellular morphology, apoptosis, and proliferation, respectively. Red arrowheads highlight areas of interest. e, f, Quantitative evaluations of the tumor sections. The number of TUNEL-positive cells (E) and Ki67-positive cells (F) are represented. Cell numbers were quantified from six random areas within the tissue, with each experiment conducted with n = 6 samples. Statistical evaluations were executed using one-way ANOVA accompanied by a Dunnett post-test in comparison with the untreated control groups. Levels of significance are denoted as: *P < 0.05, ***P < 0.001, ****P < 0.0001, ns (not significant). g, Visual representation of the xenografted tumors at day 14 post various treatments, showcasing the size reduction in response to intracellular polymerization. h, Comprehensive hematoxylin and eosin (H&E) stained lung sections taken from mice on day 19 post-treatment. These illustrations offer insights into potential metastatic spread, with metastatic sites demarcated by black dashed lines. Scale notations: For H&E images in (d), the scale bar corresponds to 1 mm, whereas for the TUNEL and Ki67 images, the scale bar represents 200 μm and 2 mm in (h). Panels b–h reproduced with permission from ref. 20, American Chemical Society.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and Table 1.
Source data
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abdelrahim, M., Gao, Q., Zhang, Y. et al. Light-mediated intracellular polymerization. Nat Protoc (2024). https://doi.org/10.1038/s41596-024-00970-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41596-024-00970-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.